-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaNODULE INCEPTION Directly Targets Subunit Genes to Regulate Essential Processes of Root Nodule Development in
The interactions of legumes with symbiotic nitrogen-fixing bacteria cause the formation of specialized lateral root organs called root nodules. It has been postulated that this root nodule symbiosis system has recruited factors that act in early signaling pathways (common SYM genes) partly from the ancestral mycorrhizal symbiosis. However, the origins of factors needed for root nodule organogenesis are largely unknown. NODULE INCEPTION (NIN) is a nodulation-specific gene that encodes a putative transcription factor and acts downstream of the common SYM genes. Here, we identified two Nuclear Factor-Y (NF-Y) subunit genes, LjNF-YA1 and LjNF-YB1, as transcriptional targets of NIN in Lotus japonicus. These genes are expressed in root nodule primordia and their translational products interact in plant cells, indicating that they form an NF-Y complex in root nodule primordia. The knockdown of LjNF-YA1 inhibited root nodule organogenesis, as did the loss of function of NIN. Furthermore, we found that NIN overexpression induced root nodule primordium-like structures that originated from cortical cells in the absence of bacterial symbionts. Thus, NIN is a crucial factor responsible for initiating nodulation-specific symbiotic processes. In addition, ectopic expression of either NIN or the NF-Y subunit genes caused abnormal cell division during lateral root development. This indicated that the Lotus NF-Y subunits can function to stimulate cell division. Thus, transcriptional regulation by NIN, including the activation of the NF-Y subunit genes, induces cortical cell division, which is an initial step in root nodule organogenesis. Unlike the legume-specific NIN protein, NF-Y is a major CCAAT box binding protein complex that is widespread among eukaryotes. We propose that the evolution of root nodules in legume plants was associated with changes in the function of NIN. NIN has acquired functions that allow it to divert pathways involved in the regulation of cell division to root nodule organogenesis.
Published in the journal: . PLoS Genet 9(3): e32767. doi:10.1371/journal.pgen.1003352
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003352Summary
The interactions of legumes with symbiotic nitrogen-fixing bacteria cause the formation of specialized lateral root organs called root nodules. It has been postulated that this root nodule symbiosis system has recruited factors that act in early signaling pathways (common SYM genes) partly from the ancestral mycorrhizal symbiosis. However, the origins of factors needed for root nodule organogenesis are largely unknown. NODULE INCEPTION (NIN) is a nodulation-specific gene that encodes a putative transcription factor and acts downstream of the common SYM genes. Here, we identified two Nuclear Factor-Y (NF-Y) subunit genes, LjNF-YA1 and LjNF-YB1, as transcriptional targets of NIN in Lotus japonicus. These genes are expressed in root nodule primordia and their translational products interact in plant cells, indicating that they form an NF-Y complex in root nodule primordia. The knockdown of LjNF-YA1 inhibited root nodule organogenesis, as did the loss of function of NIN. Furthermore, we found that NIN overexpression induced root nodule primordium-like structures that originated from cortical cells in the absence of bacterial symbionts. Thus, NIN is a crucial factor responsible for initiating nodulation-specific symbiotic processes. In addition, ectopic expression of either NIN or the NF-Y subunit genes caused abnormal cell division during lateral root development. This indicated that the Lotus NF-Y subunits can function to stimulate cell division. Thus, transcriptional regulation by NIN, including the activation of the NF-Y subunit genes, induces cortical cell division, which is an initial step in root nodule organogenesis. Unlike the legume-specific NIN protein, NF-Y is a major CCAAT box binding protein complex that is widespread among eukaryotes. We propose that the evolution of root nodules in legume plants was associated with changes in the function of NIN. NIN has acquired functions that allow it to divert pathways involved in the regulation of cell division to root nodule organogenesis.
Introduction
The interactions of legume plants with their bacterial symbionts, collectively called “rhizobia,” cause the formation of specialized root lateral organs called root nodules. Unlike lateral roots, of which initiation is regulated by endogenous signals, activation of the mitotic cell cycle for root nodule organogenesis is triggered by symbiont-derived signaling molecules called nodulation (Nod) factors [1]–[7]. Legumes have developed unique molecular networks that transmit exogenous signals to the regulatory factors for organ development. Over the past decade, many factors involved in nodulation processes have been identified, leading to basic models for the nodulation signaling pathways. However, the mechanisms by which nodulation signals induce cell division have not been elucidated.
Nodulation processes are initiated by the adhesion of rhizobia to root hairs in the model legume plants Lotus japonicus and Medicago truncatula [8]. Symbiotic bacteria are entrapped by curled root hairs and form microcolonies on the host epidermal cells. Subsequently, the invasion of plant tissues by the bacterial symbionts is mediated by host cell-derived tubular structures called infection threads. The infection threads develop by invagination of the plasma membrane at the infection foci, where the plant cell wall is degraded. Concomitant with the progression of infection processes at the epidermis, a fraction of cortical cells beneath the site of infection begin to divide and form a root nodule primordium.
Forward genetics studies in the two model legume plants have revealed that early signaling pathway(s) downstream of Nod factor perception are common to those required for mycorrhizal symbiosis. Genes involved in both symbiosis systems are referred to as the common SYM genes [9]. It has been postulated that the root nodule symbiosis systems have recruited functions partly from the ancestral mycorrhizal symbiosis systems. Among the proteins encoded by the common SYM genes, CCaMK (calcium/calmodulin-dependent protein kinase) plays a pivotal role in both symbioses. This protein is thought to act as a decoder of perinuclear Ca2+ oscillations triggered by Nod factor perception [7]. Gain-of-function (gof)-CCaMK rescues nodulation - and mycorrhizal infection-defective phenotypes caused by mutations in common SYM genes that are required for the Ca2+ oscillation [10], [11]. Furthermore, gof-CCaMK induces spontaneous root nodules in the absence of rhizobia [12], [13], indicating that this protein bypasses an early nodulation signaling pathway from Nod factor perception. The common SYM pathway is thought to transmit nodulation signals to pathway(s) that have evolved specifically to regulate root nodule organogenesis. In so doing it diverts the plant's general gene expression networks to execute complex nodulation processes.
In the current model, CCaMK-induced root nodule organogenesis is mediated by a cytokinin receptor called Lotus histidine kinase 1 (LHK1) [10], [11], [14], [15]. Cytokinin is a phytohormone that regulates various aspects of plant development [16]. Loss-of-function mutants of LHK1 (hit1) and its M. truncatula counterpart, CYTOKININ RESPONSE1 (MtCRE1), fail to initiate timely cortical cell division in response to rhizobial signals [15], [17]. Gof-LHK1 causes the development of spontaneous root nodules in loss-of-function ccamk mutants without rhizobial infection [10], [11], [14]. Exogenously applied cytokinin also causes the formation of spontaneous root nodules [18]. These results implicate cytokinin as an endogenous molecular signal for initiating root nodule organogenesis, and this signal must necessarily be integrated with nodulation specific pathways. NODULATION SIGNALING PATHWAY1 (NSP1), NSP2, and NODULE INCEPTION (NIN) are required for CCaMK - and LHK1 - mediated root nodule organogenesis [10], [11]. Unlike LHK1 and MtCRE1, these genes are essential for both the rhizobial infection processes in the epidermis and the cortical responses [19], [20], [21]. They are thought to function in nodulation-specific processes [20], [22], [23], although recent findings have implied that NSP1 and NSP2 also act in mycorrhizal symbiosis [24], [25]. NSP1 and NSP2 encode GRAS family transcription factors [26], [27]. NIN encodes a putative transcription factor with a RWP-RK domain [19]. The absence of NSP1, NSP2, or NIN function results in abortion of spontaneous root nodule formation by gof-CCaMK and gof-LHK1 [10], [11]. Transcriptional regulation by these factors is important for root nodule organogenesis. However, the mechanisms by which these factors mediate rhizobial infection signals to activate developmental programs underlying nodulation have not yet been elucidated.
NIN was the first gene to be genetically characterized for its role in the regulation of nodulation processes [19]. This gene is involved in multiple processes including symbiotic root hair responses and infection thread formation in the epidermis, and induction of cell division in the cortex [19], [28]. NIN is expressed during the early stages of organogenesis in root nodule primordia generated by exogenously applied cytokinin [18]. Cytokinin-induced NIN expression is downregulated by mutations in LHK1 and MtCRE1 [15], [17]. On the other hand, the expression of both NSP1 and NSP2 is less sensitive to or downregulated by exogenous cytokinin [17], [29], although expression of these genes is required for NIN expression induced by rhizobial infection [23], [28]. The two GRAS transcription factors form a heterodimer and bind to the M. truncatula NIN promoter in vitro [30]. The expression pattern of NIN implies that this gene may be a primary regulator of cortical cell division in response to cytokinin signaling. Despite the functional importance of NIN in root nodule-specific symbiotic events, our knowledge of its function is at the genetic level. The precise function of NIN in nodulation, and the molecular properties of the protein, have not yet been fully elucidated. The identification of genes that regulate cortical cell division downstream of NIN will be important for understanding the molecular mechanisms underlying root nodule organogenesis.
In this study we biochemically and biologically dissected NIN protein function. We found that NIN acts as a transcriptional activator that induces cortical cell division in the absence of bacterial symbionts. Furthermore, we determined that genes encoding different subunits of Nuclear Factor Y (NF-Y) are direct targets of NIN. NF-Y is a heterotrimeric CCAAT box binding protein complex composed of subunits A, B, and C [31]. The genes that we identified as NIN targets encode subunits A and B. We named these genes LjNF-YA1 and LjNF-YB1. The product of LjNF-YA1 is orthologous to M. truncatula HAP2-1, which is involved in meristem persistence in indeterminate-type root nodules [32]. Our functional analyses indicate that the NF-Y genes play overlapping roles with that of NIN in root nodule organogenesis. Unlike the legume-specific NIN protein, NF-Y is a general factor widespread among eukaryotes. Ectopic expression of NIN and the NF-Y subunit genes also influenced cell division in lateral root primordia that is not related to root nodule organogenesis. We propose that NIN is a mediator between rhizobial infection signals and general regulatory mechanisms associated with cell proliferation.
Results
Identification of candidate NIN target genes
NIN encodes a protein containing a RWP-RK domain, which is conserved among all plants from algae to seed plants [33]. Predictions of the secondary structure of this domain suggest that it binds to DNA [19]. Xie et al showed that NIN binds to the promoter of the L. japonicus nodulation pectate lyase gene [34]. Therefore, NIN is predicted to be a transcription factor. Supporting this idea, a NIN-GFP fusion protein that was functional in L. japonicus root epidermal responses (Figure S1C) localized to nuclei in Nicotiana benthamiana leaves (Figure S1A, S1B), as did the Arabidopsis NIN-like protein 7 [35]. In addition, the NIN protein fused to the GAL4 DNA-binding domain induced expression of a GFP-GUS reporter that had four tandem repeats of a GAL4 target nucleotide sequence in its promoter (Figure S1D–S1F) [36]. These results suggest that NIN is a positive regulator of gene expression.
To search for candidate genes that are targeted by NIN, we took advantage of the publicly available transcriptome database (http://cgi-www.cs.au.dk/cgi-compbio/Niels/index.cgi) [37] and selected 19 genes as possible candidates (Figure S2A). The NIN-dependent expression of 9 candidates was examined and validated by real time (RT)-PCR (Figure S2B).
We performed knockdown analyses of several of these candidate NIN targets to investigate their functions in nodulation processes, and found that RNA interference (RNAi) of chr5.CM0571.340.r2.m inhibited root nodule formation (see below). chr5.CM0571.340.r2.m encodes NF-YA, which forms a heterotrimeric CCAAT box-binding protein complex with other two subunits, NF-YB and NF-YC. We further found the NF-YB gene (LjSGA_022269.1) among the other candidate NIN targets (Figure S2A). NF-Y subunits are required for nodulation processes in M. truncatula [32] and Phaseolus vulgaris [38]. In particular, MtHAP2-1, which is orthologous to chr5.CM0571.340.r2.m (Figure S3), is involved in the maintenance of meristem activity in indeterminate-type root nodules [32]. Furthermore, mammalian NF-Y also functions in the regulation of cell division [39]–[42]. We expected that the Lotus NF-Y subunits would be included in the same NF-Y complex, and that this complex might function in nodulation processes downstream of NIN. Therefore, we focused on the Lotus NF-Y subunit genes for further analyses, and named them LjNF-YA1 (chr5.CM0571.340.r2.m) and LjNF-YB1 (LjSGA_022269.1).
LjNF-YA1 and LjNF-YB1 are direct transcriptional targets of NIN
To examine whether NIN overexpression induces expression of LjNF-YA1 and LjNF-YB1, we expressed a NIN-GR fusion protein, in which the glucocorticoid receptor (GR) was fused to the C-terminus of NIN, in nin-2 mutant roots. The overexpression construct was driven by the CaMV35S promoter (Pro35S). The transgenic roots were treated with or without dexamethasone (DEX) and cycloheximide (CHX). This recombinant protein is functional in Lotus roots, because when the gene construct was driven by the NIN promoter in the presence of DEX, it suppressed the infection thread-defective nin-2 phenotype (Figure S4A and S4B). RT-PCR analysis showed that DEX treatment of roots transformed with the overexpression construct induced expression of LjNF-YA1 and LjNF-YB1 within 4 h, and that further incubation (20 h) resulted in increased levels of expression (Figure 1). Therefore, NIN induces expression of these genes. CHX treatment did not repress the expression induced by DEX. In the cases of LjNF-YB1 and another candidate NIN target, chr4.CM0179.190.r2.m, which encodes a plastocyanin-like domain-containing protein (PLDP; see Figure S2A), CHX treatment enhanced the expression induced by DEX (Figure 1). These results support the idea that LjNF-YA1 and LjNF-YB1 are primary targets of NIN.
Fig. 1. Expression of candidate NIN-target genes in roots that were ectopically expressing the NIN protein. 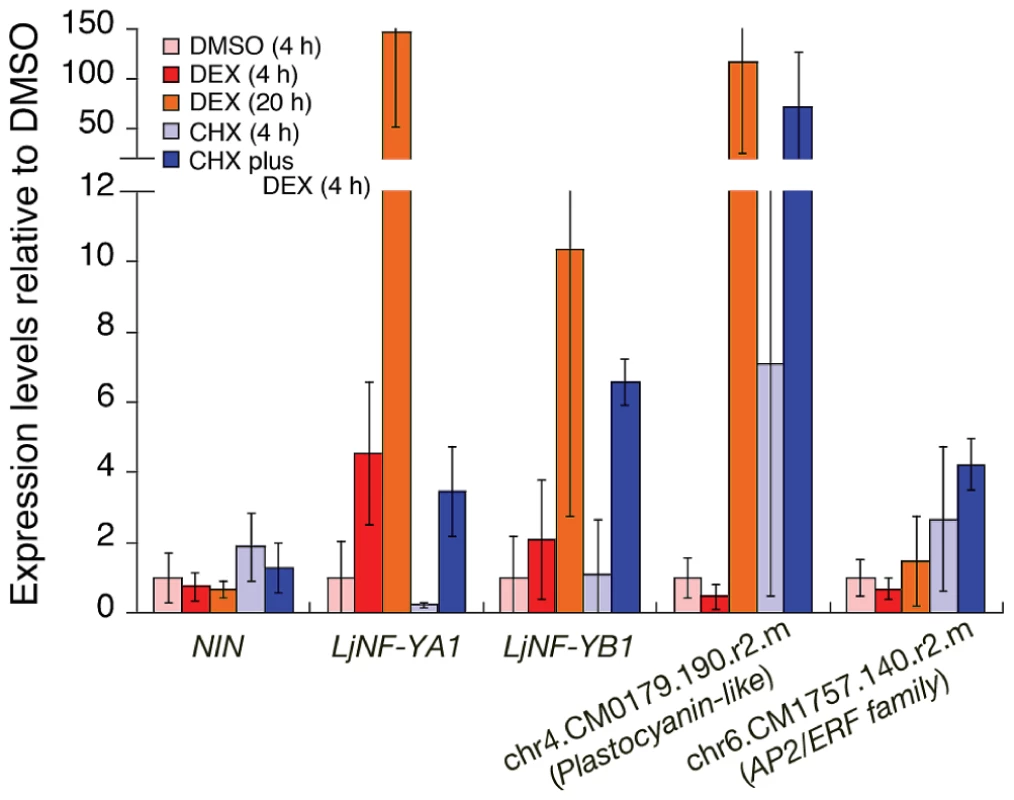
RT-PCR was used to analyze gene expression in nin-2 roots that were transformed with Pro35S-NIN-GR. Roots were treated as indicated in the figure. For CHX plus DEX, the DEX was added after pre-incubation with CHX for 30 min. chr4.CM0179.190.r2.m encodes a plastocyanin-like domain-containing protein (PLDP) and chr6.CM1757.140.r2.m encodes an AP2/ERF family protein. These genes are listed in Figure S2. The promoter of the former gene possesses NIN-binding nucleotide sequences (see Figure 2). The latter gene was used as a negative control for DEX treatment. The means and SDs from 3 biological repeats are shown. We performed a chromatin immunoprecipitation (ChIP) analysis to investigate NIN binding to the LjNF-YA1 and LjNF-YB1 promoters in vivo. Chromatin suspensions were prepared from roots that were transformed with either ProLjUb-NIN-myc or an empty vector. The NIN-myc protein was expressed from the L. japonicus polyubiquitin promoter (ProLjUb) [43]. The recombinant NIN protein suppressed the infection thread-defective nin-2 phenotype when expressed using the NIN promoter, indicating that this fusion protein is functional in planta (Figure S4C). Several primer sets for detecting LjNF-YA1 and LjNF-YB1 promoter fragments (indicated by blue lines in Figure 2A) were designed to cover the whole promoter regions that correspond to those used for spatial expression analyses using GUS reporter constructs (see Figure 3). Using anti-myc antibodies we detected enrichment of LjNF-YA1 promoter region 3 and LjNF-YB1 promoter regions 2 and 3 in immunoprecipitates of chromatin suspensions derived from roots expressing ProLjUb-NIN-myc (Figure 2A, 2B). The level of enrichment of LjNF-YB1 promoter region 3 was approximately three times greater than those of LjNF-YA1 promoter region 3 and LjNF-YB1 promoter region 2. These results indicate that NIN binds to these promoter regions in vivo.
Fig. 2. NIN directly targets LjNF-YA1 and LjNF-YB1. 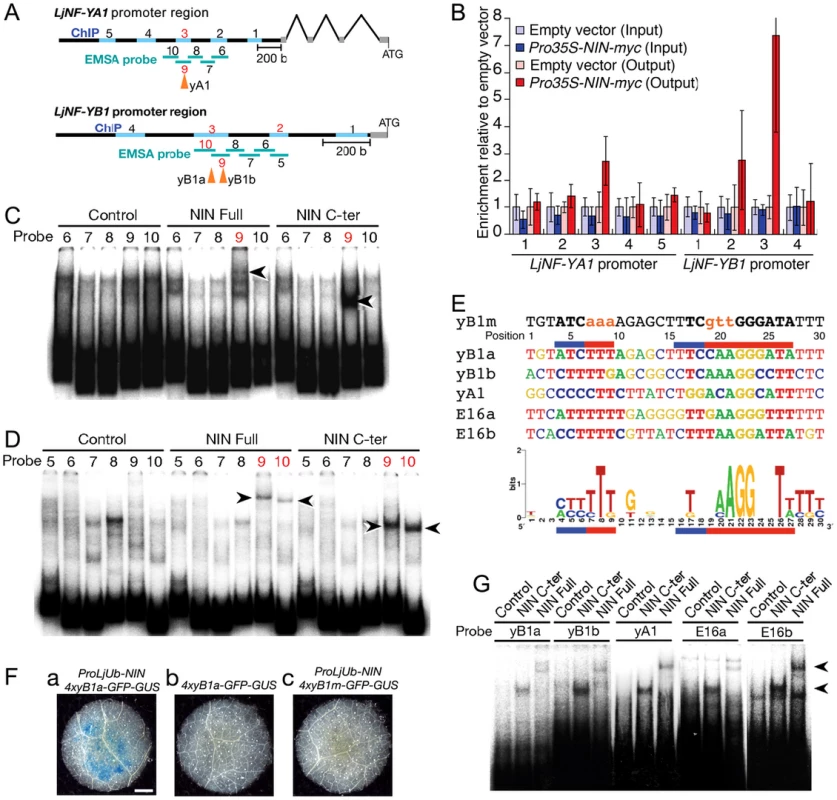
(A) A diagram of the LjNF-YA1 and LjNF-YB1 promoter regions. These regions were used for the promoter-GUS reporters (see Figure 3). Gray lines indicate 5′-UTRs. Regions analyzed by RT-PCR for the ChIP assays in (B) are shown as blue lines and probes used for EMSA in (C) and (D) are shown as green lines. The red numbers indicate regions and probes that gave positive results in the ChIP assays and EMSAs. Arrowheads indicate positions of the NIN-binding sites. (B) ChIP assays using either ProLjUb-NIN-myc roots or control (empty vector) roots. The means and SDs from 3 biological repeats are shown. (C,D) EMSAs for analyzing NIN binding to LjNF-YA1 (C) and LjNF-YB1 (D) promoter regions. NIN-myc (Full), NIN(520–878)-myc (C-ter), and in vitro translation products without templates (control) were incubated with 32P-labeled probes shown in (A). Arrowheads indicate mobility-shifted bands specifically detected when incubated with NIN proteins. (E) An alignment of the partial nucleotide sequences of probes that were bound by NIN. yB1a, yB1b, yA1, E16a, are E16b correspond to NBSs found in the promoters of LjNF-YB1, LjNF-YA1, and the PLDP-encoding gene. Red and blue lines indicate nucleotides that are required for NIN-binding to NBS-yB1a, or that influence binding, respectively (see Figure S5). NBS-yB1m is an NBS-yB1a derivative with nucleotide substitutions. Comparison of NIN-binding sequences illustrated by logo is shown at the bottom. (F) GUS expression in tobacco leaf disks transformed with the indicated constructs. Bar: 1 mm. (G) EMSA for analyzing NIN binding to NBS-yB1a and the NBS-yB1a-like sequences. Arrowheads indicate probes that were bound by NIN proteins. Fig. 3. Spatial expression patterns of LjNF-YA1 and LjNF-YB1 during root nodule organogenesis and their protein interactions in planta. 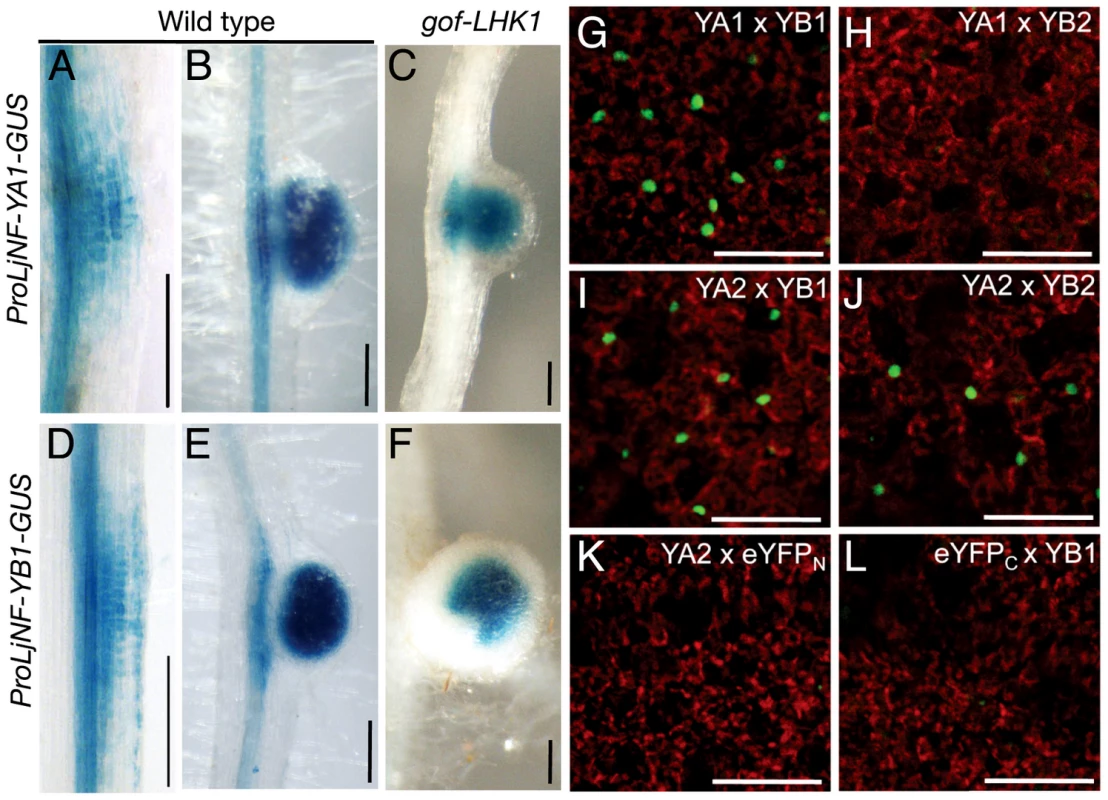
(A–F) GUS expression in roots that were transformed with either ProLjNF-YA1-GUS (A–C) or ProLjNF-YB1-GUS (D–F). GUS staining was detected in root nodule primordia (A, D), developing root nodules (B, E), and in spontaneous root nodules caused by Pro35S-gof-LHK1 (C, F). (G–L) BiFC analysis of interactions between LjNF-YA and LjNF-YB subunits in N. benthamiana leaves. Confocal images of eYFP fluorescence (green) from nuclei and chloroplast autofluorescence (red) are shown. ProLjUb-eYFPC-LjNF-YA1 was co-introduced with either ProLjUb-eYFPN-LjNF-YB1 (G) or ProLjUb-eYFPN-LjNF-YB2 (H). ProLjUb-eYFPC-LjNF-YA2 was co-introduced with either ProLjUb-eYFPN-LjNF-YB1 (I), ProLjUb-eYFPN-LjNF-YB2 (J), or ProLjUb-eYFPN (K). ProLjUb-eYFPN-LjNF-YB1 was co-introduced with ProLjUb-eYFPC (L). Bars: 0.2 mm in (A–F); 50 µm in (G–L). We also tested for in vitro binding of NIN to these promoter regions using electrophoresis mobility shift assays (EMSAs) with the NIN-myc protein and a NIN(520–878)-myc protein. The latter protein is the C-terminal half of NIN that contains the RWP-RK domain and the PB1 domain [19], [33]. Shifted bands were detected when LjNF-YA1 probe 9 and LjNF-YB1 probes 9 and 10 were incubated with the NIN proteins (Figure 2C, 2D). Differences in the mobilities between NIN-myc and NIN(520–878)-myc indicated that these mobility shifts were due to binding of the NIN proteins. These results show that NIN interacts with the LjNF-YA1 and LjNF-YB1 promoter regions that were enriched in the ChIP analysis, probably through the RWP-RK domain.
Identification of NIN-binding nucleotide sequences
We performed detailed EMSAs to identify the DNA sequences bound by NIN in the LjNF-YB1 promoter. An LjNF-YB1 promoter region corresponding to probes 9 and 10 was divided into 6 shorter sequences that were used as probes (Figure S5Aa; probes 11–16). NIN(520–878)-myc bound only to probe 14 (Figure S5B). Assays using five derivatives of probe 14, each with a six-nucleotide substitution (see Figure S5Ab; m1 to m5), narrowed down the NIN-binding site to a sequence of 31 bp covered by the mutations in m3 to m5 (Figure S5C). Further analyses using probes with three-nucleotide substitutions revealed that two separate parts of the 31 bp sequence were required for NIN binding (Figure 2E; Figure S5Ab, S5D). We refer to this NIN-binding nucleotide sequence (NBS) as NBS-yB1a. To examine whether NBS-yB1a confers NIN-dependent expression on a GFP-GUS reporter gene, a fragment with four tandem repeats of NBS-yB1a was inserted upstream of a CaMV35S minimal promoter (4xyB1a-GFP-GUS). GUS staining was detected in tobacco leaves when the reporter construct was co-introduced with ProLjUb-NIN (Figure 2Fa,b). On the other hand, a similar construct with mutations in the NBS (4xyB1m-GFP-GUS; see Figure 2E) did not show GFP-GUS expression even if ProLjUb-NIN was co-introduced (Figure 2Fc). These results indicate that 4 tandem repeats of NBS-yB1a are sufficient for NIN-dependent gene expression.
We then searched for additional DNA sequences similar to NBS-yB1a, and found two NBS-yB1a-like sequences, referred to as NBS-yA1 and -yB1b, respectively, in the LjNF-YA1 and LjNF-YB1 promoters (Figure 2E). These sequences were included in the regions that were bound by the NIN protein (Figure 2A). NBS-yB1b overlaps with the region covered by probes 11, 12, and 13 in the LjNF-YB1 promoter (Figure S5Ab). NBS-yB1a, NBS-yB1b, and NBS-yA1 are located 755, 712, and 1725 bp upstream of their respective putative translation initiation codons. We also found two NBS-yB1a-like sequences (NBS-E16a and -E16b) in the promoter of the PLDP-encoding gene that was also identified as a NIN target (Figure 2E). These sequences are located 363 and 193 bp upstream of the putative translation initiation codon. This PLDP-encoding gene was induced by the NIN-GR protein in the presence of CHX, as were LjNF-YA1 and LjNF-YB1 (Figure 1; Figure S2). ChIP analysis showed that NIN bound to the promoter of the PLDP-encoding gene in vivo (Figure S6). On the other hand, NBS-yB1a-like sequences were not found within 3 kb of the putative translation initiation codon of chr6.CM1757.140.r2.m, which encodes an AP2/ERF family protein. Expression of this gene was not induced by ectopic NIN expression (Figure 1). EMSAs showed that NIN proteins bound to probes containing the identified NBSs in vitro (Figure 2G). The specificity of NIN binding to these NBSs was confirmed using competition analyses (Figure S5E). These results are in agreement with the idea that NIN directly targets the promoters of LjNF-YA1, LjNF-YB1, and the PLDP-encoding gene and activates their transcription. A consensus of the NBSs identified here is shown in Figure 2E. A region on the left (positions 4–9) and one on the right (positions 16–27) of the consensus sequence corresponded to those required for NIN binding in NBS-yB1a. The left region was rich in T and C and was less conserved than the right region. The right region contained AGG at positions 21–23 and T at position 26, and these were present in all the NBSs that we identified in this study.
LjNF-YA1 and LjNF-YB1 are co-expressed in the root nodule primordium
We used GUS reporter constructs to investigate the expression patterns of LjNF-YA1 and LjNF-YB1 during root nodule organogenesis. The promoter regions of LjNF-YA1 and LjNF-YB1 (2.8 and 1.5 kb upstream from the putative translation initiation codon, respectively; also see Figure 2A) were inserted upstream of the GUS reporter gene. These constructs were introduced into L. japonicus roots by Agrobacterium rhizogenes-mediated transformation. Histochemical GUS analysis revealed that ProLjNF-YA1-GUS and ProLjNF-YB1-GUS were expressed in the dividing cortical cells of early root nodule primordia (Figure 3A and 3D) and in developing root nodules (Figure 3B and 3E). Both genes were also expressed in root nodules that spontaneously developed in gof-LHK1 plants [10] in the absence of Mesorhizobium loti (Figure 3C and 3F). These results suggest that expression of LjNF-YA1 and LjNF-YB1 in root nodule primordia is regulated by LHK1-mediated networks. The expression patterns were similar to that of GUS expression from the NIN promoter [13]. A construct containing 4 repeats of NBS-yB1a (4xyB1a-GFP-GUS) was also expressed in L. japonicus root nodule primordia, whereas a mutated version of the construct (4xyB1m-GFP-GUS) was not (Figure S7). This suggests that the NBS-yB1a sequence is sufficient for NIN to activate transcription in L. japonicus roots.
The overlapping expression of both NF-Y genes in root nodule primordia suggests that LjNF-YA1 and LjNF-YB1 might function together in root nodule development. We examined whether LjNF-YA1 binds to LjNF-YB1. Since binding of NF-YA to NF-YB depends on dimerization of the latter protein with NF-YC [31], [44], we performed bimolecular fluorescence complementation (BiFC) analyses in N. benthamiana leaves. We expected that the LjNF-Y subunits might form heterotrimeric complexes with the endogenous tobacco NF-YC. eYFP signals were detected in nuclei when eYFPC-LjNF-YA1 (a fusion with the C-terminal half of eYFP) and eYFPN-LjNF-YB1 (a fusion with the N-terminal half of eYFP) were transiently co-expressed in tobacco leaves (Figure 3G). However, interactions were not observed between eYFPC-LjNF-YA1 and eYFPN-LjNF-YB2, containing the LjNF-YB1 homolog, LjNF-YB2 (see Figure S3B) (Figure 3H). On the other hand, eYFPC-LjNF-YA2 (containing the LjNF-YA1 homolog LjNF-YA2; see Figure S3A), interacted with eYFPN-LjNF-YB1 and with eYFPN-LjNF-YB2, but not with eYFPN (Figure 3I–3K). eYFP signals were not detected when eYFPN-LjNF-YB1 and eYFPC were co-expressed (Figure 3L). These results suggest that LjNF-YA1 binds to LjNF-YB1 in planta.
Knockdown of LjNF-YA1 prevents root nodule formation
To investigate loss-of-function phenotypes of LjNF-YA1 and LjNF-YB1, RNAi constructs targeting the two genes were introduced into L. japonicus roots. These constructs were driven by the LjUb promoter. Although several LjNF-YB1 RNAi constructs were tested, the gene was not downregulated (Figure 4E). On the other hand, an RNAi construct containing a sequence specific to the 3′-UTR of LjNF-YA1 specifically prevented accumulation of LjNF-YA1 mRNA (Figure 4E; Figure S3A). Therefore, a phenotypic analysis was performed using roots transformed with this LjNF-YA1 RNAi construct. When inoculated with DsRed-labeled M. loti for 2 weeks, root nodules were produced in 85% of control plants that were transformed with the empty vector (Figure 4B). On the other hand, only 15% of plants with roots that were transformed with the LjNF-YA1 RNAi construct generated infected root nodules (Figure 4A). The efficiency of root nodule formation was significantly reduced in roots transformed with the RNAi construct (Figure S8A). The numbers of small bumps without M. loti invasion were also decreased by the RNAi construct, suggesting that cortical cell division was inhibited at the early stages of the root nodule development. In contrast to root nodule organogenesis, infection threads were formed in 97% of the LjNF-YA1 RNAi plants at efficiencies similar to those in control roots (Figure 4C, 4D; Figure S8A), indicating that epidermal responses resulting in infection thread growth were not influenced by the RNAi construct. Thus it appears that LjNF-YA1 is required for the regulation of cortical cell division in the development of root nodules downstream of NIN. Consistent with this, the RNAi construct also prevented spontaneous nodule formation in gof-CCaMK roots [45] in the absence of M. loti (Figure S8B, S8C).
Fig. 4. Inhibition of root nodule development by the knockdown of LjNF-YA1. 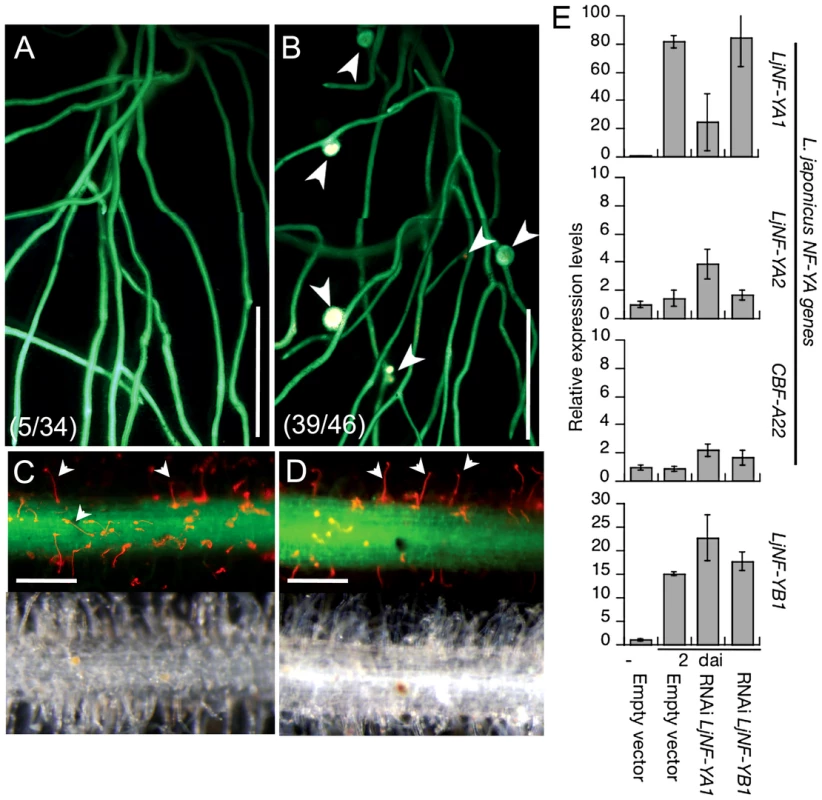
Roots that were transformed with either ProLjUb-RNAi-LjNF-YA1 (A,C) or an empty vector (B,D) were inoculated with DsRed-labeled M. loti for 14 days. (A,B) Root nodule formation in transformed roots. Fluorescence is visible from GFP (the root transformation marker) and from DsRed expressed in M. loti. Arrowheads indicate root nodules. The fractions of plants that formed root nodules on GFP-positive roots are shown in parentheses. (C,D) Infection thread formation in transformed roots. Fluorescence from GFP and DsRed is shown in the upper panels and bright field images are shown in the bottom panels of the same roots. Arrowheads indicate infection threads visualized by DsRed-labeled M. loti. Bars: 5 mm in (A,B), 0.2 mm in (C,D). (E) RT-PCR analyses of gene expression in roots that were transformed with either the empty vector, ProLjUb-RNAi-LjNF-YA1, or ProLjUb-RNAi-LjNF-YB1. Roots were inoculated with (2 dai; days after inoculation) or without (−) M. loti. Expression was analyzed for LjNF-YB1 and three L. japonicus NF-Y subunit A genes (LjNF-YA1, LjNF-YA2, and CBF-A22 [57]). The means and SDs from 3 biological repeats are shown. NIN overexpression results in the formation of root nodule primordium-like structures
It has been shown that NIN is a nodulation-specific factor essential for root nodule organogenesis. The NIN-target gene LjNF-YA1 is also required for root nodule organogenesis. To investigate whether NIN confers cortical cell division in the absence of M. loti, we produced L. japonicus roots that ectopically overexpressed NIN cDNA from the LjUb promoter (ProLjUb-NIN). In uninoculated NIN-overexpressing roots we found lateral roots with enlarged tips and bumps similar to root nodule primordia (Figure 5A, 5C). Such abnormalities were not observed in roots transformed with the empty vector (Figure 5B). In the NIN-overexpressing roots that exhibited the abnormal architecture, 74% (n = 7) of the lateral roots were malformed. The bumps occurred in roots with the abnormal lateral roots, and were often broader along the apical-basal axis than normal root nodules (Figure 5C; Figure S9). When a belt-shaped bump was counted as one, the mean bump number was 3.55±1.63 (n = 11) on roots that generated them. Microscopic analyses revealed that the bumps were formed via cortical cell division that preferentially occurred at positions opposite the protoxylem poles (6/8 bumps) (Figure 5E, 5F). This was also seen in wild-type root nodule formation (11/12 root nodules; Figure 5D). The NIN-induced bumps were anatomically similar to the root nodule primordia. Thus, NIN activity results in cortical cell division. However, unlike the infected wild-type root nodules the NIN-induced bumps did not develop peripheral vascular systems. NIN-induced bumps were also formed on transgenic roots carrying mutations in genes that function upstream of NIN, including CCaMK, LHK1, NSP1, and NSP2 (Figure S10A–S10E). The lower efficiency of bump generation in nsp2 mutants implies that NSP2 contributes to the regulation of cortical cell division independently of NIN.
Fig. 5. NIN overexpression induces cortical cell division. 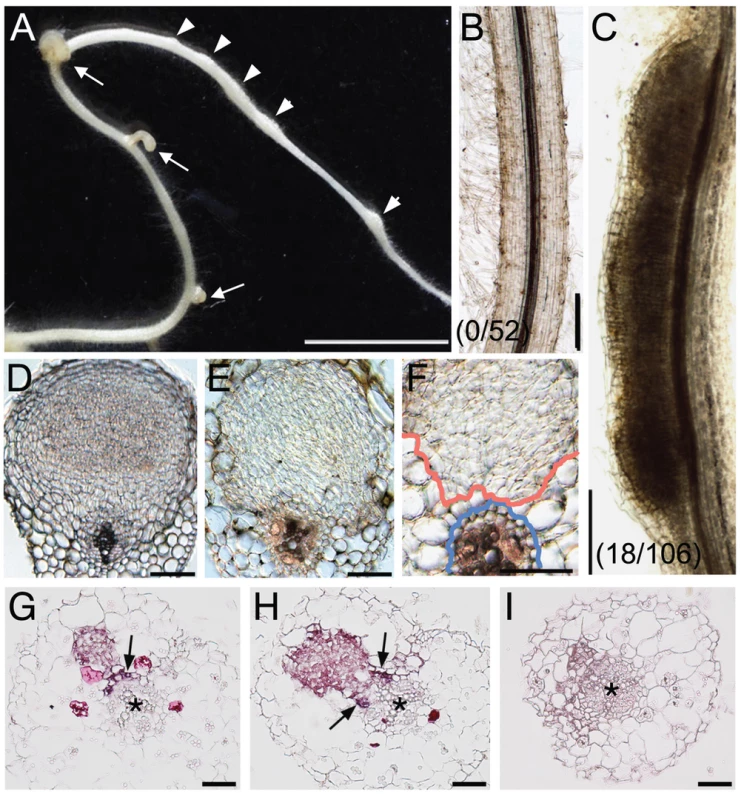
(A) A Gifu (wild-type) root was transformed with ProLjUb-NIN and then cultured for 6 weeks in the absence of M. loti. Bumps (arrowheads) and malformed lateral roots (arrows) are indicated. (B,C) Cleared roots that were transformed with either an empty vector (B) or ProLjUb-NIN (C). The fractions of plants with bumps are shown in parentheses. (D) A transverse section of a root nodule primordium (10 dai) formed on a MG-20 (wild-type) root that was transformed with the empty vector. (E,F) Transverse sections of bumps formed on uninoculated MG-20 roots that were transformed with ProLjUb-NIN. Blue and red lines in (F) represent the outer edges of the endodermis and the boundary of the region with dividing cortical cells, respectively. (G–I) in situ RNA hybridization of ENOD40-1 in transverse sections of bumps caused by NIN overexpression, using either antisense (G,H) or sense probes (I). Asterisks indicate the central xylem. Arrows indicate the pericycle with in situ signals. Bars: 5 mm in (A); 0.2 mm in (B,C); 0.1 mm in (D–F); 50 µm in (G–I). We used in situ RNA hybridization to investigate expression of an early nodulin gene, ENOD40-1, in the NIN-induced bumps. This nodulin gene is often used as a molecular maker for rhizobial infection. High levels of ENOD40-1 expression were detected in dividing cortical cells within the NIN-induced bumps, and in pericycle cells opposite to the xylem pole (Figure 5G, 5H). This expression pattern was similar to that seen in root nodules caused by Nod factors and rhizobial infection [46]–[48], indicating that the NIN-induced bumps possess a root nodule primordium-like identity at the molecular level.
Overexpression of NIN and NF-Y subunit genes enhances cell division
We next ectopically overexpressed the LjNF-YA1 and LjNF-YB1 cDNAs to examine whether the Lotus NF-Y subunits function to regulate cell proliferation. Roots overexpressing LjNF-YA1 produced lateral roots with malformed tips (Figure 6B; Figure S11C, S11F), similar to those formed as a result of NIN overexpression (Figure 5A). Such abnormalities were not observed in roots transformed with either the empty vector or ProLjUb-LjNF-YB1 alone (Figure 6A; Figure S11A, S11B, S11F). However, the co-expression of LjNF-YB1 exaggerated the root architecture abnormalities caused by LjNF-YA1 overexpression (Figure 6C; Figure S11D, S11F). This effect was not observed when LjNF-YB2 was co-expressed with LjNF-YA1 (Figure S11G–S11I), indicating a functional specificity of the interaction between LjNF-YA1 and LjNF-YB1.
Fig. 6. NIN and NF-Y subunits regulate cell division. 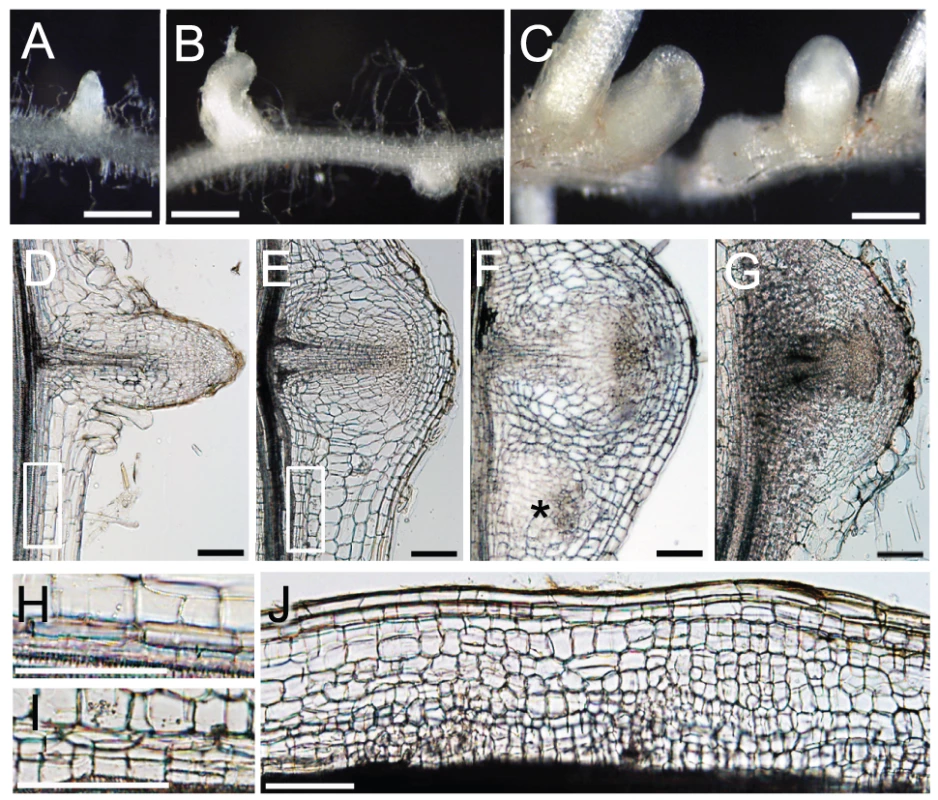
(A–C) Lateral roots formed on MG-20 roots that were transformed with either an empty vector (A), ProLjUb-LjNF-YA1 (B), or ProLjUb-LjNF-YA1 Pro35S-LjNF-YB1 (C). Roots were cultured in the absence of M. loti. (D–G) Longitudinal sections of lateral roots formed on MG-20 roots that were transformed with either the empty vector (D), ProLjUb-LjNF-YA1 Pro35S-LjNF-YB1 (E,F), or ProLjUb-NIN (G). An asterisk in (F) indicates an additional lateral root meristem-like structure. (H,I) Magnified images of the boxed regions in (D) and (E), respectively. (J) A longitudinal section of the root that was transformed with ProLjUb-LjNF-YA1 and Pro35S-LjNF-YB1. Note the presence of small cells that were generated by cortical cell division. Bars: 1 mm in (A–C); 0.1 mm in (D–G,J); 0.2 mm in (H,I). The intervals between lateral roots were shorter in roots that co-overexpressed LjNF-YA1 and LjNF-YB1 (Figure 6C; Figure S11E). Extra cell division was observed in the lateral root primordia and their proximal regions, including the pericycle, which is the origin of the lateral root primordium (Figure 6D, 6E, 6H, 6I). This pattern was also observed in the NIN-overexpressing roots (Figure 6G). In the LjNF-YA1 and LjNF-YB1 co-overexpressing roots, additional lateral root meristem-like structures emerged in the proximal regions of the lateral root primordia (Figure 6F). Although abnormal cell division occurred in the cortex of these roots (Figure 6J), they did not form visible bumps similar to those found on roots overexpressing NIN. These results indicate that NIN and the NF-Y subunits positively influence cell division. The effect of LjNF-YA1 overexpression is consistent with the role of LjNF-YA1 in root nodule primordium development (Figure 4A).
To investigate cell division activities in non-meristematic regions of roots that were transformed with either ProLjUb-NIN or ProLjUb-LjNF-YA1 Pro35S-LjNF-YB1, we produced plant lines that were stably transformed with a cell division marker, ProLjCycB1;1-CycB1;1(NT)-GUS. In this construct, a 4 kb genomic fragment of L. japonicus Cyclin B1;1 (chr1.CM0269.150.r2.m) was translationally fused to the GUS reporter coding sequence. This genomic fragment contains the 2.9 kb promoter region from the putative initiation codon and the coding region that corresponds to the N-terminal part of the protein including the destruction-box. This reporter showed dot-like expression patterns in meristem regions of root tips, lateral root primordia, and developing root nodules (Figure S12A–S12F). We introduced either ProLjUb-NIN or ProLjUb-LjNF-YA1 Pro35S-LjNF-YB1 into roots of ProLjCycB1;1-CycB1;1(NT)-GUS plants. Ectopically localized GUS staining was detected in the cortex, pericycle, and endodermis of these roots, in addition to root tip regions and lateral root primordia (Figure S12G–S12I). The number of GUS staining foci was significantly increased as compared to control roots (Figure S12J). These results indicated that overexprression of NIN and the NF-Y subunit genes resulted in increase in division activity.
We analyzed expression of the early nodulin genes ENOD40-1, ENOD40-2, and ENOD2 in roots overexpressing the NF-Y genes, and compared the results with those of roots transformed with ProLjUb-NIN or the empty vector. The expression of nodulin genes was upregulated in the NIN-overexpressing roots that had altered structures. However, nodulin gene expression levels were not significantly different from the vector control in either roots overexpressing both the NF-Y genes or roots overexpressing NIN but with no visible alterations in structure (Figure 7A, 7B, 7C, 7F). Thus, expression of the nodulin genes was correlated with alterations in root morphology in the NIN-overexpressing roots. The NF-Y genes did not upregulate expression of the nodulin genes, even if aberrant lateral roots were formed. This result indicates that the function of the Lotus NF-Y genes is not associated with root nodule primordium identity. We also found that LjNF-YB1 expression was not upregulated in NIN-overexpressing roots with no visible alterations in root structure (Figure 7D, 7E). This result is consistent with the idea that expression of both the NF-Y genes is important to stimulate cell division.
Fig. 7. RT–PCR analysis of the expression of early nodulin genes and LjNF-Y subunit genes. 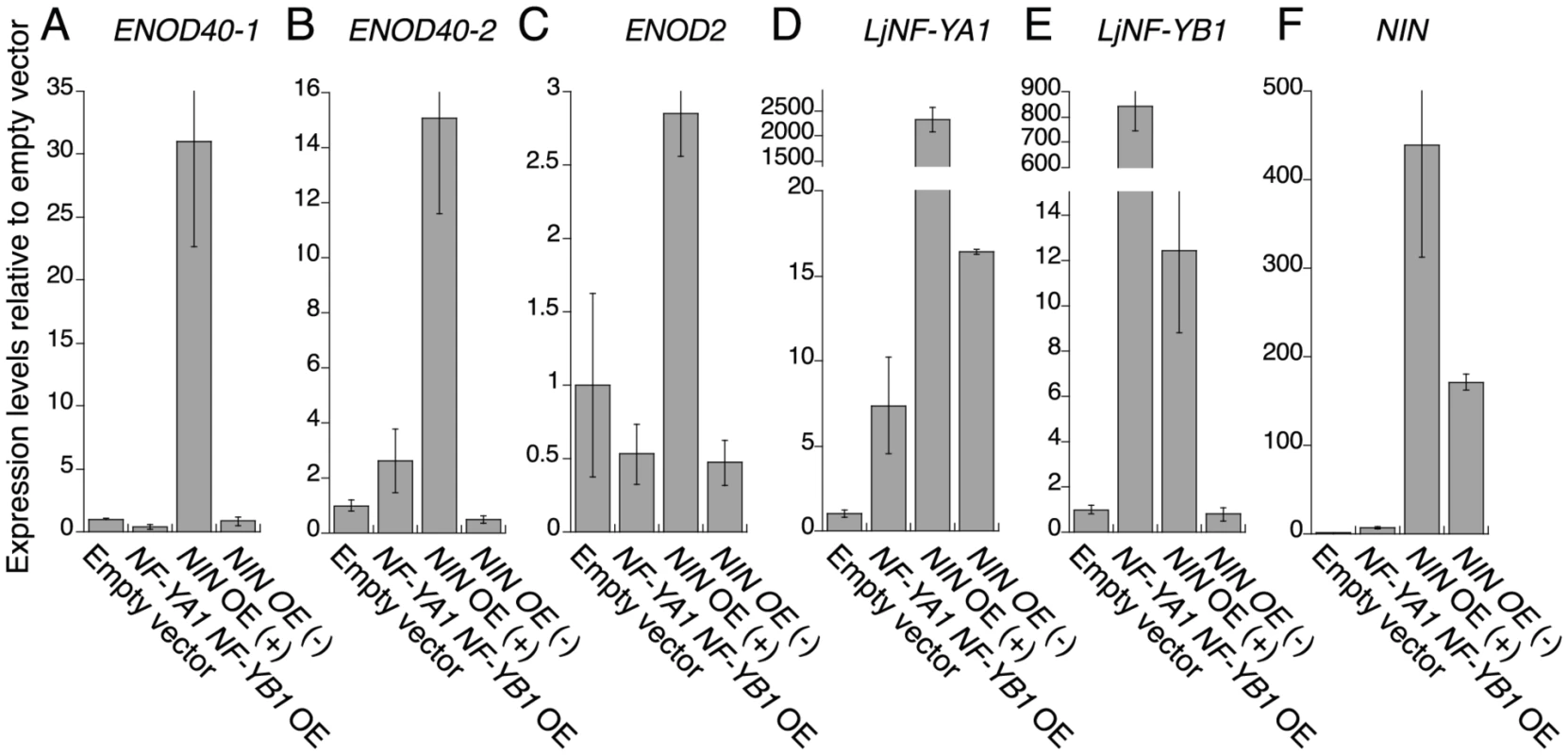
Expression was analyzed for ENOD40-1 (A), ENOD40-2 (B), ENOD2 (C), LjNF-YA1 (D), LjNF-YB1 (E), and NIN (F). Plants were cultured for 3 weeks without M. loti inoculation. Total RNA was isolated from roots transformed with either an empty vector, ProLjUb-LjNF-YA1 plus Pro35S-LjNF-YB1 (NF-YA1 NF-YB1 OE), or ProLjUb-NIN (NIN OE). ProLjUb-NIN roots that exhibited altered structures (+) were harvested separately from roots with no morphological alterations (−). The means and SDs from 3 biological repeats are shown. Discussion
We have characterized the molecular role of NIN as a transcriptional activator that directly targets the NF-Y subunit genes LjNF-YA1 and LjNF-YB1 (Figure 2). The DEX-inducible NIN protein induced expression of the NF-Y genes without M. loti inoculation (Figure 1). We further found that NIN has the ability to induce cortical cell division in the absence of bacterial symbionts (Figure 5). Functional analyses using both RNAi and overexpression indicated that the NF-Y genes play overlapping roles with that of NIN (Figure 4; Figure 6). It appears that the Lotus NF-Y subunits contribute to cortical cell division in a NIN-mediated transcriptional pathway.
NIN is a key player in root nodule organogenesis
NIN overexpression induced bumps in the absence of M. loti that were anatomically similar to root nodule primordia (Figure 5). The overexpression of M. truncatula response regulator9 induces arrested primordium-like structures in which cell division occurs in the pericycle, endodermis, and cortex [49]. Although these arrested primordia are somewhat similar to the bumps caused by NIN overexpression, the former structures are probably lateral root primordia [49]. In L. japonicus and M. truncatula, lateral root primordia originate from the pericycle and are also associated with cell division in the endodermis and cortex [49]. NIN-induced bumps, on the other hand, are initially generated by cortical cell division at a radial position where root nodule primordia are usually formed. Furthermore, ENOD40 is expressed in NIN-induced bumps with patterns characteristics of root nodules (Figure 5). We conclude that the bumps caused by NIN overexpression possess a root nodule primordium-like identity. NIN is a factor that initiates cortical cell division during root nodule organogenesis. The NIN-induced bumps were formed on roots of the symbiotic mutants ccamk-3, hit1, nsp1, and nsp2 (Figure S10), indicating that NIN functions downstream or independently of these symbiotic genes.
Bump formation resulting from NIN overexpression (17% of plants with transformed roots; Figure 5) is less efficient than the spontaneous root nodule formation observed in gof-CCaMK and gof-LHK roots (67 and 40%, respectively) [10]. Peripheral vascular systems were not formed in the NIN-induced bumps. Thus, it is unlikely that NIN is the sole factor regulating the initiation of cortical cell division and root nodule organogenesis. NSP2 is also required for root nodule organogenesis downstream of LHK1 [14]. We found that NSP2 influenced the efficiency of bump formation in NIN-overexpressing roots (Figure S10). Cytokinin signaling is involved in developmental programs that are generally conserved in plant species. It is likely that LHK1 regulates such pathways in addition to those specific to nodulation. Alternatively, NIN itself may prevent the efficient initiation of cortical cell division. NIN acts as a negative regulator in some nodulation processes. For example, M. truncatula NIN negatively influences the spatial pattern of ENOD11 expression [28]. Also in M. truncatula, a loss-of-function NIN mutation represses expression of the CLE genes, which encode small peptides that act in an inhibitory autoregulation pathway for root nodule formation [50]. Deregulated expression of NIN may activate this negative-feedback regulation system, reducing the efficiency of NIN-induced bump formation.
We also found that regions where the NIN-induced bumps emerged were often expanded along the apical-basal axis, whereas cortical cell division occurred at the correct radial position (Figure 5). These findings suggest that the radial position where cortical cell division occurs is defined independently of NIN expression. Regulatory mechanisms, by which NIN expression is restricted to the proper area along the apical-basal axis, are important for generating root nodule meristems of the correct size. The M. truncatula ethylene-insensitive sickle mutants form sequentially connected root nodules [51]. However, it is likely that the radial positioning of NIN-induced bumps is under the control of ethylene, because ethylene also influences radial positioning [52], [53]. NIN expression appears to be spatially regulated downstream of LHK1 activation. NIN expression was restricted to regions within or near dividing cortical cells in roots that were treated with exogenous cytokinin [18]. Feedback mechanisms may negatively influence the division of cortical cells surrounding root nodule primordia through modulation of the LHK1 signaling pathway.
NF-Y subunits function downstream of NIN
We searched for genes whose expression was positively regulated by NIN using the transcriptome database, and found several candidates that might be targeted by NIN. We used ChIP assays and EMSAs to show that LjNF-YA1 and LjNF-YB1 are direct NIN targets (Figure 2), and then identified the NBSs in the promoters of these genes. Previously-identified regulatory elements that control expression of nodulation-related genes [30], [54]–[56] do not contains sequences similar to the NBSs. Therefore, the NBSs may be novel elements associated with nodulation.
The Lotus NF-YA1 gene reported here is orthologous to MtHAP2-1 (Figure S3A), and identical to CBF-A01, which is induced by M. loti inoculation [57]. The knockdown of MtHAP2-1 resulted in an arrest of root nodule growth that was associated with the absence of a clear meristem zone [32]. This zone is usually present at the apical region of a tip-growing indeterminate-type root nodule. MtHAP2-1 expression is restricted to this meristem zone by post-transcriptional regulatory mechanisms [32], [58]. The Lotus counterpart, on the other hand, is required for the regulation of cortical cell division during the early stages of determinate-type root nodule development (Figure 4; Figure S8). We showed that NIN transcriptionally regulates LjNF-YA1 expression. The expression pattern is consistent with that of NIN and with the function of LjNF-YA1 in root nodule organogenesis. We further demonstrated that LjNF-YA1 interacts with LjNF-YB1. The overexpression of LjNF-YB1 exaggerated the effect of LjNF-YA1 overexpression on cell division (Figure 6; Figure S11). Our results, showing expression of both genes in the root nodule primordia and interactions between the proteins in planta (Figure 3), support the idea that the Lotus NF-Y subunits function in the same NF-Y complex during root nodule development. Thus, the two NF-Y proteins participate in the stimulation and/or promotion of cell division.
While LjNF-YA1 overexpression influenced the root architecture, LjNF-YB1 overexpression alone did not affect cell division. Similarly, overexpression of the common bean NF-YC1 did not influence root architecture apart from an increase in root nodule number [38]. Therefore, LjNF-YA1 expression is of primary importance for the control of cell division. A phylogenetic analysis suggests that the NF-YA genes in the clade that includes LjNF-YA1 and MtHAP2-1 have evolved as a result of duplication in the ancestral legume lineage (Figure S3). Interestingly, the expression of NF-YA genes that are related to LjNF-YA1 and MtHAP2-1 is strongly induced in actinorhizal root nodules, which are lateral root-like structures generated by interactions between the non-legume plants Casuarina glauca and Alnus glutinosa and their Frankia symbionts [59]. This implies that NF-Y is important for root nodule organogenesis in actinorhizal plants, and leads to the speculation that different types of root nodule symbiosis system recruited the molecular networks regulating the expression of NF-Y genes.
NF-Y complexes are general transcription factors that target CCAAT boxes. They regulate gene expression by influencing histone modifications [60]–[62]. Transcriptional activators, which work together with the NF-Y complexes, are required for the efficient expression of target genes [60], [63], [64]. The genes encoding such factors may be included among the NIN-target genes. Overexpression of the Lotus NF-Y subunit genes stimulated cell division in the lateral root primordia, resulting in the production of malformed lateral root tips at high frequencies. The ectopically expressed NF-Y subunits apparently interacted with factors other than NIN to stimulate the proliferation of cells with the competence for division in the lateral root primordia. Cortical cells, on the other hand, possess low meristematic activity in the absence of NIN activity. Factors that may act downstream of NIN would be required for the Lotus NF-Y subunits to fully stimulate cortical cell division.
NIN functions in the establishment of the root nodule symbiosis system
Rice, which engages in mycorrhizal but not root nodule symbiosis, has genes corresponding to NSP1, NSP2, and the common SYM genes [65], [66]. These are functionally equivalent to the corresponding genes in L. japonicus, whereas the closest NIN homolog in rice, OsNLP1, does not rescue the nin-2 phenotype [66]. NIN and its orthologous proteins in other legume species possess common structural characteristics distinct from other NIN-like proteins [33], [67], suggesting that NIN has acquired functions specialized for nodulation. NF-Y complexes, on the other hand, are transcription factors that are widespread among eukaryotes. The induction of extra cell division by the overexpression of the NF-Y subunits in lateral root primordia was likely due to the formation of complexes with other subunits that are expressed in dividing cells and/or cells with the competency to divide. This implies that NF-Y may be generally important for regulating cell division in plants, as it is in mammals. Importantly, overexpression of the common bean NF-YC1 upregulated the expression of genes that encode cell cycle regulators [38]. The NF-YC1 gene is ubiquitously expressed in various plant organs. NIN is thought to coordinate gene expression for onset of root nodule organogenesis by regulating the expression of genes encoding NF-Y components in addition to genes that are specifically involved in the regulation of cortical cell division. Unlike the results from overexpression of NIN and the Lotus NF-Y subunit genes, roots expressing gof-CCaMK have not been reported to show abnormally stimulated cell division during lateral root development [10]. This suggests that NIN and the NF-Y subunits are more directly involved in the regulation of cell division than CCaMK. NIN function may be related to the regulation of gene expression networks that are generally required for plant life, particularly those involved in cell division. Arabidopsis RKD proteins also stimulate cell division and possess the RWP-RK domain, suggesting that proteins containing this domain may have a general role in the transcriptional regulation of genes responsible for cell division [68], [69].
The evolution of NIN, which is thought to be specific to legume plants, may have led to the coordinated expression of the subset of genes that are involved in the generation of functionally de novo organs, the root nodules. A number of possible origins, including lateral roots, have been proposed for root nodules [70]. Our results are suggestive of common regulatory mechanisms that regulate cell proliferation during root nodule and lateral root organogenesis. A comprehensive analysis of the gene expression networks that are regulated by NIN would provide further clues about the root nodule-specific pathways that lead to the establishment of the root nodule symbiosis system.
Materials and Methods
Plant materials and bacterial strains
We used the L. japonicus accessions Gifu B-129 and MG-20 as wild-type plants, and the symbiotic mutants nin-2 [19], nsp1 (SL1795-4) [22], nsp2 (sym70) [23], ccamk-3 [13], hit1 [15], and cyclops-3 [45]. These plants were inoculated with M. loti strains MAFF303099 or one that constitutively expresses DsRed [29]. Unless otherwise indicated in a figure legend, Gifu B-129 was used as the wild type. Root transformations with A. rhizogenes and inoculation procedures with M. loti were performed as described previously [71]. An empty vector with the GFP marker for selection was used as the controls in root transformations.
Plasmid construction
Plasmids were constructed using standard molecular biology techniques. The LjNF-YA1 and LjNF-YB1 promoters were amplified by PCR from Gifu B-129 genomic DNA. The amplified fragments were inserted into an entry vector, pENTR1A (Invitrogen), that was digested with BamHI and NotI. The promoters were then transferred into pMDC162-GFP, which was produced by replacing the hygromycin resistance gene of pMDC162 [72] with GFP. PCR-amplified cDNAs of NIN, LjNF-YA1, LjNF-YB1, LjNF-YA2, and LjNF-YB2 were digested using restriction enzyme sites in the linker sequences, and the resulting fragments were cloned into pENTR-1A. To produce the GFP, GAL4DBD, GR, and myc fusions of NIN, each cDNA was inserted into the XhoI site of pENTR1A downstream of the NIN cDNA lacking a stop codon (inserted between the KpnI and NotI sites). For the BiFC analysis, cDNAs for eYFPN173-myc (tagged with the myc epitope from pSPYNE(R)173 [73]) and eYFPC155-HA (tagged with the HA epitope from pSPYCE(MR) [73]) were amplified by PCR. The former fragment was inserted upstream of the LjNF-YB1 and LjNF-YB2 cDNAs in pENTR1A. The latter fragment was inserted upstream of the LjNF-YA1 and LjNF-YA2 cDNAs. For controls, eYFPN173-myc and eYFPC155-HA were cloned into pENTR1A. The NIN, NIN-myc, LjNF-YA1, and LjNF-YB1 cDNAs were transferred from the ENTRY vectors into pUB-GW-GFP [43]; the NIN, NIN-GFP, NIN-GAL4DBD, NIN-myc, GAL4DBD, eYFPN173-LjNF-YB1, eYFPN173-LjNF-YB2, eYFP N173-myc, eYFP C155-LjNF-YA1, eYFP C155-LjNF-YA2, and eYFPC155-HA cDNAs were transferred into pUB-GW-Hyg [43]; and the NIN-GFP, NIN-GR, and NIN-myc cDNAs were transferred into ProNIN-DC-NINter [66]. To co-express LjNF-YA1 and LjNF-YB1 in L. japonicus roots, the LjNF-YA1 cDNA was transferred into pUB-GW-LjNF-YB1, which was produced by replacing the GFP gene in pUB-GW-GFP with the LjNF-YB1 cDNA. For the knockdown analyses of LjNF-YA1 and LjNF-YB1, the 3′-UTR of each gene was amplified by PCR, and cloned into pENTR/D-TOPO with reverse direction. The fragments were transferred into pUB-GWS-GFP [43]. For transcriptional activation analyses, 4xUAS with the CaMV35S minimal promoter was amplified by PCR from pTA7001 [74], and cloned into pENTR1A. 4xyB1a and 4xyB1m were synthesized by PCR and inserted between the SalI and BstXI sites of pENTR1A, with the CaMV35S minimal promoter between the NotI and XhoI sites. These synthetic promoters were transferred into pKGWFS7 [75]. For in vitro translation, cDNAs corresponding to NIN-myc and NIN(520–878)-myc were digested with SgfI and PmeI, and cloned into the pF3K-WG (BYDV) Flexi Vector (Promega). For ProLjCycB1;1-CycB1;1(NT)-GUS, the approximately 4 kb LjCycB1;1 genomic fragment was amplified from Gifu B-129 genomic DNA. The genomic fragment was subcloned into pCR-Blunt (Invitrogen), and the fragment between the HindIII and SmaI sites was then inserted upstream of the GUS gene in the binary vector pBI101.3. The sequences of primers used for plasmid construction are shown in Table S1.
GUS staining assays
Roots were washed with 100 mM NaPO4 buffer (pH 7.0), and incubated in GUS staining solution [100 mM NaPO4 (pH 7.0), 10 mM EDTA, 0.5 mg/ml 5-bromo-4-chloro-3-indolyl-b-glucuronic acid, 2 mM K4Fe(CN)6, 2 mM K3Fe(CN)6, and 0.1% Triton X-100] for 2 to 4 h at 37°C after vacuum infiltration for 10 min. To examine the transcriptional activity of NIN with the synthetic 4xyB1a and 4xyB1m promoter constructs, discs were cut out from N. benthamiana leaves 3 days after Agrobacterium infiltration, incubated in the GUS staining solution for 8 h, and then decolorized with 75% ethanol.
Microscopic analysis
Observations of roots inoculated with DsRed-labeled M. loti were performed as described previously [10]. For the BiFC and transcriptional activation analyses, N. benthamiana leaves were observed 3 days after Agrobacterium infiltration. Confocal microscopy was performed using a μRadiance confocal microscope (BioRad). GFP and YFP florescence was induced using a 488 nm argon laser and imaged using an HQ530/60 filter. Autofluorescence from chloroplasts was induced using a 514 nm green HeNe laser and imaged using an E600LP emission filter.
Expression analyses
In situ hybridizations with digoxigenin-labeled RNA probes was performed as described previously [47]. For RT-PCR, total RNAs were isolated from L. japonicus roots using the Plant RNeasy Mini kit (Qiagen). First strand cDNAs were synthesized using the QuantiTect Reverse Transcription kit (Qiagen). RT-PCR was performed in a LightCycler with the LightCycler FastStart DNA Master SYBR Green I reaction mix (Roche Applied Science). Expression levels were normalized using polyubiquitin transcripts. Primers for polyubiquitin, NIN, LjENOD40-1, LjENOD40-2, and LjENOD2 were synthesized as described previously [19], [48], [76]. Sequences of the other primers used for RT-PCR are shown in Table S1.
Electrophoresis mobility shift assays (EMSAs)
The NIN-myc and NIN(520–878)-myc proteins were synthesized using the TNT SP6 High-Yield Wheat Germ Protein Expression System (Promega). The in vitro translation products produced without templates were used as the control. Purified probes (10–30 fmol) were end-labeled with γ32P-ATP and incubated with the in vitro translation products (1.5–3 µl) in 20 µl of EMSA DNA binding buffer [20 mM Hepes-KOH (pH 7.9), 50 mM KCl, 1 mM MgCl2, 1 mM DTT, 4% glycerol, 0.1% Triton X-100] supplemented with 2 µg BSA and 400 ng dIdC, for 30 min at room temperature. Reactants were separated in 5% polyacrylamide gels with 0.5× TBE buffer at 150 V for 90 min. Radioactivity was visualized with an imaging analyzer (BAS2500; Fujifilm). The oligonucleotides that were used as probes and as primers for probe synthesis are shown in Table S1.
ChIP assays
ChIP was carried out as described previously [77] with minor modifications. Roots were transformed with either ProLjUb-NIN-myc or the empty vector then fixed with 1% formaldehyde in MC buffer for 10 min under a vacuum. The reaction was stopped by adding 0.125 M glycine, and the roots were washed three times with MC buffer. The fixed tissue (0.7–1.0 g) was powdered with a mortar and pestle in liquid nitrogen, suspended with 15 ml of M1 buffer supplemented with 1 mM PMSF and Complete Protease Inhibitor Cocktail (Roche Diagnostics), and incubated for 30 min on ice. The crude extract was filtered through two layers of Miracloth and washed with 15 ml of M1 buffer. The filtrate was centrifuged at 1,600×g for 15 min at 4°C. The pellet was washed 4 times, each with 1 ml of M2 buffer, and once with M3 buffer. After centrifugation at 2,000×g for 10 min, the pellet was resuspended in 1 ml of Sonication buffer that was supplemented with 1 mM PMSF and Protease Inhibitor Cocktail, incubated for 20 min on ice, and sonicated 9 times for 10 seconds using a Branson Sonifier 250 at 40% duty cycle, power setting 4. The chromatin suspension was centrifuged 2 times at 14,000×g for 15 min at 4°C. An equal volume of IP buffer was added to the supernatant, and 50 µl were removed to use as the as Input. Then 2 µg of anti-myc polyclonal antibodies (Santa Cruz Biotechnology, Inc) were added to the remaining suspension and the mixture was incubated for 3 h at 4°C. After centrifugation, 20 µl of proteinA-agarose (25% slurry; Santa Cruz Biotechnology, Inc) that had been pre-incubated with 0.5 mg/ml sheared salmon sperm DNA was added to the supernatant, and this mixture was rotated for 1 h at 4°C. After washing 4 times with IP buffer, the immunoprecipitate was eluted twice with 100 µl of elution buffer, then 150 µl of 1 M Tris buffer (pH 9.0) was added into the combined eluates. The DNA reverse cross-linking procedure was performed as described previously [77]. After ethanol precipitation, the DNA was dissolved in 50 µl of 5 mM Tris-HCl (pH 8.0). RT-PCR was performed to determine whether the target promoter regions had been enriched. The PCR products were quantified by comparison with products amplified using primers specific to the 5S rRNA gene. The sequences of primers used to amplify promoter fragments from LjNF-YA1, LjNF-YB1, the rRNA gene, chr4.CM0179.190.r2.m, and chr6.CM1757.140.r2.m are shown in Table S1.
Accession numbers
The sequences of genes listed in Figure S2A can be found at http://www.kazusa.or.jp/lotus/.
Supporting Information
Zdroje
1. DownieJA, WalkerSA (1999) Plant responses to nodulation factors. Curr Opin Plant Biol 2 : 483–489.
2. MadsenEB, MadsenLH, RadutoiuS, OlbrytM, RakwalskaM, et al. (2003) A receptor kinase gene of the LysM type is involved in legume perception of rhizobial signals. Nature 425 : 637–640.
3. RadutoiuS, MadsenLH, MadsenEB, FelleHH, UmeharaY, et al. (2003) Plant recognition of symbiotic bacteria requires two LysM receptor-like kinases. Nature 425 : 585–592.
4. ArrighiJF, BarreA, Ben AmorB, BersoultA, SorianoLC, et al. (2006) The Medicago truncatula lysin motif-receptor-like kinase gene family includes NFP and new nodule-expressed genes. Plant Physiol 142 : 265–279.
5. RadutoiuS, MadsenLH, MadsenEB, JurkiewiczA, FukaiE, et al. (2007) LysM domains mediate lipochitin–oligosaccharide recognition and Nfr genes extend the symbiotic host range. EMBO J 26 : 3923–3935.
6. SmitP, LimpensE, GeurtsR, FedorovaE, DolgikhF, et al. (2007) Medicago LYK3, an entry receptor in rhizobial nodulation factor signaling. Plant Physiol 145 : 183–191.
7. EhrhardtDW, WaisR, LongSR (1996) Calcium spiking in plant root hairs responding to Rhizobium nodulation signals. Cell 85 : 673–681.
8. OldroydGED, DownieJA (2008) Coordinating nodule morphogenesis with rhizobial infection in legumes. Annu Rev Plant Biol 59 : 519–546.
9. KistnerC, ParniskeM (2002) Evolution of signal transduction in intracellular symbiosis. Trends Plant Sci 7 : 511–518.
10. HayashiT, BanbaM, ShimodaY, KouchiH, HayashiM, et al. (2010) A dominant function of CCaMK in intracellular accommodation of bacterial and fungal endosymbionts. Plant J 63 : 141–154.
11. MadsenLH, TirichineL, JurkiewiczA, SullivanJT, HeckmannAB, et al. (2010) The molecular network governing nodule organogenesis and infection in the model legume Lotus japonicus. Nature Commun 1 : 10.
12. GleasonC, ChaudhuriS, YangT, MuñozA, PoovaiahBW, et al. (2006) Nodulation independent of rhizobia induced by a calcium-activated kinase lacking autoinhibition. Nature 441 : 1149–1152.
13. TirichineL, Imaizumi-AnrakuH, YoshidaS, MurakamiY, MadsenLH, et al. (2006) Deregulation of a Ca2+ calmodulin-dependent kinase leads to spontaneous nodule development. Nature 441 : 1153–1156.
14. TirichineL, SandalN, MadsenLH, RadutoiuS, AlbrektsenAS, et al. (2007) A gain-of-function mutation in a cytokinin receptor triggers spontaneous root nodule organogenesis. Science 315 : 104–107.
15. MurrayJD, KarasBJ, SatoS, TabataS, AmyotL, et al. (2007) A cytokinin perception mutant colonized by Rhizobium in the absence of nodule organogenesis. Science 315 : 101–104.
16. HwangI, SheenJ, MullerB (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63 : 353–80.
17. PletJ, WassonA, ArielF, Le SignorC, BakerD, et al. (2011) MtCRE1-dependent cytokinin signaling integrates bacterial and plant cues to coordinate symbiotic nodule organogenesis in Medicago truncatula. Plant J 65 : 622–633.
18. HeckmannAB, SandalN, BekAS, MadsenLH, JurkiewiczA, et al. (2011) Cytokinin induction of root nodule primordia in Lotus japonicus is regulated by a mechanism operating in the root cortex. Mol Plant-Microbe Interact 24 : 1385–1395.
19. SchauserL, RoussisA, StillerJ, StougaardJ (1999) A plant regulator controlling development of symbiotic root nodules. Nature 402 : 191–195.
20. CatoiraR, GaleraC, de BillyF, PenmetsaRV, JournetEP, et al. (2000) Four genes of Medicago truncatula controlling components of a Nod factor transduction pathway. Plant Cell 12 : 1647–1665.
21. OldroydGED, LongSR (2003) Identification and characterization of Nodulation-Signaling Pathway 2, a Gene of Medicago truncatula involved in Nod factor signaling. Plant Physiol 131 : 1027–1032.
22. HeckmannAB, LombardoF, MiwaH, PerryJA, BunnewellS, et al. (2006) Lotus japonicus nodulation requires two GRAS domain regulators, one of which is functionally conserved in a non-legume. Plant Physiol 142 : 1739–1750.
23. MurakamiY, MiwaH, Imaizumi-AnrakuH, KouchiH, DownieJA, et al. (2006) Positional cloning identifies Lotus japonicus NSP2, a putative transcription factor of the GRAS family, required for NIN and ENOD40 gene expression in nodule initiation. DNA Res 13 : 255–265.
24. MailletF, PoinsotV, AndréO, Puech-PagésV, HaouyA, et al. (2011) Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza. Nature 469 : 58–63.
25. LiuW, KohlenW, LilloA, Op den CampR, IvanovS, et al. (2011) Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 23 : 3853–3865.
26. KalóP, GleasonC, EdwardsA, MarshJ, MitraRM, et al. (2005) Nodulation signaling in legumes requires NSP2, a member of the GRAS family of transcriptional regulators. Science 308 : 1786–1789.
27. SmitP, RaedtsJ, PortyankoV, DebelléF, GoughC, et al. (2005) NSP1 of the GRAS protein family is essential for Rhizobial Nod factor–induced transcription. Science 308 : 1789–1791.
28. MarshJF, RakocevicA, MitraRM, BrocardL, SunJ, et al. (2007) Medicago truncatula NIN is essential for rhizobial-independent nodule organogenesis induced by autoactive calcium/calmodulin-dependent protein kinase. Plant Physiol 144 : 324–335.
29. MaekawaT, Maekawa-YoshikawaM, TakedaN, Imaizumi-AnrakuH, MurookaY, et al. (2009) Gibberellin controls the nodulation signaling pathway in Lotus japonicus. Plant J 58 : 183–194.
30. HirschS, KimJ, MuñozA, HeckmannAB, DownieJA, et al. (2009) GRAS proteins form a DNA binding complex to induce gene expression during nodulation signaling in Medicago truncatula. Plant Cell 21 : 545–557.
31. MantovaniR (1999) The molecular biology of the CCAAT-binding factor NF-Y. Gene 239 : 15–27.
32. CombierJP, FrugierF, de BillyF, BoualemA, El-YahyaouiF, et al. (2006) MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev 20 : 3084–3088.
33. SchauserL, WielochW, StougaardJ (2005) Evolution of NIN-like proteins in Arabidopsis, Rice, and Lotus japonicus. J Mol Evol 60 : 229–237.
34. XieF, MurrayJD, KimJ, HeckmannAB, EdwardsA, et al. (2012) Legume pectate lyase required for root infection by rhizobia. Proc Natl Acad Sci USA 109 : 633–638.
35. CastaingsL, CamargoA, PocholleD, GaudonV, TexierY, et al. (2008) The nodule inception-like protein 7 modulates nitrate sensing and metabolism in Arabidopsis. Plant J 57 : 426–435.
36. HaseloffJ (1999) GFP variants for multispectral imaging in living cells. Methods Cell Biol 58 : 139–151.
37. HøgslundN, RadutoiuS, KrusellL, VoroshilovaV, HannahMA, et al. (2009) Dissection of symbiosis and organ development by integrated transcriptome analysis of Lotus japonicus mutant and wild-type plants. PLoS ONE 4: e6556 doi:10.1371/journal.pone.0006556
38. ZanettiME, BlancoFA, BekerMP, BattagliaM, AguilarOM (2010) A C subunit of the plant nuclear factor NF-Y required for rhizobial infection and nodule development affects partner selection in the common bean–Rhizobium etli Symbiosis. Plant Cell 22 : 4142–4157.
39. ElkonR, LinhartC, SharanR, ShamirR, ShilohY (2003) Genome-wide in silico identification of transcriptional regulators controlling the cell cycle in human cells. Genome Res 13 : 773–780.
40. HuQ, LuJF, LuoR, SenS, MaitySN (2006) Inhibition of CBF/NF-Y mediated transcription activation arrests cells at G2/M phase and suppresses expression of genes activated at G2/M phase of the cell cycle. Nucleic Acids Res 34 : 6272–6285.
41. GrskovicM, ChaivorapolC, Gaspar-MaiaA, LiH, Ramalho-SantosM (2007) Systematic identification of cis-regulatory sequences active in mouse and human embryonic stem cells. PLoS Genet 3: e145 doi:10.1371/journal.pgen.0030145
42. BenattiP, DolfiniD, ViganòA, RavoM, WeiszA, et al. (2011) Specific inhibition of NF-Y subunits triggers different cell proliferation defects. Nucleic Acids Res 39 : 5356–5368.
43. MaekawaT, KusakabeM, ShimodaY, SatoS, TabataS, et al. (2008) Polyubiquitin promoter-based binary vectors for overexpression and gene silencing in Lotus japonicus. Mol Plant-Microbe Interact 21 : 375–382.
44. ThirumuruganT, ItoY, KuboT, SerizawaA, KurataN (2008) Identification, characterization and interaction of HAP family genes in rice. Mol Genet Genomics 279 : 279–289.
45. YanoK, YoshidaS, MüllerJ, SinghS, BanbaM, et al. (2008) CYCLOPS, a mediator of symbiotic intracellular accommodation. Proc Natl Acad Sci USA 105 : 20540–20545.
46. MinamiE, KouchiH, CohnJR, OgawaT, StaceyG (1996) Expression of the early nodulin, ENOD40, in soybean roots in response to various lipo-chitin signal molecules. Plant J 10 : 23–32.
47. NiwaS, KawaguchiM, Imaizumi-AnrakuH, ChechetkaSA, IshizakaM, et al. (2001) Responses of a model legume Lotus japonicus to lipochitin oligosaccharide nodulation factors purified from Mesorhizobium loti JRL501. Mol Plant-Microbe Interact 14 : 848–856.
48. TakedaN, OkamotoS, HayashiM, MurookaY (2005) Expression of LjENOD40 genes in response to symbiotic and non-symbiotic signals: LjENOD40–1 and LjENOD40–2 are differentially regulated in Lotus japonicus. Plant Cell Physiol 46 : 1291–1298.
49. Op den CampRH, De MitaS, LilloA, CaoQ, LimpensE, et al. (2011) A phylogenetic strategy based on a legume-specific whole genome duplication yields symbiotic cytokinin type-A response regulators. Plant Physiol 157 : 2013–2022.
50. MortierV, Den HerderG, WhitfordR, Van de VeldeW, RombautsS, et al. (2010) CLE peptides control Medicago truncatula nodulation locally and systemically. Plant Physiol 153 : 222–237.
51. PenmetsaRV, CookDR (1997) A legume ethylene-insensitive mutant hyperinfected by its rhizobial symbiont. Science 275 : 527–530.
52. HeidstraR, YangWC, YalcinY, PeckS, EmonsAM, et al. (1997) Ethylene provides positional information on cortical cell division but is not involved in Nod factor-induced root hair tip growth in Rhizobium-legume interaction. Development 124 : 1781–1787.
53. PenmetsaRV, FrugoliJA, SmithLS, LongSR, CookDR (2003) Dual genetic pathways controlling nodule number in Medicago truncatula. Plant Physiol 131 : 998–1008.
54. LaursenNB, LarsenK, KnudsenJY, HoffmannHJ, PoulsenC, et al. (1994) A protein binding AT-rich sequence in the soybean leghemoglobin c3 promoter is a general cis element that requires proximal DNA elements to stimulate transcription. Plant Cell 6 : 659–668.
55. FehlbergV, ViewegMF, DohmannEN, HohnjecN, PühlerA, et al. (2005) The promoter of the leghaemoglobin gene VfLb29: functional analysis and identification of modules necessary for its activation in the infected cells of root nodules and in the arbuscule-containing cells of mycorrhizal roots. J Exp Bot 56 : 799–806.
56. AndriankajaA, Boisson-DernierA, FrancesL, SauviacL, JauneauA, et al. (2007) AP2-ERF transcription factors mediate Nod factor–dependent Mt ENOD11 activation in root hairs via a novel cis-regulatory motif. Plant Cell 19 : 2866–2885.
57. AsamizuE, ShimodaY, KouchiH, TabataS, SatoS (2008) A positive regulatory role for LjERF1 in the nodulation process is revealed by systematic analysis of nodule-associated transcription factors of Lotus japonicus. Plant Physiol 147 : 2030–2040.
58. CombierJP, de BillyF, GamasP, NiebelA, RivasS (2008) Trans-regulation of the expression of the transcription factor MtHAP2-1 by a uORF controls root nodule development. Genes Dev 22 : 1549–1559.
59. HocherV, AlloisioN, AuguyF, FournierP, DoumasP, et al. (2011) Transcriptomics of actinorhizal symbioses reveals homologs of the whole common symbiotic signaling cascade. Plant physiol 156 : 700–711.
60. de SilvioA, ImbrianoC, MantovaniR (1999) Dissection of the NF-Y transcriptional activation potential. Nucleic Acids Res 27 : 2578–2584.
61. DonatiG, GattaR, DolfiniD, FossatiA, CeribelliM, et al. (2008) An NF-Y-dependent switch of positive and negative histone methyl marks on CCAAT promoters. PLoS ONE 3: e2066 doi:10.1371/journal.pone.0002066
62. GurtnerA, FuschiP, MagiF, ColussiC, GaetanoC, et al. (2008) NF-Y dependent epigenetic modifications discriminate between proliferating and postmitotic tissue. PLoS ONE 3: e2047 doi:10.1371/journal.pone.0002047
63. YamamotoA, KagayaY, ToyoshimaR, KagayaM, TakedaS, et al. (2009) Arabidopsis NF-YB subunits LEC1 and LEC1-LIKE activate transcription by interacting with seed-specific ABRE-binding factors. Plant J 58 : 843–856.
64. LiuJX, HowellSH (2010) bZIP28 and NF-Y transcription factors are activated by ER stress and assemble into a transcriptional complex to regulate stress response genes in Arabidopsis. Plant Cell 22 : 782–796.
65. BanbaM, GutjahrC, MiyaoA, HirochikaH, PaszkowskiU, et al. (2008) Divergence of evolutionary ways among common sym genes: CASTOR and CCaMK show functional conservation signaling pathway. Plant Cell Physiol 49 : 1659–1671.
66. YokotaK, SoyanoT, KouchiH, HayashiM (2010) Function of GRAS proteins in root nodule symbiosis is retained in homologs of a non-legume, rice. Plant Cell Physiol 51 : 1436–1442.
67. YokotaK, HayashiM (2011) Function and evolution of nodulation genes in legumes. Cell Mol. Life Sci 68 : 1341–1351.
68. KöszegiD, JohnstonAJ, RuttenT, CzihalA, AltschmiedL, et al. (2011) Members of the RKD transcription factor family induce an egg cell-like gene expression program. Plant J 67 : 280–291.
69. WakiT, HikiT, WatanabeR, HashimotoH, NakajimaK (2011) The Arabidopsis RWP-RK protein RKD4 triggers gene expression and pattern formation in early embryogenesis. Curr Biol 21 : 1277–1281.
70. HirschAM, LarueTA, DoyleJ (1997) Is the legume nodule a modified root or stem or an organ sui generis? Crit Rev Plant Sci 16 : 361–392.
71. Díaz CL, Grønlund M, Schlaman HRM, Spaink HP (2005) Induction of hairy roots for symbiotic gene expression studies. In: Marquez AJ. Editor. Lotus japonicus Handbook. Dordrecht: Springer. pp. 261–277.
72. CurtisMD, GrossniklausU (2003) A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 133 : 462–469.
73. WaadtR, SchmidtLK, LohseM, HashimotoK, BockR, et al. (2008) Multicolor bimolecular fluorescence complementation reveals simultaneous formation of alternative CBL/CIPK complexes in planta. Plant J 56 : 505–516.
74. AoyamaT, ChuaNH (1997) A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J 11 : 605–612.
75. KarimiM, InzéD, DepickerA (2002) GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7 : 193–195.
76. FlemetakisE, KavroulakisN, QuaedvliegNE, SpainkHP, DimouM, et al. (2000) Lotus japonicus contains two distinct ENOD40 genes that are expressed in symbiotic, nonsymbiotic, and embryonic tissues. Mol Plant–Microbe Interact 13 : 987–994.
77. WangH, TangW, ZhuC, PerrySE (2002) A chromatin immunoprecipitation (ChIP) approach to isolate genes regulated by AGL15, a MADS domain protein that preferentially accumulates in embryos. Plant J 32 : 831–843.
78. LohmannGV, ShimodaY, NielsenMW, JørgensenFG, GrossmannC, et al. (2010) Evolution and regulation of the Lotus japonicus LysM receptor gene family. Mol Plant-Microbe Interact 23 : 510–521.
79. RzhetskyA, NeiM (1992) A simple method for estimating and testing minimum evolution trees. Mol Biol Evol 9 : 945–967.
80. TamuraK, PetersonD, PetersonN, StecherG, NeiM, et al. (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28 : 2731–2739.
81. Colón-CarmonaA, YouR, Haimovitch-GalT, DoernerP (1999) Spatio-temporal analysis of mitotic activity with a labile cyclin-GUS fusion protein. Plant J 20 : 503–508.
82. KouchiH, SekineM, HataS (1995) Distinct classes of mitotic cyclins are differentially expressed in the soybean shoot apex during the cell cycle. Plant Cell 7 : 1143–1155.
Štítky
Genetika Reprodukční medicína
Článek Ubiquitous Polygenicity of Human Complex Traits: Genome-Wide Analysis of 49 Traits in KoreansČlánek Alternative Splicing and Subfunctionalization Generates Functional Diversity in Fungal ProteomesČlánek RFX Transcription Factor DAF-19 Regulates 5-HT and Innate Immune Responses to Pathogenic Bacteria inČlánek Surveillance-Activated Defenses Block the ROS–Induced Mitochondrial Unfolded Protein ResponseČlánek Deficiency Reduces Adipose OXPHOS Capacity and Triggers Inflammation and Insulin Resistance in Mice
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 3- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- Power and Predictive Accuracy of Polygenic Risk Scores
- Rare Copy Number Variants Are a Common Cause of Short Stature
- Coordination of Flower Maturation by a Regulatory Circuit of Three MicroRNAs
- Ubiquitous Polygenicity of Human Complex Traits: Genome-Wide Analysis of 49 Traits in Koreans
- Genomic Evidence for Island Population Conversion Resolves Conflicting Theories of Polar Bear Evolution
- Mechanistic Insight into the Pathology of Polyalanine Expansion Disorders Revealed by a Mouse Model for X Linked Hypopituitarism
- Genome-Wide Association Study and Gene Expression Analysis Identifies as a Predictor of Response to Etanercept Therapy in Rheumatoid Arthritis
- Problem Solved: An Interview with Sir Edwin Southern
- Long Interspersed Element–1 (LINE-1): Passenger or Driver in Human Neoplasms?
- Mouse HFM1/Mer3 Is Required for Crossover Formation and Complete Synapsis of Homologous Chromosomes during Meiosis
- Alternative Splicing and Subfunctionalization Generates Functional Diversity in Fungal Proteomes
- A WRKY Transcription Factor Recruits the SYG1-Like Protein SHB1 to Activate Gene Expression and Seed Cavity Enlargement
- Microhomology-Mediated Mechanisms Underlie Non-Recurrent Disease-Causing Microdeletions of the Gene or Its Regulatory Domain
- Ancient Evolutionary Trade-Offs between Yeast Ploidy States
- Differential Evolutionary Fate of an Ancestral Primate Endogenous Retrovirus Envelope Gene, the EnvV , Captured for a Function in Placentation
- A Feed-Forward Loop Coupling Extracellular BMP Transport and Morphogenesis in Wing
- The Tomato Yellow Leaf Curl Virus Resistance Genes and Are Allelic and Code for DFDGD-Class RNA–Dependent RNA Polymerases
- The U-Box E3 Ubiquitin Ligase TUD1 Functions with a Heterotrimeric G α Subunit to Regulate Brassinosteroid-Mediated Growth in Rice
- Role of the DSC1 Channel in Regulating Neuronal Excitability in : Extending Nervous System Stability under Stress
- –Independent Phenotypic Switching in and a Dual Role for Wor1 in Regulating Switching and Filamentation
- Pax6 Regulates Gene Expression in the Vertebrate Lens through miR-204
- Blood-Informative Transcripts Define Nine Common Axes of Peripheral Blood Gene Expression
- Genetic Architecture of Skin and Eye Color in an African-European Admixed Population
- Fine Characterisation of a Recombination Hotspot at the Locus and Resolution of the Paradoxical Excess of Duplications over Deletions in the General Population
- Estrogen Mediated-Activation of miR-191/425 Cluster Modulates Tumorigenicity of Breast Cancer Cells Depending on Estrogen Receptor Status
- Complex Patterns of Genomic Admixture within Southern Africa
- Yap- and Cdc42-Dependent Nephrogenesis and Morphogenesis during Mouse Kidney Development
- Molecular Networks of Human Muscle Adaptation to Exercise and Age
- Alp/Enigma Family Proteins Cooperate in Z-Disc Formation and Myofibril Assembly
- Polycomb Group Gene Regulates Rice () Seed Development and Grain Filling via a Mechanism Distinct from
- RFX Transcription Factor DAF-19 Regulates 5-HT and Innate Immune Responses to Pathogenic Bacteria in
- Distinct Molecular Strategies for Hox-Mediated Limb Suppression in : From Cooperativity to Dispensability/Antagonism in TALE Partnership
- A Natural Polymorphism in rDNA Replication Origins Links Origin Activation with Calorie Restriction and Lifespan
- TDP2–Dependent Non-Homologous End-Joining Protects against Topoisomerase II–Induced DNA Breaks and Genome Instability in Cells and
- Recurrent Rearrangement during Adaptive Evolution in an Interspecific Yeast Hybrid Suggests a Model for Rapid Introgression
- Genome-Wide Association Study in Mutation Carriers Identifies Novel Loci Associated with Breast and Ovarian Cancer Risk
- Coincident Resection at Both Ends of Random, γ–Induced Double-Strand Breaks Requires MRX (MRN), Sae2 (Ctp1), and Mre11-Nuclease
- Identification of a -Specific Modifier Locus at 6p24 Related to Breast Cancer Risk
- A Novel Function for the Hox Gene in the Male Accessory Gland Regulates the Long-Term Female Post-Mating Response in
- Tdp2: A Means to Fixing the Ends
- A Novel Role for the RNA–Binding Protein FXR1P in Myoblasts Cell-Cycle Progression by Modulating mRNA Stability
- Association Mapping and the Genomic Consequences of Selection in Sunflower
- Histone Deacetylase 2 (HDAC2) Regulates Chromosome Segregation and Kinetochore Function via H4K16 Deacetylation during Oocyte Maturation in Mouse
- A Novel Mutation in the Upstream Open Reading Frame of the Gene Causes a MEN4 Phenotype
- Ataxin1L Is a Regulator of HSC Function Highlighting the Utility of Cross-Tissue Comparisons for Gene Discovery
- Human Spermatogenic Failure Purges Deleterious Mutation Load from the Autosomes and Both Sex Chromosomes, including the Gene
- A Conserved Upstream Motif Orchestrates Autonomous, Germline-Enriched Expression of piRNAs
- Statistical Analysis Reveals Co-Expression Patterns of Many Pairs of Genes in Yeast Are Jointly Regulated by Interacting Loci
- Matefin/SUN-1 Phosphorylation Is Part of a Surveillance Mechanism to Coordinate Chromosome Synapsis and Recombination with Meiotic Progression and Chromosome Movement
- A Role for the Malignant Brain Tumour (MBT) Domain Protein LIN-61 in DNA Double-Strand Break Repair by Homologous Recombination
- The Population and Evolutionary Dynamics of Phage and Bacteria with CRISPR–Mediated Immunity
- Long Noncoding RNA MALAT1 Controls Cell Cycle Progression by Regulating the Expression of Oncogenic Transcription Factor B-MYB
- Surveillance-Activated Defenses Block the ROS–Induced Mitochondrial Unfolded Protein Response
- DNA Topoisomerase III Localizes to Centromeres and Affects Centromeric CENP-A Levels in Fission Yeast
- Genome-Wide Control of RNA Polymerase II Activity by Cohesin
- Divergent Selection Drives Genetic Differentiation in an R2R3-MYB Transcription Factor That Contributes to Incipient Speciation in
- NODULE INCEPTION Directly Targets Subunit Genes to Regulate Essential Processes of Root Nodule Development in
- Spreading of a Prion Domain from Cell-to-Cell by Vesicular Transport in
- Deficiency in Origin Licensing Proteins Impairs Cilia Formation: Implications for the Aetiology of Meier-Gorlin Syndrome
- Deficiency Reduces Adipose OXPHOS Capacity and Triggers Inflammation and Insulin Resistance in Mice
- The Conserved SKN-1/Nrf2 Stress Response Pathway Regulates Synaptic Function in
- Functional Genomic Analysis of the Regulatory Network in
- Astakine 2—the Dark Knight Linking Melatonin to Circadian Regulation in Crustaceans
- CRL2 E3-Ligase Regulates Proliferation and Progression through Meiosis in the Germline
- Both the Caspase CSP-1 and a Caspase-Independent Pathway Promote Programmed Cell Death in Parallel to the Canonical Pathway for Apoptosis in
- PRMT4 Is a Novel Coactivator of c-Myb-Dependent Transcription in Haematopoietic Cell Lines
- A Copy Number Variant at the Locus Likely Confers Risk for Canine Squamous Cell Carcinoma of the Digit
- Evidence of Gene–Environment Interactions between Common Breast Cancer Susceptibility Loci and Established Environmental Risk Factors
- HIV Infection Disrupts the Sympatric Host–Pathogen Relationship in Human Tuberculosis
- Trans-Ethnic Fine-Mapping of Lipid Loci Identifies Population-Specific Signals and Allelic Heterogeneity That Increases the Trait Variance Explained
- A Gene Transfer Agent and a Dynamic Repertoire of Secretion Systems Hold the Keys to the Explosive Radiation of the Emerging Pathogen
- The Role of ATM in the Deficiency in Nonhomologous End-Joining near Telomeres in a Human Cancer Cell Line
- Dynamic Circadian Protein–Protein Interaction Networks Predict Temporal Organization of Cellular Functions
- Nuclear Myosin 1c Facilitates the Chromatin Modifications Required to Activate rRNA Gene Transcription and Cell Cycle Progression
- Robust Prediction of Expression Differences among Human Individuals Using Only Genotype Information
- A Single Cohesin Complex Performs Mitotic and Meiotic Functions in the Protist
- The Role of the Arabidopsis Exosome in siRNA–Independent Silencing of Heterochromatic Loci
- Elevated Expression of the Integrin-Associated Protein PINCH Suppresses the Defects of Muscle Hypercontraction Mutants
- Twist1 Controls a Cell-Specification Switch Governing Cell Fate Decisions within the Cardiac Neural Crest
- Genome-Wide Testing of Putative Functional Exonic Variants in Relationship with Breast and Prostate Cancer Risk in a Multiethnic Population
- Heteroduplex DNA Position Defines the Roles of the Sgs1, Srs2, and Mph1 Helicases in Promoting Distinct Recombination Outcomes
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Fine Characterisation of a Recombination Hotspot at the Locus and Resolution of the Paradoxical Excess of Duplications over Deletions in the General Population
- Molecular Networks of Human Muscle Adaptation to Exercise and Age
- Recurrent Rearrangement during Adaptive Evolution in an Interspecific Yeast Hybrid Suggests a Model for Rapid Introgression
- Genome-Wide Association Study and Gene Expression Analysis Identifies as a Predictor of Response to Etanercept Therapy in Rheumatoid Arthritis
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



