-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Hsp70-Hsp40 Chaperone Complex Functions in Controlling Polarized Growth by Repressing Hsf1-Driven Heat Stress-Associated Transcription
How the molecular mechanisms of stress response are integrated at the cellular level remains obscure. Here we show that the cellular polarity machinery in the fission yeast Schizosaccharomyces pombe undergoes dynamic adaptation to thermal stress resulting in a period of decreased Cdc42 activity and altered, monopolar growth. Cells where the heat stress-associated transcription was genetically upregulated exhibit similar growth patterning in the absence of temperature insults. We identify the Ssa2-Mas5/Hsp70-Hsp40 chaperone complex as repressor of the heat shock transcription factor Hsf1. Cells lacking this chaperone activity constitutively activate the heat-stress-associated transcriptional program. Interestingly, they also exhibit intermittent monopolar growth within a physiological temperature range and are unable to adapt to heat stress. We propose that by negatively regulating the heat stress-associated transcription, the Ssa2-Mas5 chaperone system could optimize cellular growth under different temperature regiments.
Published in the journal: . PLoS Genet 9(10): e32767. doi:10.1371/journal.pgen.1003886
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003886Summary
How the molecular mechanisms of stress response are integrated at the cellular level remains obscure. Here we show that the cellular polarity machinery in the fission yeast Schizosaccharomyces pombe undergoes dynamic adaptation to thermal stress resulting in a period of decreased Cdc42 activity and altered, monopolar growth. Cells where the heat stress-associated transcription was genetically upregulated exhibit similar growth patterning in the absence of temperature insults. We identify the Ssa2-Mas5/Hsp70-Hsp40 chaperone complex as repressor of the heat shock transcription factor Hsf1. Cells lacking this chaperone activity constitutively activate the heat-stress-associated transcriptional program. Interestingly, they also exhibit intermittent monopolar growth within a physiological temperature range and are unable to adapt to heat stress. We propose that by negatively regulating the heat stress-associated transcription, the Ssa2-Mas5 chaperone system could optimize cellular growth under different temperature regiments.
Introduction
Sensing ecological parameters and mounting an appropriate adaptive response allows organisms to thrive in changing habitats. Free-living microorganisms, such as yeast, can maintain regular patterns of growth over a relatively wide range of environmental temperatures. Cellular adaptation to increased temperature is one of the most robust and evolutionarily conserved mechanisms (reviewed in [1]). This response involves temperature-induced activation of genes encoding so-called heat shock proteins (HSPs) [1]–[4]. Molecular chaperones constitute a predominant class of HSPs and their up-regulation is essential for maintaining protein homeostasis at elevated temperatures (reviewed in [1]). Although much of heat shock-associated transcription is transient, recent analyses indicate deviation from this dynamic pattern and existence of genes with expression constitutively regulated by temperature [1], [5], [6].
One of the early events in the heat stress pathway ultimately setting off an extensive downstream gene expression program involves activation of critical transcriptional regulators, the heat shock factors (HSFs) [7], [8]. HSFs are evolutionary conserved winged helix-turn-helix proteins that bind to cis-acting DNA promoter sequences called heat shock elements (HSEs) [9]. HSFs are essential for viability in many fungi and control important developmental processes in higher eukaryotes suggesting that they may regulate basal transcription in addition to their function in stress response [10]–[14]. The ability of HSFs to respond to cellular stresses is under negative regulation by chaperones, modulation of nucleocytoplasmic shuttling, post-translational modifications and, in higher eukaryotes, transition from monomeric to trimeric state (reviewed in [15]). HSF repression by the Hsp90 and the Hsp70s-Hsp40 chaperone complexes [16]–[18] could provide a negative feedback loop titrating production of chaperones to allow for optimal protein folding [19]. Although essential for orchestrating an acute response to changing environment, constitutive Hsf1 transcriptional induction is known to retard cellular growth [20]. However, molecular mechanisms reducing HSF activity following the original heat-induced surge are poorly understood. Although fascinating, the cellular biology of adaptation to growth at elevated temperatures is largely unchartered.
The fission yeast Schizosaccharomyces pombe (S. pombe) are rod-shaped cells that grow at cell tips and divide by placing a medial septum. Upon division, each daughter cell initiates growth at the cell tip inherited from the mother, the so-called old-end (Fig. 1A). During the G2 phase of the cell cycle, cells transition to bipolar growth in a process termed New-End-Take-Off (NETO) [21]. NETO occurs when growth sets a sufficient distance between the cell tips [21], [22] and requires the function of a large protein complex called polarisome and active actin cytoskeleton remodeling [23]–[25]. The polarisome, built around the kelch-repeat protein Tea1 and its binding partner Tea4, ensures the fidelity of polarized growth through recruitment of downstream polarity factors (Fig. 1B and [23], [24], [26], [27]).
Fig. 1. Heat-stress induces transient longitudinal growth arrest succeeded by monopolar growth. 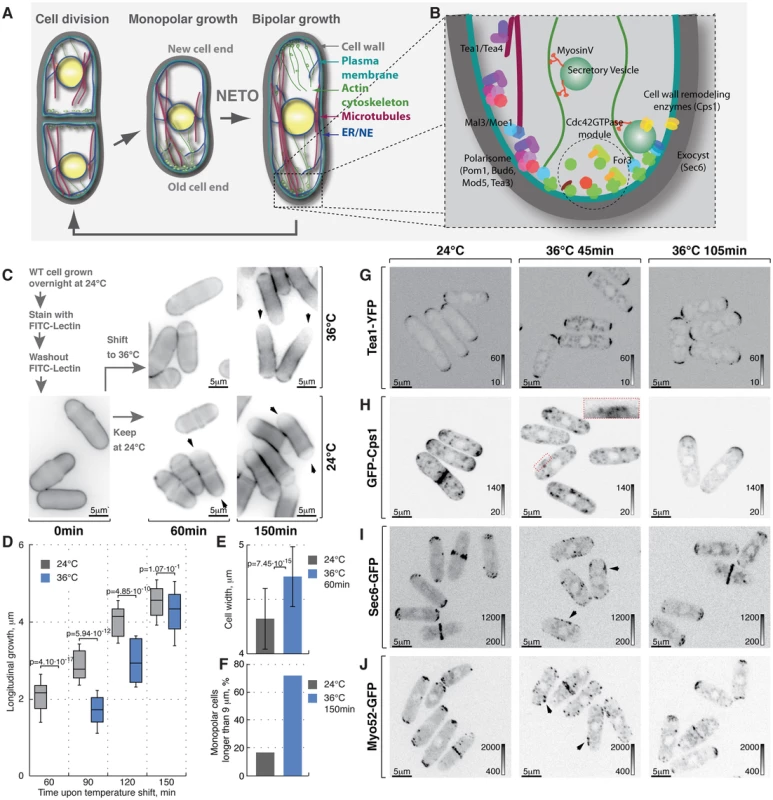
(A) Schematic of fission yeast cell cycle and (B) key regulators of cell growth and polarity. (C) Shown are single z-plane epifluorescence micrographs of FITC-Lectin stained cells grown at indicated temperatures and imaged immediately (left panel), 60 min (middle panels) and 150 min upon dye washout (right panels). (D) Quantification of longitudinal cell growth of cells represented in (C). Presented are box plots with standard deviation for cells grown at 24°C (gray) or shifted to 36°C (blue) for indicated time points; n>25 per sample per time point, p-values derived by Welch's test. (E) Cell width in cells grown at 24°C (gray) or shifted to 36°C for 60 min (blue). (F) Percentage of monopolar late G2 phase cells in a cell suspensions treated as described in (C), n>250 per sample. (G–J) Shown are whole cell maximum intensity z-projections of confocal micrographs of log-phase wild type cells expressing indicated fluorophore tagged marker proteins grown at 24°C (left panels) or shifted to 36°C for 45 min (middle panels) or 105 min (right panels). The arrows point out localization of the marker proteins to lateral cell cortex. Inset represents a zoom-in of the indicated region. Image contrast is reported using corresponding gray wedges. Scale bars, 5 µm. Polarized growth requires local plasma membrane (PM) and cell wall remodeling. The targeted secretion of cell wall enzymes, such as the β-glucan synthase Cps1 [28] relies on long-range exocytic vesicle transport along actin cables and the tethering of the secretory vesicles to the PM by the exocyst complex [29]. Polarized assembly of both actin cables and the exocyst depends on the Rho family GTPase Cdc42 [30]–[34].
The guanine-nucleotide-exchange-factors (GEFs) promote Cdc42 activity while the GTPase-activating proteins (GAPs) repress it [35], [36]. The fission yeast GEFs, Scd1 and Gef1, are enriched at the cell tips [37], [38], while the Cdc42 GAP Rga4 localizes to the lateral cortex [39]. This results in predominant activation of Cdc42 at the cell poles. Recent high-end imaging techniques have uncovered oscillations in Cdc42 activity at the PM suggesting that spatiotemporal activation of Cdc42 is governed by a series of positive and negative feedback loops [40]–[43]. The polarisome and the Cdc42 circuitries appear to intersect at various levels [26], [39], [44].
The mechanisms regulating polarized growth exhibit a high degree of plasticity [45], [46]. Yeast cells readily orient growth towards their mating partners [47], [48] and relocalize the growth machinery to the site of cell injury [49]. In Saccharomyces cerevisiae (S. cerevisiae), heat stress activates the cell wall integrity pathway and promotes nucleotide exchange of the Rho-family GTPase Rho1, which in turn initiates F-actin dependent redistribution of β-glucan synthase throughout the entire cellular cortex [50], [51]. Repolarization of actin as cells adapt to elevated temperature appears to depend on a negative feedback mechanism where the protein kinase C Pkc1 activates MAPK cascade to inhibit the Rho1-GEF [52]. In fission yeast several types of stress, including heat and osmolytes, also cause F-actin redistribution throughout the cellular cortex [53]–[56] and lead to activation of MAP kinases [57]–[59].
Here we utilize a multi-disciplinary approach to explore how the heat stress and associated transcription impact polarized growth in fission yeast. We identify the Mas5-Ssa2 chaperone complex as a negative regulator of the heat shock transcription factor Hsf1. Using cells lacking the chaperone function we probe the physiological outcome of constitutively upregulated heat stress response.
Results
Heat stress triggers transient loss of polarity followed by a period of monopolar growth in fission yeast cells exposed to elevated temperature
Lectins bind polysaccharides of the yeast cell wall with high affinity but without impairing cell growth [60]. Thus, sites of cell wall deposition subsequent to fluorophore-coupled lectin staining can be readily visualized as a dip in fluorescence intensity [61]. To monitor cell growth upon heat stress we stained log-phase wild type cells growing at 24°C with FITC-Lectin, washed-out the excess dye and immediately shifted cells to 36°C or allowed them to grow at 24°C (Fig. 1C). Samples were collected at 30-minute intervals, fixed and subjected to epifluorescence microscopy. Cells continuously growing at 24°C elongated at a rate of 2.51±0.77 mm per hour after the dye washout. Conversely, cells shifted to 36°C for one hour fully arrested longitudinal growth (Fig. 1C and 1D, n>60 cells per sample, pWelch's t-test = 4.10⋅10−17). Interestingly the diameter of heat-stressed cells increased (Fig. 1C and 1E, width increased by 9±7%, n>60 cells per sample, pWelch's t-test = 7.45⋅10−15) and the overall FITC-Lectin staining of the cell visibly decreased (data not shown) suggesting that the cell wall underwent remodeling throughout the cell periphery rather than at the cell tips.
After a period of depolarized cell wall deposition, cells resumed polarized growth and subsequently elongated at a rate of ∼28% greater than cells grown at 24°C (Fig. 1D). Most control cells in the G2 phase of the cell cycle exhibited bipolar growth pattern (Fig. 1C and 1F). In line with previous observations [21], the majority of heat-stressed cells elongated only at one cell tip (Fig. 1C and 1F). When propagated overnight at 36°C, most wild type cells became bipolar (83%; Fig. S1A). Thus, the heat stress leads to a transient arrest of longitudinal growth followed by regaining cellular polarity through an intermediate monopolar growth phase.
Temperature had no effect on the distribution of the polarisome components Tea1-YFP, Tea4-GFP and Pom1-GFP as these markers exhibited bipolar localization in late G2 phase cells at both 24°C and 36°C within the observed timeframe (Fig. 1G, S1B, S1C, see Fig. S1F for quantifications). In contrast, β-glucan synthase GFP-Cps1, which predominantly localized to both cell tips in late G2 cells at 24°C, could be observed in intracellular vesicle-like structures and at the lateral cell cortex 45 minutes after cells were shifted to 36°C (Fig. 1H). Longer incubation at 36°C (105 minutes) allowed all cells to repolarize GFP-Cps1 to the cell tips, albeit in a monopolar fashion (Fig. 1H and S1F). Similarly, the exocyst component Sec6-GFP localized to both cell tips in unstressed late G2 phase cells but was observed throughout the cell cortex in cells shifted to 36°C for 45 minutes (Fig. 1I). Cells growing at 36°C for 105 minutes did repolarize Sec6-GFP but approximately half of late G2 phase cells confined it to only one tip (Fig. 1I and S1F).
Polarization of actin structures typical for yeast cells grown at normal temperatures was completely abolished following incubation at 36°C for 45 minutes. Cells were able to re-polarize F-actin cytoskeleton after 105 minutes at 36°C but it was unequally distributed between the two tips of most late G2 phase cells (Fig. S1D, S1F and S1G). Similarly, both fluorophore-tagged formin For3 and the type-V myosin Myo52 relocalized from the tips to the entire cortex upon heat stress but then re-established polarity after 105 minutes at 36°C, although in a monopolar fashion (Fig. 1J, S1E and S1F).
Taken together our results suggest that processes confining the growth machinery to the polarisome-demarcated cell tips are rapidly perturbed by temperature increase. Cells become monopolar in the adaptive phase following the heat stress but eventually resume normal bipolar growth.
Cortical association of GTP-bound Cdc42 and its GEFs and GAP is modulated by temperature
Both F-actin distribution and exocyst localization are governed by the activity of a Rho-family GTPase Cdc42 [29]. To monitor the active Cdc42 in vivo during heat-stress we used the GFP-tagged CRIB (Cdc42/Rac-interactive binding) domain reported to interact with the GTP-bound Cdc42 [39]. G2 phase cells grown at 24°C focus the Cdc42 activity to the cell tips with moderate oscillations in its levels ([40] and Fig. 2A, top panel). Minutes after cells were shifted from 24°C to 36°C, CRIB-GFP was seen spreading from the cell tips towards the cell equator establishing zones of Cdc42-activity at the cell sides (Fig. 2A, middle panel). After a period of depolarization, cells restricted CRIB-GFP to just one tip and initiated monopolar growth (Fig. 2A, bottom panel). In contrast to the bipolar localization in almost all of G2 phase cells grown at 24°C, a third of G2 phase cells grown at 36°C for 105 minutes exhibited a strictly monopolar CRIB-GFP pattern (Fig. 2B and S2A). While total fluorescence signal of CRIB-GFP was not perturbed by temperature, its cortical association was decreased as compared to unstressed cells (Fig. 2C).
Fig. 2. Cortical association of GTP-bound Cdc42 and its GEFs and GAP is modulated by temperature. 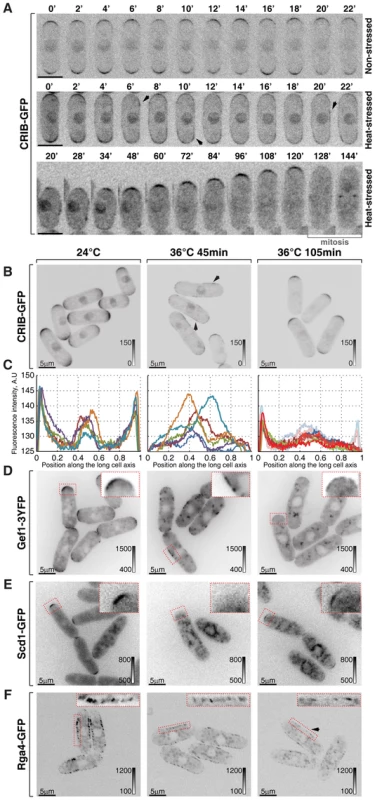
(A) Single z-plane spinning disk confocal microscopy time-lapse images of CRIB-GFP expressing wild type cells grown at room temperature (top panel) or shifted to 36°C for the indicated period of time (middle and bottom panels). Spreading of CRIB-GFP signal immediately following heat stress (middle panel) is indicated by arrowheads. A heat-stressed cell shown in the bottom panel repolarizes CRIB-GFP to one cell tip and eventually enters mitosis. (B) Shown are whole cell maximum intensity z-projections of scanning confocal micrographs of log-phase wild type cells expressing CRIB-GFP grown at 24°C (left panels) or shifted to 36°C for 45 min (middle panels) or 105 min (right panels). The arrows point out localization of the marker protein to lateral cell cortex. (C) Quantification of CRIB-GFP intensities along the long cell axis in log-phase wild type cells grown under indicated temperature regiments. Individual lines correspond to individual cells. Low opacity lines refer to heat-stressed cells from the subpopulation exhibiting bipolar CRIB-GFP localization after 105 min at 36°C. (D–F) Shown are single z-plane micrographs of log-phase wild type cells expressing indicated fluorophore-tagged Cdc42 regulators grown at 24°C (left panels) or shifted to 36°C for 45 min (middle panels) or 105 min (right panels). The insets represent zoomed-in and re-contrasted main image segments outlined in red dotted line. Image contrast is reported using corresponding gray wedges. Scale bars, 5 µm. Both Cdc42-GEFs exhibited a bipolar localization pattern in unstressed late G2 cells (Fig. 2D, 2E and S2A). Forty-five minutes upon temperature shift from 24°C to 36°C, Gef1-3YFP localized to patches distributed throughout the plasma membrane whereas Scd1-GFP was no longer detectable at the cell cortex (Fig. 2D and 2E). Both Gef1-3YFP and Scd1-GFP also localized to intracellular structures upon heat stress. After 105 minutes at 36°C, both GEFs became confined to cell tips but in a monopolar fashion (Fig. 2D, 2E and S2A). Interestingly, Gef1-3YFP and Scd1-GFP levels were severely diminished at the cortex of cells repolarizing growth at elevated temperature. The average intensity of Scd1-GFP at a single cell tip decreased by 49±23% (pWelch's t-test = 1.43⋅10−7, n>25 cells per sample), whereas Gef1-3YFP signal dipped by 65±17% (pWelch's t-test = 7.34⋅10−12, n>25 cells per sample).
The only known fission yeast Cdc42-GAP, Rga4, localizes to the lateral cell cortex and to the new cell end in pre-NETO cells [39]. Upon heat stress, the Rga4-GFP localization remained restricted to the cell sides throughout the course of the experiment although its absolute amount at the cortex progressively decreased (Fig. 2F; after 105 minutes at 36°C the cortex-associated Rga4-GFP signal fell by 34±22%, pWelch's t-test = 7.35⋅10−8, n>25 cells per sample). The diminished cortical levels of Cdc42 regulators in cells shifted to the elevated temperature were not a consequence of a gross decrease in total amounts of these proteins (Fig. S2B).
Since heat-stressed fission yeast cells eventually resume bipolar growth (Fig. S1A), we analyzed cortical levels of Cdc42-GAP and GEFs in cells growing overnight at 36°C. Gef1-3YFP and Scd1-GFP did assume bipolar localization in late G2 cells (Fig. S2E–F). Surprisingly, the average signal intensity per cell tip remained relatively low, as did Rga4-GFP levels at the lateral cortex. In fact, even though levels of active Cdc42 at the cortex were similar in cells growing at 18°C, 24°C and 30°C (Fig. S2C and S2D), the cortical levels of the three Cdc42 regulators inversely correlated with the growth temperature (Fig. S2E–G). Immunoblot analyses of cells grown to log-phase at 18°C, 24°C, 30°C and 36°C showed a similar trend in total protein levels of Scd1-GFP, Gef1-3YFP and Rga4-GFP (Fig. S2H).
Thus, cortical Cdc42 activity appears to be extensively regulated at both initial and adaptive phases of the heat-stress response and during steady state growth at different temperatures.
Overexpression of the heat shock transcription factor Hsf1 leads to monopolar growth
Cells respond to stress by adjusting their transcriptional output [1], [62]. Heat-stress associated transcription is primarily mediated by a temporary activation of the HSF family of transcription factors (reviewed in [63]), Hsf1 is the only HSF homologue present in the fission yeast genome and it was reported to be essential not only for heat shock response but also for vegetative growth [11], [64]. We confirmed that cells depleted of Hsf1 ceased growth (Fig. S3A). We wondered if cell polarity could be affected by the Hsf1 transcriptional activity alone, without the temperature change. To this aim, we replaced the fission yeast hsf1 genomic promoter with a strong, thiamine responsive nmt1 promoter. Cells with high levels of Hsf1 eventually arrested growth (Fig. S3B). Overexpression of Hsf1 was sufficient to promote its transcriptional activity since we observed vastly increased levels of the heat stress-induced protein Hsp104 (Fig. S3C–D) and the GFP reporter expressed under hsp104 regulatory elements (Fig. 3A and S3E, see Materials and Methods for details on reporter construction). Strikingly, Calcofluor staining revealed that Hsf1-overexpressing cells became increasingly monopolar (Fig. 3B and 3B; on average the new-end growth decreased by 59±22% in nmt1:hsf1 cells as compared to wild type cells, pWelch's t-test = 4.58⋅10−10, n>50 cells per sample). Furthermore, cells overexpressing Hsf1 exhibited decreased tip association of CRIB-GFP that predominantly localized to only one cell tip (Fig. 3D and 3E). Taken together, it appears that high Hsf1 activity prevents normal growth patterning and eventually leads to growth arrest in fission yeast.
Fig. 3. Hsf1-mediated transcription regulates growth patterning in fission yeast. 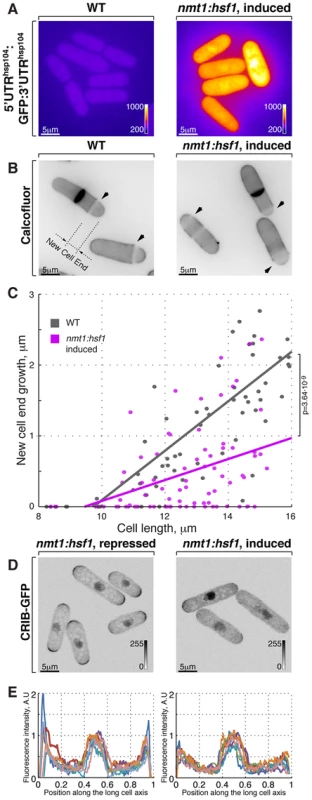
(A) GFP reporter expression is driven by hsp104 promoter in cells where Hsf1 ORF is under the control of either wild type or nmt1 promoter grown to log phase in minimal medium without thiamine. Shown are pseudocolored average intensity whole cell z-projections of epifluorescence image stacks. Image contrast is reported using corresponding color wedges. (B) Single z-plane images of calcofluor stained wild type and cells with Hsf1 over-expression driven by nmt1 promoter induced for 20 hours by removal of thiamine. Arrowheads point to birth-scars; the new-cell-end length was determined as a distance from the birth-scar to the proximal cell tip. (C) New-cell-end length as a function of cell length for wild type (gray) and cells over-expressing Hsf1 from the thiamine responsive nmt1 promoter (magenta). Dots represent individual late G2 phase cells and lines represent linear regressions for cells analyzed (n>30). p-values were obtained through ANCOVA analysis. (D) Shown are whole cell maximum intensity z-projections of scanning confocal micrographs of nmt1::hsf1 cells expressing CRIB-GFP and grown in the presence (left panel) or absence (right panel) of thiamine. (E) Quantification of CRIB-GFP intensities along the long cell axis in log-phase cells grown under conditions described in (D). Gray wedges report image contrasting. Scale bars, 5 µm. The Ssa2-Mas5 chaperone complex modulates adaptation to heat stress in fission yeast
Hsp90 and Hsp70-Hsp40 chaperones are known repressors of Hsf1 activity but both Hsp90 and Hsp70 proteins are known to engage a large number of client proteins [65]. To isolate specificity factor(s) for Hsp70 that could function in this process, we screened the deletion library of sixteen non-essential genes encoding nucleocytoplasmic Hsp40 chaperones (Supplementary Table S2). Two screening strategies were employed. One approach was based on our finding that Hsf1 overexpression leads to decreased growth rates and eventually a growth arrest. We hypothesized that mutant strains with elevated levels of Hsf1-associated transcription would display decreased growth rates already at 24°C and would not be able to grow at 36°C due to high Hsf1 activity. Within the deletion library of nonessential Hsp70-binding proteins only the strain lacking the DnaJ chaperone Mas5 exhibited such growth pattern (Fig. 4A).
Fig. 4. Screening for chaperone mutant cells with elevated levels of heat stress-associated transcription. 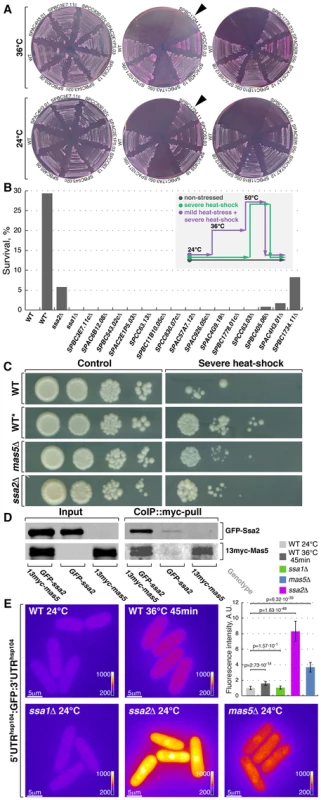
(A) Fission yeast strains lacking indicated non-essential nucleocytoplasmic Hsp40 proteins were streaked out onto YES plates containing phloxine B and grown at 24°C or 36°C. Arrowheads points to the strain lacking DnaJ chaperone Mas5. (B) Survival rate upon extreme heat-shock (50°C for 15 min) of wild type cells (WT), wild type cells pre-exposed to mild heat stress (WT*) and cells lacking individual SSA subfamily chaperones or indicated non-essential nucleocytoplasmic Hsp40 proteins. The inset outlines the screening strategy used to identify strains with elevated levels of heat-stress associated transcription: cells were grown to log-phase at 24°C and aliquots were either allowed to continue growth at 24°C (gray line) or shifted to 50°C for 15 min (green line). An additional sample of wild type cells was shifted to 36°C for 45 min before being transferred to 50°C for 15 min (purple line). Number of colony forming units was measured for each sample and used to calculate survival rates. (C) Dilution spotting assay of wild type cells (WT), wild type cells pre-exposed to mild heat-stress (WT*) and cells lacking Mas5 or Ssa2 grown at 24°C (left panels) and exposed to 50°C for 15 min prior to plating (right panels). (D) Lysates prepared from cells with indicated genotypes were incubated with anti-myc antibodies and subsequently with beads coupled to Protein-G. Proteins that remained associated with the beads after multiple buffer washes were resolved by SDS-PAGE and prepared for Western Blotting with anti-myc and anti-GFP antibodies. (E) Pseudocolored average z-projection epifluorescence images of the GFP under the regulation of hsp104 regulatory elements in wild type, ssa1Δ, ssa2Δ and mas5Δ cells grown in indicated conditions. Image contrast is reported using corresponding color wedges. Scale bars, 5 µm. Top right panel histogram quantifies the fluorescence signal in indicated strains. The second screening strategy exploited the fact that a mild heat stress increases survival of cells exposed to a subsequent severe transient heat shock, likely due to higher expression of protective factors in pre-stressed cells [66]. As outlined in Fig. 4B, wild type cells and deletion mutants were grown to log-phase at 24°C and culture aliquots were either allowed to continue growth at 24°C or shifted to 50°C for 15 minutes before returning them to 24°C. Additionally, a sample of wild type cells was shifted to 36°C for 45 minutes before being exposed to 50°C. Survival was measured by assessing the number of colony forming units in each culture (Fig. 4B) and confirmed by a dilution spotting assay (Fig. 4C). Wild type and most tested mutant strains exhibited a survival rate below 0.5% when shifted from 24°C to 50°C for 15 minutes (n>200 CFU per strain). Conversely, 29% of wild type cells survived if a mild heat stress preceded the severe heat shock. Interestingly, cells lacking Mas5 showed an 8% survival rate when shifted directly from 24°C to 50°C for 15 minutes (Fig. 4B and 4C).
Both major nucleocytoplasmic Hsp70 chaperones, Ssa1 and Ssa2, physically interacted with Mas5 as shown by co-immunoprecipitation experiments (Fig. 4D and S4A). Cells lacking Ssa2 but not Ssa1 exhibited increased survival following a transient heat shock, similar to mas5Δ cells (Fig. 4B and 4C). In line with this, the fluorophore-tagged marker proteins Hsp104, Swo1 and Psi1 that are known to be strongly upregulated upon heat stress were highly abundant in mas5Δ and ssa2Δ but not in ssa1Δ cells, again suggesting that Ssa2-Mas5 complex may have a specific function in heat stress response (Fig. S4B and data not shown). Excluding the possibility that elevated fluorescence levels were due to selective stabilization of heat-responsive proteins in mutant cells, the UTRsHsp104:GFP reporter was also strongly elevated in both ssa2Δ and mas5Δ but not in ssa1Δ cells (Fig. 4E). Thus, mas5Δ and ssa2Δ cells express high levels of heat stress markers already at 24°C, which may increase their chances of survival following transient heat shock.
Cells lacking the Mas5-Ssa2 chaperone complex exhibit an intermittent, monopolar growth pattern
Doubling time of log-phase mas5Δ and ssa2Δ cultures growing at 24°C increased approximately 1.3-fold and 2-fold as compared to the wild type (Fig. S5A left panel, pWelch's t-test = 1.61⋅10−2 and 7.16⋅10−3, n = 3). Moreover, cells lacking Mas5 and Ssa2 divided at a consistently shorter length as compared to the wild type (Fig. S5A, right panel). Calcofluor staining suggested that both ssa2Δ and mas5Δ cells exhibited significantly decreased new-cell-end growth (Fig. 5A and 5B, on average, mas5Δ and ssa2Δ cells reduced the new-end growth by 50±24% and 78±93% respectively, pWelch's t-test = 2.28⋅10−28 and 2.46⋅10−11, n>60 per genotype).
Fig. 5. Cells with impaired Ssa2p•Mas5 complex have decreased cortical Cdc42 activity and exhibit monopolar intermittent growth pattern. 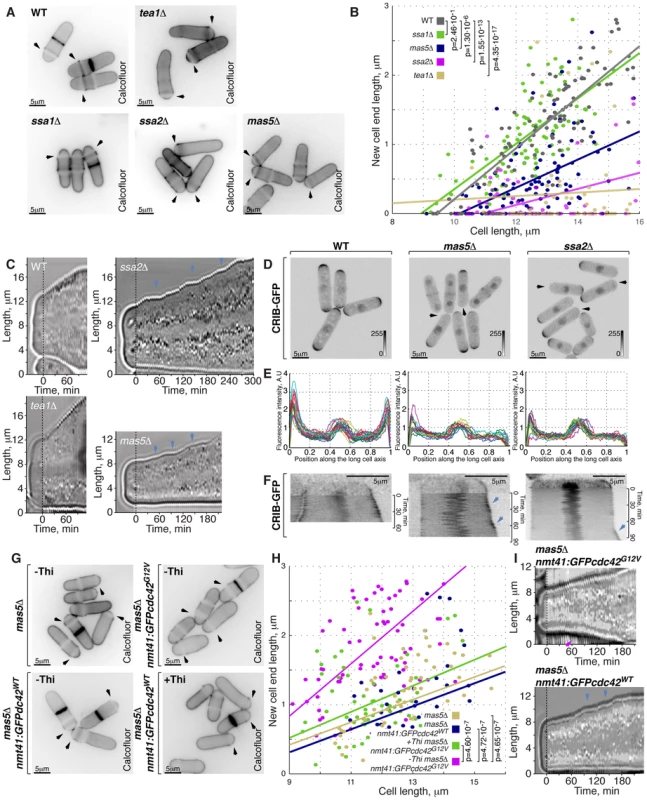
(A) Single z-plane images of calcofluor stained log-phase cells with indicated genotypes. Arrows point to birth-scars. (B) New-cell-end length as a function of cell length for cells with indicated genotypes. Points represent individual cells, lines represent the linear regression of a sample and p-values were obtained through ANCOVA. (C) Kymographs of bright-field microscopy time-lapse analyses of cells with the indicated genetic backgrounds grown in a perfusion chamber at room temperature. Arrowheads point to growth pauses characteristic to ssa2Δ and mas5Δ cells. (D) Shown are whole cell maximum intensity z-projections of scanning confocal micrographs of log-phase cells with indicated genotypes expressing CRIB-GFP, arrowheads point to cell tips lacking Cdc42 activity. (E) Quantification of CRIB-GFP intensities along the long cell axis in log-phase wild type (left panel), mas5Δ (middle panel) and ssa2Δ cells (right panels). Individual lines correspond to individual cells. (F) Kymographs of single z-plane spinning disk confocal microscopy time-lapse of wild type (left panel), mas5Δ (middle panel) and ssa2Δ cells (right panels) grown at room temperature. Arrows point to intermittent Cdc42 activity bursts associated with longitudinal growth seen in mas5Δ and ssa2Δ cells. (G) Single z-plane images of calcofluor stained log-phase cells with indicated genotypes grown in minimal medium in the presence or absence of thiamine for 30 hours and shifted to YE medium for 7 hours prior to fixation and staining. Arrows point to birth-scars. (H) New-cell-end length as a function of total cell length for cells presented in (G). Points represent individual cells, lines represent the linear regression of a sample and p-values were obtained through ANCOVA. (I) Kymographs of bright-field microscopy time-lapse analyses of cells with the indicated genetic backgrounds grown in a perfusion chamber at room temperature. Blue arrowheads point to growth pauses and a magenta arrowhead indicates onset of bipolar growth. Scale bars, 5 µm. Progression through the cell cycle was not affected in mas5Δ and ssa2Δ mutants, as evaluated by FACS (Fluorescence Activated Cell Sorting) analysis (Fig. S5B), suggesting that the monopolar growth in these mutants could not be explained by delaying entry into the G2 phase of the cell cycle.
Subcellular distribution of the polarisome components Tea1-YFP, Tea4-GFP and Pom1-GFP was unaffected by mas5 gene deletion (Fig. S5C–S5F). In contrast, localization of the β-glucan synthase GFP-Cps1 was predominantly monopolar late into the G2 phase of the cell cycle (Fig. S5C and S5G). Similarly, exocyst subunit Sec6 localized to only one cell tip in a majority of mas5Δ cells (Fig. S5G and S5K). Imaging of LifeAct-GFP [67] showed that the actin cytoskeletal structures were also enriched at just one cell tip of late G2 phase mas5Δ cells (Fig. S5I, S5C, S5J). Consistently, For3-GFP and Myo52-GFP were also monopolar in most mas5Δ cells longer than 9 µm (Fig. S5C, S5K and S5L).
Next, we performed live imaging of wild type, tea1Δ, ssa2Δ and mas5Δ cells grown in a perfusion chamber at room temperature (Supplemental Movie 1, see Materials and Methods). After an initial phase of monopolar growth, the newly born wild type cells underwent NETO and assumed bipolar growth pattern clearly seen in the kymograph (Fig. 5C). As expected [27] tea1Δ cells maintained a constitutively monopolar growth pattern and were rarely seen undergoing NETO (Fig. 5C). Consistent with the data presented earlier, over 90% of ssa2Δ and mas5Δ cells grew only at the old cell tip. Surprisingly, all mutant cells exhibited an intermittent growth pattern (Fig. 5C, n>10 cells) with periods of complete growth arrest lasting from 5 to over 30 minutes followed by growth bursts of 5 to over 30 minutes.
Active Cdc42, as visualized by CRIB-GFP, also localized to one cell tip in cells lacking Mas5 or Ssa2 (Fig. 5D, 3% of wild type, 93% of ssa2Δ and 58% of mas5Δ late G2 cells were monopolar). Furthermore, the absolute levels of CRIB-GFP signal at the polar cortex were diminished in ssa2Δ and mas5Δ cells (Fig. 5E). The time-lapse spinning-disk confocal microscopy revealed that the CRIB-GFP recruitment to the cell tips was unstable in mas5Δ and ssa2Δ cells (Fig. 5F). In wild type cells, CRIB-GFP signal at the continuously growing old cell end displayed only minor intensity oscillations during the course of imaging. On the other hand, CRIB-GFP levels at the new cell end underwent major on-off oscillations prior to NETO but eventually stabilized exhibiting only minor oscillations (Fig. 5F, left panel, also see [40]). However, CRIB-GFP levels underwent on-off oscillations at both tips throughout the cell cycle in mas5Δ and ssa2Δ cells. We observed that high CRIB-GFP levels at the old cell end coincided with periods of tip growth (Fig. 5F, middle and right panels). CRIB-GFP levels at the new cell end of ssa2Δ and mas5Δ cells underwent on-off oscillations that would rarely stabilize to allow for noticeable tip elongation during the course of imaging. These results suggest that the intermittent growth pattern of ssa2Δ and mas5Δ cells is a consequence of the unstable Cdc42 activity at the cell tips. Furthermore, the decreased Cdc42 activity at the new cell end could explain the monopolar growth pattern of mutants lacking Ssa2-Mas5 chaperone complex.
The low levels of active Cdc42 in mas5Δ cells prompted us to explore the distribution of the Cdc42-GAP and GEFs in the mutant. Both Gef1-3YFP and Scd1-GFP exhibited bipolar localization in majority of late wild type cells but were predominantly monopolar in cells lacking Mas5 (Fig. S5M, S5N and S5C). The Cdc42-GAP, Rga4-GFP, remained restricted to the lateral cell cortex in ∼90% in mas5Δ cells (Fig. S5O). Levels of cortically associated Cdc42 regulators appeared decreased in mas5Δ cells as compared to the wild type. The average intensity of Gef1-3YFP at a cell tip was decreased by 38±28% whereas Scd1-GFP signal diminished by 53±27% (pWelch's t-test = 2.51⋅10−6 and 7.94⋅10−7 respectively, n>25 cells per sample). A 20±22% decrease in cortically associated levels was also observed for the Rga4-GFP in cells lacking Mas5 (pWelch's t-test = 2.71⋅10−4, n>25 cells per sample). The total protein levels of the three examined Cdc42 regulators were also diminished in mas5Δ cells (Fig. S5P). Gef1-3YFP and Scd1-GFP levels decreased by 18% and 48% respectively and a 21% decrease was observed for Rga4-GFP.
These results suggested that the lack of Ssa2-Mas5 chaperone complex resulted in monopolar growth due to insufficient function of the Cdc42 GTPase module. Indeed, mild overexpression of the constitutively active Cdc42-G12V mutant but not wild type Cdc42 in mas5Δ cells led to its enrichment at the cell tips (Fig. S5Q) and a significant increase in bipolar growth (Fig. 5G and 5H, p = 4.65⋅10−7). Interestingly, upregulation of Cdc42 activity also largely rescued the intermittent growth phenotype of mas5Δ cells, as observed by time-lapse microscopy (Fig. 5I, continuous growth was sustained in 11/20 cells overexpressing Cdc42-G12V and only 3/20 cells overexpressing the wild type Cdc42).
We wondered if the abnormal monopolar growth observed in cells lacking Mas5-Ssa2 chaperone complex could be a manifestation of an upregulated heat stress response.
Cells lacking Mas5-Ssa2 chaperone complex exhibit elevated levels of heat stress-associated transcription
We investigated the gene expression profiles of mas5Δ, ssa2Δ and heat-stressed wild type S. pombe cells by RNAseq analysis (See Materials and Methods for experimental details). Interestingly, we observed a striking similarity between their gene expression programs - majority of genes that >2-fold up - or down-regulated in the chaperone mutants exhibited similar regulation in heat-stressed cells (Fig. 6A).
Fig. 6. Cells with impaired function of the Ssa2p•Mas5 chaperone complex exhibit elevated levels of heat-stress associated transcription. 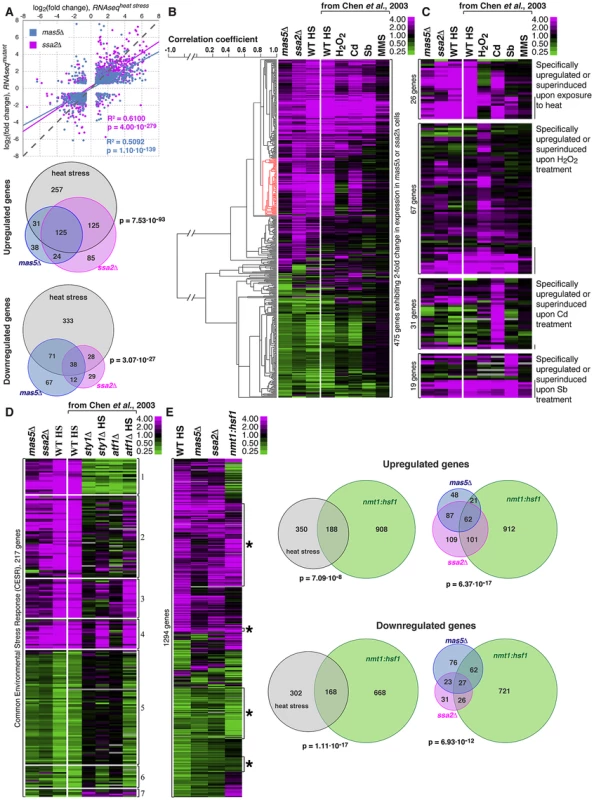
(A) Top panel represents correlation analysis between gene expression profiles of mas5Δ (in blue) or ssa2Δ (in magenta) cells and wild type cells exposed to heat stress. Individual dots represent individual genes. Pearson's correlation coefficients and statistical significance are indicated. Venn diagrams (bottom panel) analyze the overlap in >2-fold differentially regulated genes between mas5Δ (in blue), ssa2Δ (in magenta) cells and heat-stressed wild type cells (in gray). (B) Comparison between gene expression profiles of mas5Δ, ssa2Δ and wild type cells under indicated environmental stresses; gene clustering presented on the left. (C) Expression profiles of genes specifically upregulated by distinct environmental stresses in mas5Δ and ssa2Δ cells and wild type cells under indicated environmental stresses. (D) Analysis of expression of common environmental stress response genes in exponentially growing mas5Δ and ssa2Δ cells, and sty1Δ, atf1Δ and wild type cells under indicated conditions. (E) Left panel shows the comparison between gene expression profiles of wild type heat-stressed cells, mas5Δ, ssa2Δ and cells overexpressing Hsf1, as detected by RNAseq analysis. Venn diagrams (right panel) analyze the overlap in >2-fold differentially regulated genes between mas5Δ (in blue), ssa2Δ (in magenta) cells, heat-stressed wild type cells (in gray) and cells overexpressing Hsf1 (in green). Our RNAseq-based profiling of heat stress exhibited remarkable overlap with the microarray data on different abiotic stresses [62] allowing direct comparison between two technology platforms (Fig. S6A). The correlation between expression profiles in mas5Δ and ssa2Δ strains and wild type cells under distinct stress conditions confirmed similarities between expression profiles of the chaperone mutants and wild type cells exposed to heat and also suggested some similarities to cadmium-treated cultures (Fig. 6B and Fig. S6B–C). Gene expression linkage analysis identified a cluster enriched in genes reported as exclusively induced or super-induced in response to heat stress (Fig. 6B, red cluster, p = 2.65⋅10−7) [62]. Moreover, when we considered genes distinctly induced by different abiotic stresses, only the heat stress-specific genes were consistently upregulated in mas5Δ and ssa2Δ cells (Fig. 6C, pmas5Δ = 1.44⋅10−2, pssa2Δ = 5.25⋅10−6 for 1.5-fold cut-off). No significant overlaps were detected for genes specifically up-regulated by other types of stress (Fig. 6C; pbinomial test>0.25 for oxidative, osmotic stress and cadmium treatment). The microarray and qPCR analysis confirmed that genes specifically induced in response to heat stress were upregulated in mas5Δ cells and genes downregulated during heat-stress exhibited similar behavior in mas5Δ cells (Fig. S6D; See Supplementary Table S5 for detailed microarray analysis). We concluded that the transcriptional profiles of the chaperone mutants qualitatively matched the heat stress gene expression signature.
Both mas5Δ and ssa2Δ mutants exhibited differential regulation of most common environmental stress response (CESR) genes (Fig. 6D and [62]). Stress response is activated in part by MAP kinase (MAPK) signaling through the MAP kinase Sty1 and Atf1 transcription factor [59], [68], [69]. Interestingly, the most consistently up - or down-regulated CESR gene clusters were those independent of either Sty1 and/or Atf1 (Fig. 6D, clusters 4 and 6; Fig. S6E). While many Sty1 - and Atf1-dependent CESR genes were also differentially regulated in the chaperone mutants (Fig. 6D), we failed to observe synthetic genetic interactions between mas5Δ and cells lacking Sty1 [70] or between mas5Δ and mutants carrying the constitutively activated MAPK kinase allele wis1.DD [59]. The phenotype of mas5Δ cells was also unchanged in the presence of Polo kinase mutant alleles plo1S402A and plo1S402E known to be defective in some aspects of the MAPK signaling [55].
We next wondered how the gene expression programs of heat-stressed cells and the chaperone mutants compared to that of cells overexpressing Hsf1. To this end, we performed RNAseq analysis of nmt1:hsf1 cells grown in the absence of thiamine for 20 hours, at a time point when most Hsf1-overexpressing cells exhibited monopolar growth (Fig. 3). We observed a significant overlap between the sets of genes regulated by Hsf1 overexpression and the heat stress (Fig. 6E). Importantly, almost half of the differentially expressed genes in mas5Δ and/or ssa2Δ cells were similarly regulated in cells overexpressing Hsf1 (Fig. 6E). Many of those genes appeared the heat stress-responsive Hsf1-targets (Fig. S6F). It is possible that genes up-/down-regulated in the chaperone mutants that were not affected by Hsf1 overexpression could be regulated through other pathways. Alternatively, overexpression of Hsf1 could fail to fully recapitulate its native post-transcriptional activation.
Taken together, our gene expression analyses imply that cells lacking the Mas5-Ssa2 chaperone complex exhibit elevated levels of heat stress-associated transcription, at least in part due to abnormal activation of Hsf1.
High Hsf1 activity accounts for the growth pattering defects in cells lacking Ssa2-Mas5 chaperone complex
We wondered if polarized growth defects observed in the chaperone mutants were in fact related to high Hsf1 activity. To test this hypothesis, we sought to down-regulate Hsf1 expression in mas5Δ cells by promoter replacement. The nmt81-hsf1 transcriptional shut-off allows full depletion of Hsf1 but the strains carrying this construct do not germinate and therefore are not suitable for genetic analyses (Fig. S3A and data not shown). Thus we replaced the native hsf1 promoter with a recently reported uracil-regulatable purg1 promoter [71]. Repression of the purg1-driven Hsf1 in wild type cells did not lead to growth arrest, suggesting that even in its repressed, basal state, purg1 promoter allows Hsf1 expression sufficient for supporting vegetative growth. In line with this, we did not observe differences in the basal levels of Hsp104-GFP reporter upon repression of purg1-hsf1 in unstressed wild type cells (Fig. 7A–B). However, Hsp104-GFP levels were strongly reduced in mas5Δ cells when the purg1-driven Hsf1 expression was repressed (Fig. 7A–B).
Fig. 7. Repression of Hsf1 expression in mas5Δ cells rescues the growth patterning defects. 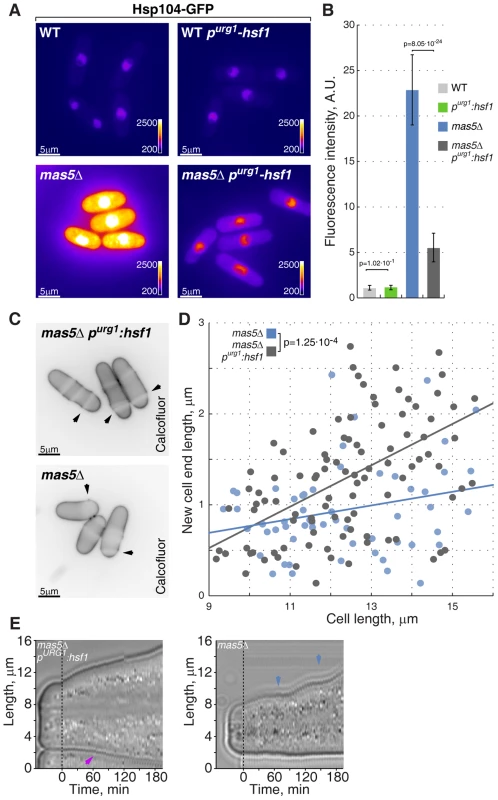
(A) Hsp104-GFP expressing wild type and mas5Δ cells where Hsf1 expression was under the regulation of the native or inducible purg1 promoter were grown in minimal medium lacking uracil for 36 hours at 24°C. Shown are pseudocolored epifluorescence images. Image contrast is reported using color wedges. Scale bars, 5 µm. (B) Quantification of GFP fluorescence in cells presented in (A). (C) Single z-plane images of calcofluor stained log-phase cells with indicated genotypes grown as described in (A). Arrowheads point to birth-scars. (D) New-cell-end length as a function of total cell length for cells with indicated genotypes. Points represent individual cells, lines represent the linear regression of a sample and the p-value was obtained through ANCOVA. (E) Kymographs of bright-field microscopy time-lapse analyses of cells with the indicated genetic backgrounds grown in a perfusion chamber at room temperature. Blue arrowheads point to growth pauses and a magenta arrowhead indicates onset of bipolar growth. Importantly, down-regulation of Hsf1 largely corrected the growth patterning defects of the chaperone mutant cells. Calcofluor staining revealed a significant increase in bipolar growth in mas5Δ cells with repressed purg1-hsf1 expression as compared to those carrying the wild type hsf1 allele (Fig. 7C–D). Curiously, down-regulation of Hsf1 also rescued the intermittent growth phenotype, with mas5Δ purg1-hsf1 cells demonstrating continuous tip extension when the promoter was repressed (Fig. 7E; 12/20 mas5Δ purg1-hsf1 cells sustained continuous tip extension vs 5/20 mas5Δ cells).
Taken together, our results indicate that high Hsf1 activity leads to downregulation of Cdc42 module resulting in inability to support normal bipolar growth. Following stress-triggered Hsf1 activation, the Hsp70-Hsp40 chaperone pair Ssa2 and Mas5 could function to repress it, thus allowing cells to reset their transcriptional output and return to normal growth.
Discussion
Organisms rapidly adjust their physiology in response to environmental stimuli such as changes in nutrient and mate availability, injury and abiotic stresses. A variety of stresses, including heat and exposure to osmolytes, trigger transient depolarization of the actin cytoskeleton and the cell wall remodeling machinery [25], [50], [53], [55]. After a period of adaptation to new conditions, the polarized growth is restored (Fig. 1 and S1). Our work focuses on this latter phase of stress response when cells resume polarized growth in an altered environment.
Following heat stress, the GTP-bound, active Cdc42 is no longer confined to the cell poles and redistributes throughout the entire cortex (Fig. 2). Cdc42 functions at the vertex of cellular growth (reviewed in [23], [72]) so its mis-localization likely propagates down to the cell wall remodeling machinery by delocalizing exocyst and the actin cytoskeleton (Fig. 1 and S1). Redistribution of the growth machinery throughout the cortex has been proposed to function in strengthening the cell wall and possibly to fix injuries caused by a temperature increase [49], [73], [74], although we did not readily detect major changes in cell wall appearance in heat-stressed S. pombe cells by electron microscopy (data not shown). The reported diffusion rates of the membrane-bound Cdc42 are relatively low [75] suggesting that mis-localization is likely due to Cdc42 activation occurring throughout the cortex rather than the lateral movement of active GTPase from the cell tips. Overexpression of Gef1 is sufficient to promote Cdc42 activation throughout the entire plasma membrane [76] suggesting that modulation of Gef1 activity and/or membrane recruitment upon heat stress could allow for ectopic activation of Cdc42. Interestingly, inhibition of the evolutionary conserved NDR kinase Orb6 promotes invasion of Cdc42 into the lateral cortical domain possibly through direct phosphorylation of Gef1 [37]. Activity of GEFs could be also modulated by interaction with plasma membrane proteins sensing the environment or the structural integrity of the cell wall. For instance, in budding yeast the cell wall sensors Wsc1 and Mid2 have been shown to interact with the GEF for Rho1, Rom2, and stimulate the Rho1 nucleotide exchange [51]. Since all landmark proteins including Pom1, the direct modulator of Rga4/Cdc42-GAP activity [39], remain properly polarized during heat stress response (Fig. 1), the mechanisms regulating delocalization of active Cdc42 are likely to function independently of the polarisome.
Once the initial phase of stress response is over, cells must return to normal growth in new conditions. Curiously, transient depolarization of the growth machinery upon mild heat stress is followed by a prolonged period of monopolar growth ([21], Fig. 1). Since the entire cell population becomes predominantly monopolar it means that heat stress could delay both the onset of NETO in newly born cells and re-initiation of persistent bipolar growth in late G2 phase of the cell cycle. The predominantly monopolar growth pattern in cells adapting to heat stress could be possibly explained by the recently developed model correlating the cell length to the bipolar Cdc42 activation [40]. The salient prediction of the model is that reduction of the positive relative to the negative Cdc42 feedback would result in an increased incidence of monopolar cells. Reduction in cortical levels of Cdc42-GEFs in heat-stressed cells could indeed weaken the positive feedback [42], [77], [78]. It is known that monopolar growth does result from the perturbed regulation of Cdc42 activity. S. pombe cells lacking Cdc42-GEF Gef1 have been shown to delay NETO [76]. Similarly, mutants in the Cdc42 effector PAK kinase that likely functions as a part of Cdc42 activity feedback [40] are monopolar [79]. The nature of negative feedback is less clear but it is unlikely to be limited to Cdc42-GAP [40], [41]. Alternatively, the heat-induced chaperones could directly modulate the cytoskeletal dynamics [80], [81] and in turn, affect the Cdc42 module activity.
Of note, cells with high levels of heat stress associated transcription either due to overexpression of the heat shock transcription factor Hsf1 or lacking the chaperone complex Ssa2-Mas5, also grow in a monopolar fashion and exhibit decreased cortical levels of active Cdc42 (Fig. 3 and Fig. 5). Unlike in a polarisome-deficient monopolar mutant tea1Δ, the active Cdc42 can be recruited to both tips of mas5Δ cells (Fig. S5R). Yet, only the old cell end that was inherited from the mother is capable of maintaining Cdc42 activity levels sufficient for growth (Fig. 5F). Consistent with an idea that low Cdc42 activity does not allow bipolar growth in cells where the heat stress response is on, mild overexpression of the constitutively active Cdc42 mutant rescued the growth patterning defects of mas5Δ cells (Fig. 5G–I).
Once wild type cells attain the new steady state growth at elevated temperature, the cortical activity of Cdc42 returns to normal and the bipolar growth is resumed (Fig. S2). Interestingly, the abundance of Cdc42 regulators is inversely correlated with environmental temperature (Fig. S2H), which may suggest a degree of temperature compensation in the Cdc42 module. The eukaryotic circadian clock is a typical example of a complex feedback network that buffers the period of oscillations against temperature (reviewed in [82]). The temperature compensation of the Neurospora clock relies on phosphorylation of the key regulator Frq by the casein kinase 2 [83], an ortholog of the gene isolated as orb5 in the fission yeast morphogenesis screen [84]. Furthermore, Hsf1 itself is known to regulate temperature compensation of the mammalian circadian clock [85]. The decrease in the Cdc42 activity in cells lacking the Ssa2-Mas5 complex could be potentially explained by over-compensation of Cdc42 module in the absence of the actual temperature increase.
Our work suggests that in fission yeast, the Hsp40 protein Mas5 functions in repressing Hsf1 activity as a specificity co-factor for the Hsp70 chaperone Ssa2. Heat shock transcription factors are extensively regulated [63] and both Hsp90 and Hsp70 chaperones have been proposed to function as their repressors [18], [19], [86]. We show that transcriptional depletion of Hsf1 indeed suppresses the heat-stress associated transcription in mas5Δ cells and also rescues the polarized growth defects in this chaperone mutant (Fig. 7). In line with this, the slow growth of budding yeast lacking Ssa1 and Ssa2 is rescued by a partial loss-of-function mutation in Hsf1 [87]. An interesting twist to the chaperone-mediated regulation is that mRNA abundance of these chaperones increases upon heat stress (1.9-fold and 3.3-fold increase, respectively), possibly placing Hsf1 activity within a negative feedback loop. The cytosolic Hsp70 chaperones could also act as primary sensors of stress at least in some conditions, e. g. Ssa1 in budding yeast has been shown to release Hsf1 from the inhibitory complex upon binding to the thiol-reactive compounds [88]. The human Mas5 and Ssa2 orthologs have been shown to bind Hsf1 in vitro [89], an interaction that we were unable to detect in fission yeast, possibly due to its transient nature.
Several reports suggested that subcellular distribution of Hsf1 could affect its activity (surveyed in [90]). Interestingly, we found that during the steady state growth in S. pombe, Hsf1 exhibited a predominantly nuclear localization but it was fully exported into the cytoplasm following heat stress (Fig. S7). However, in the absence of Mas5 and Ssa2 Hsf1 failed to exit the nuclei of heat-stressed cells, suggesting that the chaperone complex could also regulate Hsf1 through controlling its subcellular distribution (Fig. S7). The temporally resolved studies of Hsf1-driven transcription triggered by heat stress suggest that the initial burst of Hsf1 activity is rapidly followed by its repression [6], [8], [62], [91]. Removing Hsf1 from the nucleus could be one of the mechanisms to attenuate its activity.
The Mas5-like Hsp40 chaperones are highly promiscuous in their interactions with client proteins [65]. This property could potentially allow them to serve as sensors of global folding status linking the machineries handling environmental stress response and the normal cellular growth. Indeed, Ydj1, the budding yeast ortholog of Mas5, has been proposed to function as a growth rate sensor regulating G1/S entry [92], [93]. Curiously, bursts of fairly normal tip extension in individual mas5Δ or ssa2Δ cells are interspersed with periods of rest (Fig. 5), suggesting that one or more factors could become limiting for growth when availability of these chaperones is decreased. Given that the instantaneous growth rate in yeast has been negatively correlated to upregulation of heat-responsive genes [94], the Hsf1-chaperone interaction could function in a feedback loop sustaining the normal cellular growth. It is worth noting that while both upregulation of Cdc42 activity and partial downregulation of Hsf1 in the chaperone mutant cells rescued the polarity-related phenotypes, neither genetic manipulation ameliorated the slow doubling time of mas5Δ cells.
Placing the central transcriptional activator of the heat shock response under negative regulation by chaperones with a wide client base may ensure efficient and highly sensitive means of adapting to frequently changing environment. Moreover, this circuitry appears to be coopted in regulating the normal cell cycle progression as a function of growth rate. By regulating Hsf1 activity through the Ssa2-Mas5 chaperone system, fission yeast cells could optimize Cdc42-driven polarized growth under different temperature regiments. Given that the heat stress associated transcription modulates diverse cellular functions [1], similar adaptive dynamics are likely to operate in other physiological processes.
Materials and Methods
S. pombe strains and reagents
S. pombe strains used in this study and their genotypes are listed in Supplementary Table S1. Growth media and genetic methods were as described in [95]. The actin marker Lifeact-GFP was a kind gift from M. Balasubramanian. The cell wall dyes Calcofluor White and FITC-Lectin were obtained from Sigma-Aldrich (St. Louis, MO, USA). We placed GFP under the control of the hsp104 promoter region (1.5 kb upstream of the start codon) and the 3′ UTR (0.7 kb downstream of the stop codone) to construct a reporter of heat-activated expression. The construct was integrated into hsp104 chromosomal locus without disrupting the Hsp104 ORF. For Hsf1 overexpression and thiamine-responsive transcriptional shut-off we replaced the hsf1 promoter with nmt1 and nmt81 promoters respectively at the native chromosomal locus. Similarly, for uracil-responsive modulation of Hsf1 expression we replaced the hsf1 promoter with urg1 promoter (675 bp upstream of the start codon) at the native chromosomal locus.
Fluorescence Activated Cell Sorting (FACS)
FACS was performed on MACSQuant Analyzer (Miltenyi Biotec GmbH, Bergisch Gladbach, Germany) using a protocol described by [96] with RNase obtained from Roche (Basel, Switzerland) and propidium-iodide from Sigma-Aldrich (St. Louis, MO, USA).
Microscopy
Epifluorescence images were collected using mercury lamp as an illumination source with appropriate sets of filters on a Zeiss Axiovert 200M microscope (Carl Zeiss, Inc.) using an Plan Apochromat 100X, 1.4 N.A. objective lens, CoolSnap camera (Photometrics, Tucson, AZ) and Uniblitz shutter driver (Photonics, Rochester, NY) under the control of Metamorph software package (Universal Imaging, Sunnyvale, CA). We acquired either single z-planes or whole cell image stacks that consisted of 9 z-sections with 0.5 µm spacing. The z-stack maximum or average projection images were obtained by ImageJ 1.46b software package (http://rsb.info.nih.gov/ij/; National Institutes of Health, Bethesda, MD, USA). The same setup was also used in DIC microscopy on fission yeast cells grown in liquid YES medium using ONIX perfusion chambers (CellASIC, Hayward, CA, USA) under the control of the proprietor software and the flow of 4 psi.
Scanning confocal microscopy was performed on a LSM510 microscope equipped with a Plan Apochromat 100X, 1.4 N.A. objective lens, a 488-nm argon laser and a 543-nm HeNe laser. We acquired either single z-planes or whole cell image stacks that consisted of 9 z-sections with 0.5 µm spacing. The z-stack maximum or average projection images were obtained by ImageJ 1.46b software package.
Time-lapse fluorescence microscopy images were generated on Nikon TiE system (CFI Plan Apochromat VC 100XH 1.4 N.A. objective) equipped with Yokogawa CSU-X1-A1 spinning disk unit, the Photometrics CoolSNAP HQ2 camera and a DPSS 491 nm 100 mW and DPSS561 nm 50 mW laser illumination under the control of MetaMorph Premier Ver. 7.7.5. We acquired single z-plane images. Imaging was performed on S. pombe cells placed in sealed growth chambers containing 2% agarose YES medium.
Image analyses
Longitudinal growth in lectin stained cells was assessed manually as a distance from the cell tip to the lectin signal along the long cell axis.
New-cell-end growth was quantified from calcofluor-stained cells as the distance from the birth scar to the proximal cell tip and expressed as a function of total cell length. We employed ANCOVA analysis to assess the statistical significance of variation in new-end growth between cells with different genotypes (n>30 cells per genotype). For simplified presentation we also estimated average new-end length in cells 12–15 µm long while making sure that the average cell length between samples was comparable, p-values were obtained using Welch's t-test.
ImageJ in-built line-scan module was used to create an intensity profile along the long axis of the cells expressing CRIB-GFP. The positional information was normalized as the percentage of the cell length. Cellular fluorescence profiles were calculated from background-subtracted images and the nuclear signal was set as one arbitrary unit since it did not vary significantly between samples. In Fig. 2C unscaled fluorescence intensities are reported and were obtained from cells with the same levels of total CRIB-GFP fluorescence.
Fluorescence levels of Cdc42 regulators were analyzed from background subtracted images by manually outlining the whole cells and cell tips. Reported relative changes in fluorescence intensities are based on the average calculated values.
ImageJ 1.46b software package was used for all image quantifications.
Total protein extraction, co-immunoprecipitation
For co-immunoprecipitation, yeast cells were grown to log phase, harvested and washed with Buffer A (6 mM Na2HPO4, 4 mM NaH2PO4.H2O, 150 mM NaCl, 2 mM EDTA, 50 mM NaF, 0.1 mM Na3VO4, protease inhibitor cocktail (Roche)). Cell pellets were resuspended in 200 µL of buffer A, mixed with glass beads and homogenized in a Mini Bead Beater (Biospec, Bartlesville, OK, USA) at 4°C. Total cell lysate were harvested and centrifuged (16,000× g, 10 min) to remove cell debris. Soluble fractions were adjusted to same total protein concentration using Buffer A and 350 µl were incubated with either mouse anti-myc (Milipore, Billerica, MA, USA) antibodies and Protein A beads (Invitrogen, Carlsbad, CA, USA) or just GFP-Trap beads (ChromoTek, Munich, Germany) for 1 hour. Beads were washed 6 times with 1 mL of buffer A and resuspended in 50 µL of SDS-Loading buffer.
For total protein quantifications, cells were pelleted and resuspended in 200 µl of BufferA (1.85M NaOH, 1M β-mercaptoethanol, 5× EDTA-free Complete Protease Inhibitor Cocktail (Roche)) and kept on ice for 10 min. We added 450 µl of water and 350 µl of 6.1N trichloroacetic acid. The samples were incubated in ice for 10 min and centrifuged for 10 min at 4°C. The obtained pellet was washed with 0.5 M Tris-base, dissolved in gel loading buffer and kept at 65°C for 10 min and 95°C for 3 min.
Protein samples were subjected SDS-PAGE and standard Western Blotting. Proteins of interest were probed by mouse anti-GFP (Roche), rabbit anti-HA (Roche) and mouse anti-myc (Milipore). Rabbit anti-HistoneH3 (Abcam, Cambridge, MA, USA) probing served to monitor sample loading. IRDye800 conjugated anti-mouse and IRDye700 conjugated anti-rabbit antibodies were used prior to analysis on the Odyssey Infrared Imaging system, all purchased from LI-COR Biosciences (Lincoln, NE, USA).
RNA sequencing (RNAseq) analyses
Cells were grown in standard YE (wild-type, mas5Δ, ssa2Δ) or MM media (wild type, nmt1:hsf1 without thiamine, nmt81:hsf1 with thiamine) at 24°C. A sample of wild type cells was also shifted from 24°C to 36°C for 15 minutes. RNA was extracted using Qiagen RNeasy Mini Kit (Qiagen, Venlo, Netherlands) following manufacturer's protocol. Samples were sequenced by single end 50 bp sequencing at the University of Utah's Bioinformatics and Microarray Next Generation Sequencing Shared Resources (Salt Lake City, Utah, USA) after Illumina TruSeq RNA Sample Prep with oligo-dT selection. The RNA sequencing results were analyzed using Galaxy software package [97]–[99]. Briefly, upon quality control using the FastQ Groomer [100] module, reads were mapped to the fission yeast genome using the Tophat module [101] and differential gene expression was assessed by the Cufflinks module [102]. Analyses of differential gene expression are presented in Supplementary Table S4. The relevant raw RNAseq data have been deposited in NCBI's Gene Expression Omnibus [103] and are accessible through GEO Series accession number GSE50156 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE50156).
Quantitative PCR
The RNA samples from mas5Δ and wild type cells grown at 24°C or shifted to 36°C for 45 min were prepared using phenol-chlorophorm extraction described by [104] and reverse transcribed using SuperScript III (Invitrogen) and random decamer primers (N10) at 50°C for 60 min. The reverse transcriptase was inactivated at 70°C for 10 min. cDNA samples were analyzed by quantitative real-time PCR (qPCR) using a StepOnePlus real-time PCR system (Applied Biosystems) and Fast SYBR Green Master Mix (Applied Biosystems) with 0.15 µM of each forward and reverse primers. The following cycling program was used: 95°C for 20 s, followed by 45 cycles of a three-step reaction, denaturation at 95°C for 5 s, annealing and extension at 60°C for 45 s. The primers used are listed in Supplementary Table S3. Data shown were normalized to the expression levels of actin (act1) and similar results were obtained when using GAPDH (gpd1, data not shown) to normalize the data.
Microarray analysis
The gene expression profile of mas5Δ cells was obtained as previously described by [104]. The significance of differential gene regulation between wild type and mas5Δ cell was assessed by the Welch's t-test (Supplementary Table S5). Only genes with p<0.05 and fold change >1.5 were included in subsequent analysis. The expression profiles for stress conditions were taken from [62].
Clustering analysis was performed using Gene Cluster [5] using uncentered correlation similarity metrics average-linkage method. The visualization was done using TreeView software [105].
Correspondence at the top plot was derived from [106]. Briefly, the genes were sorted into a list according to how pronounced their up/down-regulation was in mas5Δ as compared to wild type cells. The binomial test was then employed to generate p-values as a function of an increasing number of genes included into the comparison between mas5Δ expression profile and that of stressed cells.
Supporting Information
Zdroje
1. RichterK, HaslbeckM, BuchnerJ (2010) The heat shock response: life on the verge of death. Mol Cell 40 : 253–266.
2. AshburnerM (1970) Patterns of puffing activity in the salivary gland chromosomes of Drosophila. V. Responses to environmental treatments. Chromosoma 31 : 356–376.
3. AshburnerM, BonnerJJ (1979) The induction of gene activity in drosophilia by heat shock. Cell 17 : 241–254.
4. TissieresA, MitchellHK, TracyUM (1974) Protein synthesis in salivary glands of Drosophila melanogaster: relation to chromosome puffs. J Mol Biol 84 : 389–398.
5. EisenMB, SpellmanPT, BrownPO, BotsteinD (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 95 : 14863–14868.
6. GaschAP, SpellmanPT, KaoCM, Carmel-HarelO, EisenMB, et al. (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11 : 4241–4257.
7. Kloster-LandsbergM, HerbomelG, WangI, DerouardJ, Vourc'hC, et al. (2012) Cellular response to heat shock studied by multiconfocal fluorescence correlation spectroscopy. Biophys J 103 : 1110–1119.
8. ZobeckKL, BuckleyMS, ZipfelWR, LisJT (2010) Recruitment timing and dynamics of transcription factors at the Hsp70 loci in living cells. Mol Cell 40 : 965–975.
9. HarrisonCJ, BohmAA, NelsonHC (1994) Crystal structure of the DNA binding domain of the heat shock transcription factor. Science 263 : 224–227.
10. SorgerPK, PelhamHR (1988) Yeast heat shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell 54 : 855–864.
11. GalloGJ, PrenticeH, KingstonRE (1993) Heat shock factor is required for growth at normal temperatures in the fission yeast Schizosaccharomyces pombe. Molecular and cellular biology 13 : 749–761.
12. ThompsonS, CroftNJ, SotiriouA, PigginsHD, CrosthwaiteSK (2008) Neurospora crassa heat shock factor 1 Is an essential gene; a second heat shock factor-like gene, hsf2, is required for asexual spore formation. Eukaryotic cell 7 : 1573–1581.
13. JedlickaP, MortinMA, WuC (1997) Multiple functions of Drosophila heat shock transcription factor in vivo. The EMBO journal 16 : 2452–2462.
14. XiaoX, ZuoX, DavisAA, McMillanDR, CurryBB, et al. (1999) HSF1 is required for extra-embryonic development, postnatal growth and protection during inflammatory responses in mice. The EMBO journal 18 : 5943–5952.
15. AnckarJ, SistonenL (2011) Regulation of HSF1 function in the heat stress response: implications in aging and disease. Annu Rev Biochem 80 : 1089–1115.
16. AbravayaK, MyersMP, MurphySP, MorimotoRI (1992) The human heat shock protein hsp70 interacts with HSF, the transcription factor that regulates heat shock gene expression. Genes Dev 6 : 1153–1164.
17. BalerR, WelchWJ, VoellmyR (1992) Heat shock gene regulation by nascent polypeptides and denatured proteins: hsp70 as a potential autoregulatory factor. J Cell Biol 117 : 1151–1159.
18. ZouJ, GuoY, GuettoucheT, SmithDF, VoellmyR (1998) Repression of heat shock transcription factor HSF1 activation by HSP90 (HSP90 complex) that forms a stress-sensitive complex with HSF1. Cell 94 : 471–480.
19. GuoY, GuettoucheT, FennaM, BoellmannF, PrattWB, et al. (2001) Evidence for a mechanism of repression of heat shock factor 1 transcriptional activity by a multichaperone complex. J Biol Chem 276 : 45791–45799.
20. EspinetC, de la TorreMA, AldeaM, HerreroE (1995) An efficient method to isolate yeast genes causing overexpression-mediated growth arrest. Yeast 11 : 25–32.
21. MitchisonJM, NurseP (1985) Growth in cell length in the fission yeast Schizosaccharomyces pombe. J Cell Sci 75 : 357–376.
22. CastagnettiS, NovakB, NurseP (2007) Microtubules offset growth site from the cell centre in fission yeast. Journal of cell science 120 : 2205–2213.
23. ChangF, MartinSG (2009) Shaping fission yeast with microtubules. Cold Spring Harb Perspect Biol 1: a001347.
24. FeierbachB, VerdeF, ChangF (2004) Regulation of a formin complex by the microtubule plus end protein tea1p. The Journal of cell biology 165 : 697–707.
25. RupesI, JiaZ, YoungPG (1999) Ssp1 promotes actin depolymerization and is involved in stress response and new end take-off control in fission yeast. Mol Biol Cell 10 : 1495–1510.
26. MartinSG, McDonaldWH, YatesJR (2005) Tea4p links microtubule plus ends with the formin for3p in the establishment of cell polarity. Dev Cell 8 : 479–491.
27. MataJ, NurseP (1997) tea1 and the microtubular cytoskeleton are important for generating global spatial order within the fission yeast cell. Cell 89 : 939–949.
28. IshiguroJ, SaitouA, DuranA, RibasJC (1997) cps1+, a Schizosaccharomyces pombe gene homolog of Saccharomyces cerevisiae FKS genes whose mutation confers hypersensitivity to cyclosporin A and papulacandin B. Journal of bacteriology 179 : 7653–7662.
29. BendezuFO, MartinSG (2011) Actin cables and the exocyst form two independent morphogenesis pathways in the fission yeast. Mol Biol Cell 22 : 44–53.
30. BendezuFO, VincenzettiV, MartinSG (2012) Fission yeast Sec3 and Exo70 are transported on actin cables and localize the exocyst complex to cell poles. PLoS One 7: e40248.
31. EvangelistaM, BlundellK, LongtineMS, ChowCJ, AdamesN, et al. (1997) Bni1p, a yeast formin linking cdc42p and the actin cytoskeleton during polarized morphogenesis. Science 276 : 118–122.
32. MartinSG, RinconSA, BasuR, PerezP, ChangF (2007) Regulation of the formin for3p by cdc42p and bud6p. Mol Biol Cell 18 : 4155–4167.
33. RinconSA, YeY, Villar-TajaduraMA, SantosB, MartinSG, et al. (2009) Pob1 participates in the Cdc42 regulation of fission yeast actin cytoskeleton. Mol Biol Cell 20 : 4390–4399.
34. ZhangX, OrlandoK, HeB, XiF, ZhangJ, et al. (2008) Membrane association and functional regulation of Sec3 by phospholipids and Cdc42. J Cell Biol 180 : 145–158.
35. RossmanKL, DerCJ, SondekJ (2005) GEF means go: turning on RHO GTPases with guanine nucleotide-exchange factors. Nature reviews Molecular cell biology 6 : 167–180.
36. SinhaS, YangW (2008) Cellular signaling for activation of Rho GTPase Cdc42. Cell Signal 20 : 1927–1934.
37. DasM, WileyDJ, ChenX, ShahK, VerdeF (2009) The conserved NDR kinase Orb6 controls polarized cell growth by spatial regulation of the small GTPase Cdc42. Curr Biol 19 : 1314–1319.
38. RinconS, CollPM, PerezP (2007) Spatial regulation of Cdc42 during cytokinesis. Cell cycle 6 : 1687–1691.
39. TatebeH, NakanoK, MaximoR, ShiozakiK (2008) Pom1 DYRK regulates localization of the Rga4 GAP to ensure bipolar activation of Cdc42 in fission yeast. Curr Biol 18 : 322–330.
40. DasM, DrakeT, WileyDJ, BuchwaldP, VavylonisD, et al. (2012) Oscillatory dynamics of Cdc42 GTPase in the control of polarized growth. Science 337 : 239–243.
41. HowellAS, JinM, WuCF, ZylaTR, ElstonTC, et al. (2012) Negative feedback enhances robustness in the yeast polarity establishment circuit. Cell 149 : 322–333.
42. KozubowskiL, SaitoK, JohnsonJM, HowellAS, ZylaTR, et al. (2008) Symmetry-breaking polarization driven by a Cdc42p GEF-PAK complex. Curr Biol 18 : 1719–1726.
43. Wedlich-SoldnerR, AltschulerS, WuL, LiR (2003) Spontaneous cell polarization through actomyosin-based delivery of the Cdc42 GTPase. Science 299 : 1231–1235.
44. KimH, YangP, CatanutoP, VerdeF, LaiH, et al. (2003) The kelch repeat protein, Tea1, is a potential substrate target of the p21-activated kinase, Shk1, in the fission yeast, Schizosaccharomyces pombe. J Biol Chem 278 : 30074–30082.
45. Gomis-RuthS, WierengaCJ, BradkeF (2008) Plasticity of polarization: changing dendrites into axons in neurons integrated in neuronal circuits. Curr Biol 18 : 992–1000.
46. SlutskyB, BuffoJ, SollDR (1985) High-frequency switching of colony morphology in Candida albicans. Science 230 : 666–669.
47. BendezuFO, MartinSG (2013) Cdc42 explores the cell periphery for mate selection in fission yeast. Current biology : CB 23 : 42–47.
48. DyerJM, SavageNS, JinM, ZylaTR, ElstonTC, et al. (2013) Tracking shallow chemical gradients by actin-driven wandering of the polarization site. Current biology 23 : 32–41.
49. KonoK, SaekiY, YoshidaS, TanakaK, PellmanD (2012) Proteasomal degradation resolves competition between cell polarization and cellular wound healing. Cell 150 : 151–164.
50. DelleyPA, HallMN (1999) Cell wall stress depolarizes cell growth via hyperactivation of RHO1. J Cell Biol 147 : 163–174.
51. PhilipB, LevinDE (2001) Wsc1 and Mid2 are cell surface sensors for cell wall integrity signaling that act through Rom2, a guanine nucleotide exchange factor for Rho1. Mol Cell Biol 21 : 271–280.
52. GuoS, ShenX, YanG, MaD, BaiX, et al. (2009) A MAP kinase dependent feedback mechanism controls Rho1 GTPase and actin distribution in yeast. PLoS One 4: e6089.
53. BaoS, QyangY, YangP, KimH, DuH, et al. (2001) The highly conserved protein methyltransferase, Skb1, is a mediator of hyperosmotic stress response in the fission yeast Schizosaccharomyces pombe. J Biol Chem 276 : 14549–14552.
54. KanaiM, KumeK, MiyaharaK, SakaiK, NakamuraK, et al. (2005) Fission yeast MO25 protein is localized at SPB and septum and is essential for cell morphogenesis. EMBO J 24 : 3012–3025.
55. PetersenJ, HaganIM (2005) Polo kinase links the stress pathway to cell cycle control and tip growth in fission yeast. Nature 435 : 507–512.
56. RobertsonAM, HaganIM (2008) Stress-regulated kinase pathways in the recovery of tip growth and microtubule dynamics following osmotic stress in S. pombe. J Cell Sci 121 : 4055–4068.
57. GarciaP, TajaduraV, SanchezY (2009) The Rho1p exchange factor Rgf1p signals upstream from the Pmk1 mitogen-activated protein kinase pathway in fission yeast. Mol Biol Cell 20 : 721–731.
58. MaY, KunoT, KitaA, AsayamaY, SugiuraR (2006) Rho2 is a target of the farnesyltransferase Cpp1 and acts upstream of Pmk1 mitogen-activated protein kinase signaling in fission yeast. Mol Biol Cell 17 : 5028–5037.
59. ShiozakiK, ShiozakiM, RussellP (1998) Heat stress activates fission yeast Spc1/StyI MAPK by a MEKK-independent mechanism. Molecular biology of the cell 9 : 1339–1349.
60. HayesCE, GoldsteinIJ (1974) An alpha-D-galactosyl-binding lectin from Bandeiraea simplicifolia seeds. Isolation by affinity chromatography and characterization. J Biol Chem 249 : 1904–1914.
61. MayJW, MitchisonJM (1986) Length growth in fission yeast cells measured by two novel techniques. Nature 322 : 752–754.
62. ChenD, TooneWM, MataJ, LyneR, BurnsG, et al. (2003) Global transcriptional responses of fission yeast to environmental stress. Mol Biol Cell 14 : 214–229.
63. AkerfeltM, MorimotoRI, SistonenL (2010) Heat shock factors: integrators of cell stress, development and lifespan. Nat Rev Mol Cell Biol 11 : 545–555.
64. GalloGJ, SchuetzTJ, KingstonRE (1991) Regulation of heat shock factor in Schizosaccharomyces pombe more closely resembles regulation in mammals than in Saccharomyces cerevisiae. Molecular and cellular biology 11 : 281–288.
65. GongY, KakiharaY, KroganN, GreenblattJ, EmiliA, et al. (2009) An atlas of chaperone-protein interactions in Saccharomyces cerevisiae: implications to protein folding pathways in the cell. Mol Syst Biol 5 : 275.
66. LiGC, WerbZ (1982) Correlation between synthesis of heat shock proteins and development of thermotolerance in Chinese hamster fibroblasts. Proc Natl Acad Sci U S A 79 : 3218–3222.
67. RiedlJ, CrevennaAH, KessenbrockK, YuJH, NeukirchenD, et al. (2008) Lifeact: a versatile marker to visualize F-actin. Nat Methods 5 : 605–607.
68. LevinDE (2011) Regulation of cell wall biogenesis in Saccharomyces cerevisiae: the cell wall integrity signaling pathway. Genetics 189 : 1145–1175.
69. TodaT, DhutS, Superti-FurgaG, GotohY, NishidaE, et al. (1996) The fission yeast pmk1+ gene encodes a novel mitogen-activated protein kinase homolog which regulates cell integrity and functions coordinately with the protein kinase C pathway. Molecular and cellular biology 16 : 6752–6764.
70. ShiozakiK, RussellP (1995) Cell-cycle control linked to extracellular environment by MAP kinase pathway in fission yeast. Nature 378 : 739–743.
71. WattS, MataJ, Lopez-MauryL, MargueratS, BurnsG, et al. (2008) urg1: a uracil-regulatable promoter system for fission yeast with short induction and repression times. PLoS One 3: e1428.
72. PerezP, RinconSA (2010) Rho GTPases: regulation of cell polarity and growth in yeasts. The Biochemical journal 426 : 243–253.
73. KamadaY, JungUS, PiotrowskiJ, LevinDE (1995) The protein kinase C-activated MAP kinase pathway of Saccharomyces cerevisiae mediates a novel aspect of the heat shock response. Genes & development 9 : 1559–1571.
74. LevinDE (2005) Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiology and molecular biology reviews 69 : 262–291.
75. MarcoE, Wedlich-SoldnerR, LiR, AltschulerSJ, WuLF (2007) Endocytosis optimizes the dynamic localization of membrane proteins that regulate cortical polarity. Cell 129 : 411–422.
76. CollPM, TrilloY, AmetzazurraA, PerezP (2003) Gef1p, a new guanine nucleotide exchange factor for Cdc42p, regulates polarity in Schizosaccharomyces pombe. Mol Biol Cell 14 : 313–323.
77. EndoM, ShirouzuM, YokoyamaS (2003) The Cdc42 binding and scaffolding activities of the fission yeast adaptor protein Scd2. The Journal of biological chemistry 278 : 843–852.
78. IrazoquiJE, GladfelterAS, LewDJ (2003) Scaffold-mediated symmetry breaking by Cdc42p. Nature cell biology 5 : 1062–1070.
79. VerdeF, MataJ, NurseP (1995) Fission yeast cell morphogenesis: identification of new genes and analysis of their role during the cell cycle. J Cell Biol 131 : 1529–1538.
80. LiangP, MacRaeTH (1997) Molecular chaperones and the cytoskeleton. Journal of cell science 110 (Pt 13) 1431–1440.
81. MounierN, ArrigoAP (2002) Actin cytoskeleton and small heat shock proteins: how do they interact? Cell stress & chaperones 7 : 167–176.
82. HogeneschJB, UedaHR (2011) Understanding systems-level properties: timely stories from the study of clocks. Nat Rev Genet 12 : 407–416.
83. MehraA, ShiM, BakerCL, ColotHV, LorosJJ, et al. (2009) A role for casein kinase 2 in the mechanism underlying circadian temperature compensation. Cell 137 : 749–760.
84. SnellV, NurseP (1994) Genetic analysis of cell morphogenesis in fission yeast–a role for casein kinase II in the establishment of polarized growth. EMBO J 13 : 2066–2074.
85. BuhrED, YooSH, TakahashiJS (2010) Temperature as a universal resetting cue for mammalian circadian oscillators. Science 330 : 379–385.
86. LeachMD, BudgeS, WalkerL, MunroC, CowenLE, et al. (2012) Hsp90 Orchestrates Transcriptional Regulation by Hsf1 and Cell Wall Remodelling by MAPK Signalling during Thermal Adaptation in a Pathogenic Yeast. PLoS pathogens 8: e1003069.
87. HalladayJT, CraigEA (1995) A heat shock transcription factor with reduced activity suppresses a yeast HSP70 mutant. Mol Cell Biol 15 : 4890–4897.
88. WangY, GibneyPA, WestJD, MoranoKA (2012) The yeast Hsp70 Ssa1 is a sensor for activation of the heat shock response by thiol-reactive compounds. Mol Biol Cell 23 : 3290–3298.
89. ShiY, MosserDD, MorimotoRI (1998) Molecular chaperones as HSF1-specific transcriptional repressors. Genes & development 12 : 654–666.
90. VujanacM, FenaroliA, ZimarinoV (2005) Constitutive nuclear import and stress-regulated nucleocytoplasmic shuttling of mammalian heat-shock factor 1. Traffic 6 : 214–229.
91. RiegerTR, MorimotoRI, HatzimanikatisV (2005) Mathematical modeling of the eukaryotic heat-shock response: dynamics of the hsp70 promoter. Biophys J 88 : 1646–1658.
92. FerrezueloF, ColominaN, PalmisanoA, GariE, GallegoC, et al. (2012) The critical size is set at a single-cell level by growth rate to attain homeostasis and adaptation. Nat Commun 3 : 1012.
93. VergesE, ColominaN, GariE, GallegoC, AldeaM (2007) Cyclin Cln3 is retained at the ER and released by the J chaperone Ydj1 in late G1 to trigger cell cycle entry. Mol Cell 26 : 649–662.
94. LuC, BrauerMJ, BotsteinD (2009) Slow growth induces heat-shock resistance in normal and respiratory-deficient yeast. Mol Biol Cell 20 : 891–903.
95. GouldKL (2004) Protocols for experimentation with Schizosaccharomyces pombe. Methods 33 : 187–188.
96. SabatinosSA, ForsburgSL (2009) Measuring DNA content by flow cytometry in fission yeast. Methods Mol Biol 521 : 449–461.
97. GoecksJ, NekrutenkoA, TaylorJ (2010) Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome biology 11: R86.
98. BlankenbergD, Von KusterG, CoraorN, AnandaG, LazarusR, et al. (2010) Galaxy: a web-based genome analysis tool for experimentalists. Current protocols in molecular biology/edited by Frederick M Ausubel [et al] Chapter 19: Unit 19 10 11–21.
99. GiardineB, RiemerC, HardisonRC, BurhansR, ElnitskiL, et al. (2005) Galaxy: a platform for interactive large-scale genome analysis. Genome research 15 : 1451–1455.
100. BlankenbergD, GordonA, Von KusterG, CoraorN, TaylorJ, et al. (2010) Manipulation of FASTQ data with Galaxy. Bioinformatics 26 : 1783–1785.
101. TrapnellC, PachterL, SalzbergSL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25 : 1105–1111.
102. TrapnellC, WilliamsBA, PerteaG, MortazaviA, KwanG, et al. (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature biotechnology 28 : 511–515.
103. EdgarR, DomrachevM, LashAE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30 : 207–210.
104. PengX, KaruturiRK, MillerLD, LinK, JiaY, et al. (2005) Identification of cell cycle-regulated genes in fission yeast. Mol Biol Cell 16 : 1026–1042.
105. SaldanhaAJ (2004) Java Treeview–extensible visualization of microarray data. Bioinformatics 20 : 3246–3248.
106. IrizarryRA, WarrenD, SpencerF, KimIF, BiswalS, et al. (2005) Multiple-laboratory comparison of microarray platforms. Nat Methods 2 : 345–350.
Štítky
Genetika Reprodukční medicína
Článek Defending Sperm FunctionČlánek How to Choose the Right MateČlánek Conserved Translatome Remodeling in Nematode Species Executing a Shared Developmental TransitionČlánek Genome-Wide and Cell-Specific Epigenetic Analysis Challenges the Role of Polycomb in SpermatogenesisČlánek The Integrator Complex Subunit 6 (Ints6) Confines the Dorsal Organizer in Vertebrate EmbryogenesisČlánek Multiple bHLH Proteins form Heterodimers to Mediate CRY2-Dependent Regulation of Flowering-Time inČlánek Playing the Field: Sox10 Recruits Different Partners to Drive Central and Peripheral MyelinationČlánek A Minimal Nitrogen Fixation Gene Cluster from sp. WLY78 Enables Expression of Active Nitrogenase inČlánek Evolutionary Tuning of Protein Expression Levels of a Positively Autoregulated Two-Component System
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 10- Je „freeze-all“ pro všechny? Odborníci na fertilitu diskutovali na virtuálním summitu
- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
-
Všechny články tohoto čísla
- Defending Sperm Function
- How to Choose the Right Mate
- A Mutation in the Gene in Labrador Retrievers with Hereditary Nasal Parakeratosis (HNPK) Provides Insights into the Epigenetics of Keratinocyte Differentiation
- Conserved Translatome Remodeling in Nematode Species Executing a Shared Developmental Transition
- A Novel Actin mRNA Splice Variant Regulates ACTG1 Expression
- Tracking Proliferative History in Lymphocyte Development with Cre-Mediated Sister Chromatid Recombination
- Correlated Occurrence and Bypass of Frame-Shifting Insertion-Deletions (InDels) to Give Functional Proteins
- Chimeric Protein Complexes in Hybrid Species Generate Novel Phenotypes
- Loss of miR-10a Activates and Collaborates with Activated Wnt Signaling in Inducing Intestinal Neoplasia in Female Mice
- Both Rare and Copy Number Variants Are Prevalent in Agenesis of the Corpus Callosum but Not in Cerebellar Hypoplasia or Polymicrogyria
- Reverse PCA, a Systematic Approach for Identifying Genes Important for the Physical Interaction between Protein Pairs
- Partial Deletion of Chromosome 8 β-defensin Cluster Confers Sperm Dysfunction and Infertility in Male Mice
- Genome-Wide and Cell-Specific Epigenetic Analysis Challenges the Role of Polycomb in Spermatogenesis
- Coordinate Regulation of Mature Dopaminergic Axon Morphology by Macroautophagy and the PTEN Signaling Pathway
- Cooperation between RUNX1-ETO9a and Novel Transcriptional Partner KLF6 in Upregulation of in Acute Myeloid Leukemia
- Mobility of the Native Conjugative Plasmid pLS20 Is Regulated by Intercellular Signaling
- FliZ Is a Global Regulatory Protein Affecting the Expression of Flagellar and Virulence Genes in Individual Bacterial Cells
- Specific Tandem Repeats Are Sufficient for Paramutation-Induced Trans-Generational Silencing
- Condensin II Subunit dCAP-D3 Restricts Retrotransposon Mobilization in Somatic Cells
- Dominant Mutations in Identify the Mlh1-Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway
- The Insulator Homie Promotes Expression and Protects the Adjacent Gene from Repression by Polycomb Spreading
- Human Intellectual Disability Genes Form Conserved Functional Modules in
- Coordination of Cell Proliferation and Cell Fate Determination by CES-1 Snail
- ORFs in Drosophila Are Important to Organismal Fitness and Evolved Rapidly from Previously Non-coding Sequences
- Different Roles of Eukaryotic MutS and MutL Complexes in Repair of Small Insertion and Deletion Loops in Yeast
- The Spore Differentiation Pathway in the Enteric Pathogen
- Acceleration of the Glycolytic Flux by Steroid Receptor Coactivator-2 Is Essential for Endometrial Decidualization
- The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay
- Genome Wide Analysis Reveals Zic3 Interaction with Distal Regulatory Elements of Stage Specific Developmental Genes in Zebrafish
- Xbp1 Directs Global Repression of Budding Yeast Transcription during the Transition to Quiescence and Is Important for the Longevity and Reversibility of the Quiescent State
- The Integrator Complex Subunit 6 (Ints6) Confines the Dorsal Organizer in Vertebrate Embryogenesis
- Incorporating Motif Analysis into Gene Co-expression Networks Reveals Novel Modular Expression Pattern and New Signaling Pathways
- The Bacterial Response Regulator ArcA Uses a Diverse Binding Site Architecture to Regulate Carbon Oxidation Globally
- Direct Monitoring of the Strand Passage Reaction of DNA Topoisomerase II Triggers Checkpoint Activation
- Multiple bHLH Proteins form Heterodimers to Mediate CRY2-Dependent Regulation of Flowering-Time in
- A Reversible Histone H3 Acetylation Cooperates with Mismatch Repair and Replicative Polymerases in Maintaining Genome Stability
- ALS-Associated Mutations Result in Compromised Alternative Splicing and Autoregulation
- Robust Demographic Inference from Genomic and SNP Data
- Preferential Binding to Elk-1 by SLE-Associated Risk Allele Upregulates Expression
- Rad52 Sumoylation Prevents the Toxicity of Unproductive Rad51 Filaments Independently of the Anti-Recombinase Srs2
- The Serum Resistome of a Globally Disseminated Multidrug Resistant Uropathogenic Clone
- Identification of 526 Conserved Metazoan Genetic Innovations Exposes a New Role for Cofactor E-like in Neuronal Microtubule Homeostasis
- SUMO Localizes to the Central Element of Synaptonemal Complex and Is Required for the Full Synapsis of Meiotic Chromosomes in Budding Yeast
- Integrated Enrichment Analysis of Variants and Pathways in Genome-Wide Association Studies Indicates Central Role for IL-2 Signaling Genes in Type 1 Diabetes, and Cytokine Signaling Genes in Crohn's Disease
- Genome-Wide High-Resolution Mapping of UV-Induced Mitotic Recombination Events in
- Genome-Wide Analysis of Cell Type-Specific Gene Transcription during Spore Formation in
- Playing the Field: Sox10 Recruits Different Partners to Drive Central and Peripheral Myelination
- Two Portable Recombination Enhancers Direct Donor Choice in Fission Yeast Heterochromatin
- Mining the Human Phenome Using Allelic Scores That Index Biological Intermediates
- Yeast Tdh3 (Glyceraldehyde 3-Phosphate Dehydrogenase) Is a Sir2-Interacting Factor That Regulates Transcriptional Silencing and rDNA Recombination
- A Minimal Nitrogen Fixation Gene Cluster from sp. WLY78 Enables Expression of Active Nitrogenase in
- A Review of Bacteria-Animal Lateral Gene Transfer May Inform Our Understanding of Diseases like Cancer
- High Throughput Sequencing Reveals Alterations in the Recombination Signatures with Diminishing Spo11 Activity
- Partitioning the Heritability of Tourette Syndrome and Obsessive Compulsive Disorder Reveals Differences in Genetic Architecture
- Eleven Candidate Susceptibility Genes for Common Familial Colorectal Cancer
- A GDF5 Point Mutation Strikes Twice - Causing BDA1 and SYNS2
- Systematic Unraveling of the Unsolved Pathway of Nicotine Degradation in
- Natural Genetic Variation of Integrin Alpha L () Modulates Ischemic Brain Injury in Stroke
- Evolutionary Tuning of Protein Expression Levels of a Positively Autoregulated Two-Component System
- Evolutionary Change within a Bipotential Switch Shaped the Sperm/Oocyte Decision in Hermaphroditic Nematodes
- Limiting of the Innate Immune Response by SF3A-Dependent Control of MyD88 Alternative mRNA Splicing
- Multiple Signaling Pathways Coordinate to Induce a Threshold Response in a Chordate Embryo
- Distinct Regulatory Mechanisms Act to Establish and Maintain Pax3 Expression in the Developing Neural Tube
- Genome Wide Analysis of Narcolepsy in China Implicates Novel Immune Loci and Reveals Changes in Association Prior to Versus After the 2009 H1N1 Influenza Pandemic
- Mismatch Repair Genes and Modify CAG Instability in Huntington's Disease Mice: Genome-Wide and Candidate Approaches
- The Histone H3 K27 Methyltransferase KMT6 Regulates Development and Expression of Secondary Metabolite Gene Clusters
- Hsp70-Hsp40 Chaperone Complex Functions in Controlling Polarized Growth by Repressing Hsf1-Driven Heat Stress-Associated Transcription
- Function and Evolution of DNA Methylation in
- Stimulation of mTORC1 with L-leucine Rescues Defects Associated with Roberts Syndrome
- Transcription Termination and Chimeric RNA Formation Controlled by FPA
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- A GDF5 Point Mutation Strikes Twice - Causing BDA1 and SYNS2
- Dominant Mutations in Identify the Mlh1-Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway
- Eleven Candidate Susceptibility Genes for Common Familial Colorectal Cancer
- The Histone H3 K27 Methyltransferase KMT6 Regulates Development and Expression of Secondary Metabolite Gene Clusters
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



