-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Functional Diversity of Carbohydrate-Active Enzymes Enabling a Bacterium to Ferment Plant Biomass
Plant-fermenting bacteria are important for the global carbon cycle, human nutrition, and industrial production of renewable fuels and commodities from cellulosic biomass. Plants are primarily composed of heterogeneous polysaccharides, requiring plant-degrading microbes to encode many carbohydrate-active enzymes (CAZymes) to cleave different sugar linkages. Here we develop a broadly applicable method to study how microbes catabolize plant biomass by determining the combination of CAZymes that depolymerize each polysaccharide into sugars, how the cell alters global mRNA expression, and the efficiency with which each polysaccharide is metabolized. We apply this method to investigate how Clostridium phytofermentans, a bacterium encoding 171 CAZymes, ferments polysaccharides. We assimilate our results into a genetic model of how this bacterium metabolizes plant biomass and discuss how these results further our understanding of microbial plant fermentation.
Published in the journal: . PLoS Genet 10(11): e32767. doi:10.1371/journal.pgen.1004773
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004773Summary
Plant-fermenting bacteria are important for the global carbon cycle, human nutrition, and industrial production of renewable fuels and commodities from cellulosic biomass. Plants are primarily composed of heterogeneous polysaccharides, requiring plant-degrading microbes to encode many carbohydrate-active enzymes (CAZymes) to cleave different sugar linkages. Here we develop a broadly applicable method to study how microbes catabolize plant biomass by determining the combination of CAZymes that depolymerize each polysaccharide into sugars, how the cell alters global mRNA expression, and the efficiency with which each polysaccharide is metabolized. We apply this method to investigate how Clostridium phytofermentans, a bacterium encoding 171 CAZymes, ferments polysaccharides. We assimilate our results into a genetic model of how this bacterium metabolizes plant biomass and discuss how these results further our understanding of microbial plant fermentation.
Introduction
Plants annually produce 200 billion tons of lignocellulosic biomass [1], which is metabolized by specialized microbes in diverse environments. For instance, recycling of plant biomass by soil [2] and marine [3] microbes is a key part of the global carbon cycle and intestinal bacteria ferment indigestible plant fiber to short chain fatty acids that constitute 60–85% of calories in ruminants and 5–10% in humans [4]. Further, as only 2% of cellulosic biomass is currently used by humans [5], it is a vast potential feedstock that industrial microbes could convert into energy and commodities. Elucidating how microbes depolymerize and metabolize plant biomass is thus important to understand carbon flow in the environment, to promote healthy human nutrition and prevent disease [6], and to develop industrial processes based on cellulosic bioconversion.
Most of plant biomass is in the cell wall, a macromolecular network of phenolic lignin and three types of polysaccharides (cellulose, hemicelluloses, and pectins) whose relative abundances vary widely among species and tissues (Table S1). The load bearing structure of the cell wall consists of cellulose fibrils tethered by various types of hemicellulose. Hemicellulose is enriched in xylan [7] and xyloglucan [8] in dicots, arabinoxylan in monocots [9], and galacto - and glucomannans in gymnosperms [10]. Outside the cell wall, mannans also act as storage polysaccharides in seeds [11], similar to starch. Pectins are cross-linked galacturonic acid-based polysaccharides that act in cellular adhesion and primary wall extension. More than 60% of pectin is often homogalacturonan (HG) [12], which is esterified with methanol to various degrees. Rhamnogalacturonan I (RGI) [13], the second most abundant pectin, can have galactan and arabinan side chains on the rhamnose residues [14]. Because plant tissues are composed of such heterogeneous polysaccharides, plant-degrading microbes express a myriad of carbohydrate-active enzymes (CAZymes) [15], each of which modifies or cleaves a specific type of sugar linkage.
Here we demonstrate a strategy for systematic analysis of the enzymatic machinery used by microbes to degrade and metabolize plant polysaccharides. Among these microbes, the plant-fermenting clostridia are of particular interest for being a dominant group in the human gut microbiome [6] and top candidates to transform cellulosic biomass into fuels and commodities [16], [17]. We studied Clostridium phytofermentans [18], a soil bacterium with 171 CAZyme-encoding genes (Table S2) including 116 glycoside hydrolases in 44 different families. We first quantified growth on comprehensive panel of plant polysaccharides and sugars (Table S3). Strand-specific RNA sequencing revealed all genes whose expression changed on the various substrates. In particular, we focused on up-regulated CAZyme genes and determined how they are organized into regulons that respond to specific polysaccharides. A set of 56 up-regulated CAZymes were cloned, purified, and an “each enzyme versus each substrate” screen quantified their abilities to bind and cleave plant polysaccharides. These enzymes were then tested in combination to identify synergies for polysaccharide degradation. We discuss how the results can be integrated to further our knowledge of how microbes metabolize plant biomass.
Results/Discussion
Growth on polysaccharides and sugars
We developed a high resolution, microtiter anaerobic growth assay that shows C. phytofermentans ferments diverse plant polysaccharides (Fig. 1) and their constituent monosaccharides (Fig. S1), but with widely varying cell yields and growth rates (Table S4). It also forms colonies on solid medium containing each polysaccharide except arabinogalactan II (AGII) (Fig. S2). Growth was fastest on HG (Fig. 1A, generation time 0.70h), similar to rumen microbes that digest pectin more rapidly than cellulose and hemicellulose [19]. Although C. phytofermentans ferments both galacturonic acid (Fig. S1F) and rhamnose (Fig. S1H), cell yield was low on RGI (Fig. 1B). C. phytofermentans grows well on galactan (Fig. 1C), xylans (Fig. 1F–G), mannans (Fig. 1H–I), xyloglucan (Fig. 1J), and starch (Fig. 1L). Limited growth on AGII (Fig. 1E) relative to galactan supports that C. phytofermentans cleaves β-1,4 galactan, but not the β-1,3 and β-1,6-galactose bonds in AGII. Poor growth on arabinan (Fig. 1D) is similar to arabinose (Fig. S1G), suggesting this sugar is transported or metabolized inefficiently. C. phytofermentans grows well on cellulose plates (Fig. S2) and solubilizes cellulosic substrates such as filter paper and raw corn stover (Fig. S3), but weak growth on carboxymethylcellulose (CMC) might result from either lack of a suitable endoglucanase or carboxymethyl side groups inhibiting its metabolism.
Fig. 1. C. phytofermentans growth on pectic A–E, hemicellulosic F–J, and glucan K–L. 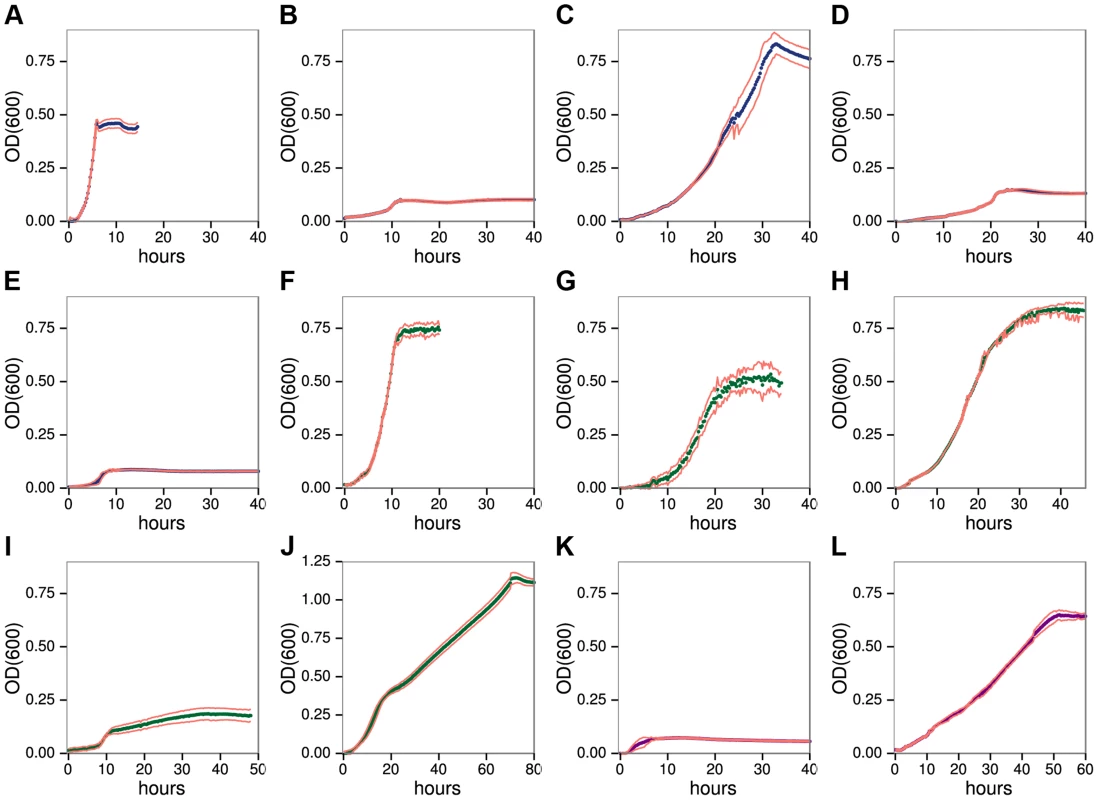
Polysaccharides: homogalacturonan A, rhamnogalacturonan I B, galactan C, arabinan D, arabinogalactan II E, xylan F, arabinoxylan G, glucomannan H, galactomannan I, xyloglucan J, carboxymethylcellulose K, starch L. Growth was measured as OD600 every 15 minutes. Each point is the mean of six cultures; red lines show one standard deviation. C. phytofermentans shows diauxic growth on the mixed sugar polysaccharides galactomannan (Fig. 1I) and xyloglucan (Fig. 1J). For each of these substrates, one of the component sugars (galactose or glucose) supports faster growth than the other (mannose or xylose) (Fig. S1, Table S4). Growth on various mixtures of galactose/mannose (Fig. S4) and of glucose/xylose (Fig. S5) shows rapid metabolism of the preferred sugar followed by slower growth on the other one. However, in both cases when the favored sugar reached 75% of the total, the other sugar does not appear to be metabolized.
Similar to some ruminal [19] and human gut microbes [20], C. phytofermentans often grows faster on polysaccharides than the constituent sugars (Table S4). When presented with mixtures of xylan and xylose, this bacterium shows diauxic growth with preferential metabolism of xylan (Fig. S6), which is surprising because xylan must be cleaved to xylose to be metabolized. Growth on polysaccharides could be energetically favorable if significant ATP is saved by simultaneous transport of multiple sugar units in a single oligosaccharide [21] or by intracellular phosphorolysis of oligosaccharides [22]. C. phytofermentans encodes at least a dozen phosphorylases [23] [24], which cleave oligosaccharides without using ATP. Although the mechanisms regulating sugar metabolism in C. phytofermentans are unknown, diauxic growth supports carbon catabolite repression prioritizes growth on preferred sugars and polysaccharides.
Gene expression
We quantified mRNA expression by strand-specific RNA sequencing during log-phase growth on 8 polysaccharides, 3 monosaccharides, and raw corn stover as a complex biomass substrate. An average of 17.3 million mRNA reads were mapped per sample (Table S5), yielding expression (RPKM) values (Table S6) that were highly correlated (r2 = 0.96–0.99) between duplicate cultures for all conditions (Fig. S7). The reads were also highly strand-specific (Fig. S8), which will facilitate their future use for de novo transcriptome assembly, gene annotation and detection of antisense transcription. The fraction of reads mapping to CAZymes during growth on glucose was 2.0%, but this increased greatly on polysaccharides, especially cellulose (11.9%) and stover (31.0%). We assessed which genes were significantly differentially expressed on each polysaccharide relative to glucose using DESeq [25] (Table S7). Expression of CAZyme genes on polysaccharides relative to glucose (Fig. 2) shows that between 15 (cellobiose) and 40 (stover) CAZymes were significantly up-regulated per treatment (Table S8) with a total of 92 CAZymes up-regulated on at least one polysaccharide.
Fig. 2. mRNA expression of all 171 CAZymes during growth on pectins A–C, hemicelluloses D–E, glucans F–H, and raw corn stover I relative to expression on glucose. 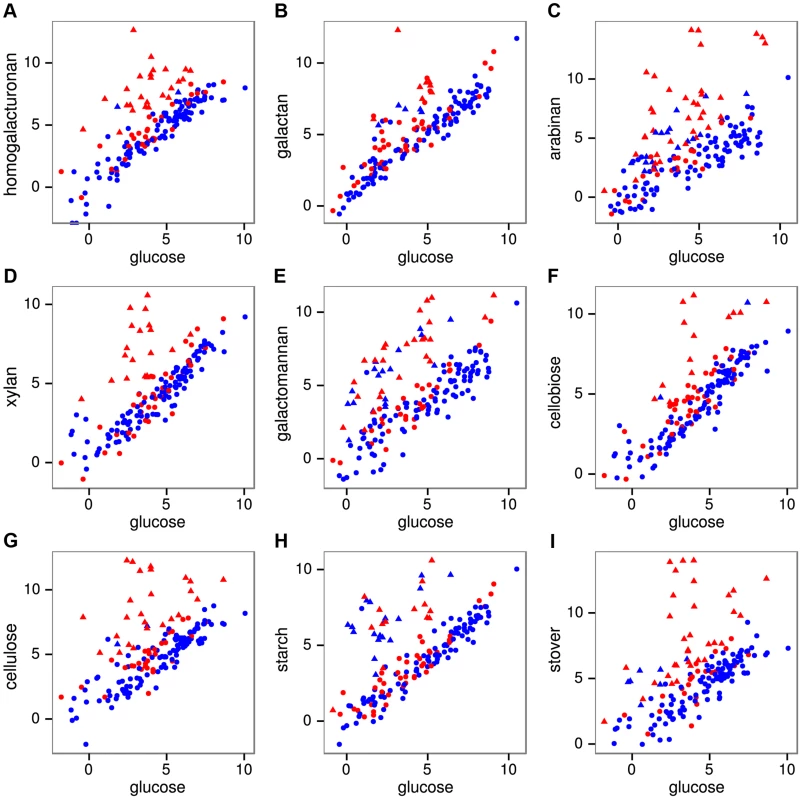
Expression was quantified as log2(RPKM) with significantly differentially expressed genes on a given polysaccharide shown as triangles and unchanged genes as circles. The 56 purified CAZymes are red and others are blue. The differentially expressed CAZymes are putatively classified by the CAZy database as 67 glycoside hydrolases, 6 carbohydrate esterases, 4 polysaccharide lyases, 14 glycosyl transferases, and 2 CBM proteins. We analyzed the specificity of the CAZyme transcriptional response by K-means clustering the expression profiles of these genes (Fig. 3, Table S9). Cluster A consists six genes that were highly up-regulated on multiple substrates: the GH26 cphy1071, the GH11 cphy2105, two GH18 chitinases cphy1799 and cphy1800 [26], the GH9 cellulase cphy3367 [27] [28] and the GH48 cellulase cphy3368 [29]. Clusters B–F respond to specific polysaccharides such as homogalacturonan (clusters B,C), starch (cluster D), xylan (cluster E), and cellulose/arabinan (cluster F). C. phytofermentans thus perceives signals from individual polysaccharides and responds by up-regulating specific transcriptional regulons that enable it to tailor its complement of CAZymes to the polysaccharide substrate.
Fig. 3. CAZymes clustered based on gene expression patterns (clusters A–I) show that some genes respond to multiple carbon sources while others are substrate-specific. 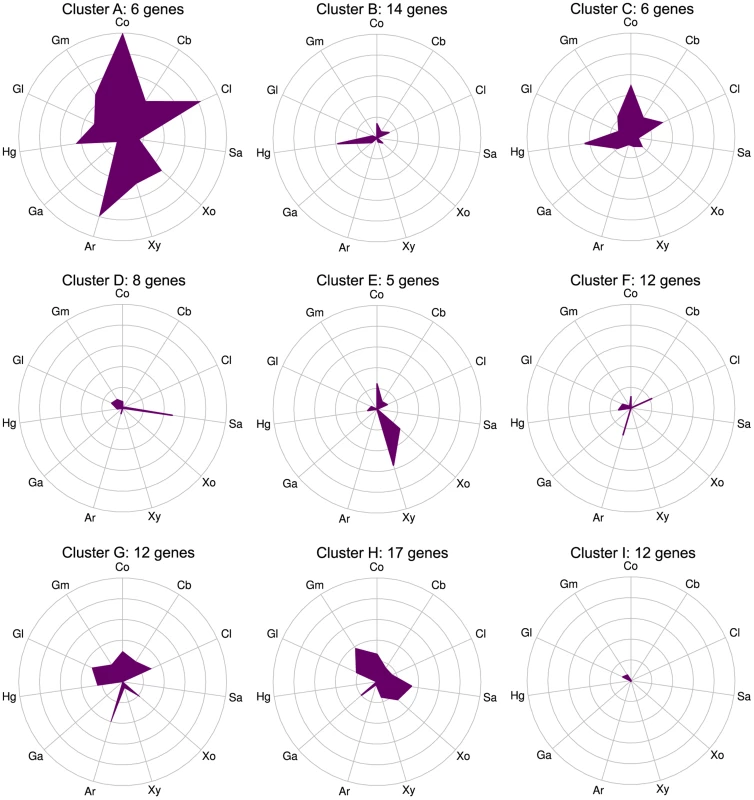
mRNA expression changes (log2 expression ratios relative to glucose) for all 92 CAZyme genes differentially expressed on at least 1 polysaccharide relative to glucose were separated into nine clusters using K-means. Plot centers are expression on glucose and concentric rings show log2 up-regulation on the following carbon sources: cellobiose (Cb), filter paper cellulose (Cl), starch (Sa), xylose (Xo), xylan (Xy), arabinan (Ar), galacturonic acid (Ga), homogalacturonan (Hg), galactan (Gl), galactomannan (Gm), raw corn stover (Co). Gene membership of clusters is shown in Table S9. CAZyme activities
A set of 56 CAZymes up-regulated on polysaccharides were His-tagged, overexpressed, purified, and their abilities to bind and cleave polysaccharides were quantified. The CAZy database classifies these enzymes putatively as 47 glycoside hydrolases, 4 polysaccharide lyases, and 4 carbohydrate esterases (Table S2); putative glycosyltransferases were not examined as they are not involved in polysaccharide catabolism [6]. Thirty-two enzymes have significant cleavage or binding activities (Fig. 4, Table S11). Some substrates such as β-1,4-galactan appear to be cut by a single, highly active enzyme, while multiple CAZymes from the same family degrade other substrates such as xylan (GH10), mannan (GH26), starch (GH13), and HG (PL9). CAZymes from multiple families together depolymerize substrates such as xyloglucan (GH2,5,12,31), glucomannan (GH5,GH9,GH26) and galactomannan (GH5,GH26).
Fig. 4. Cleavage A, binding B, and CAZy database classification C of purified enzymes. 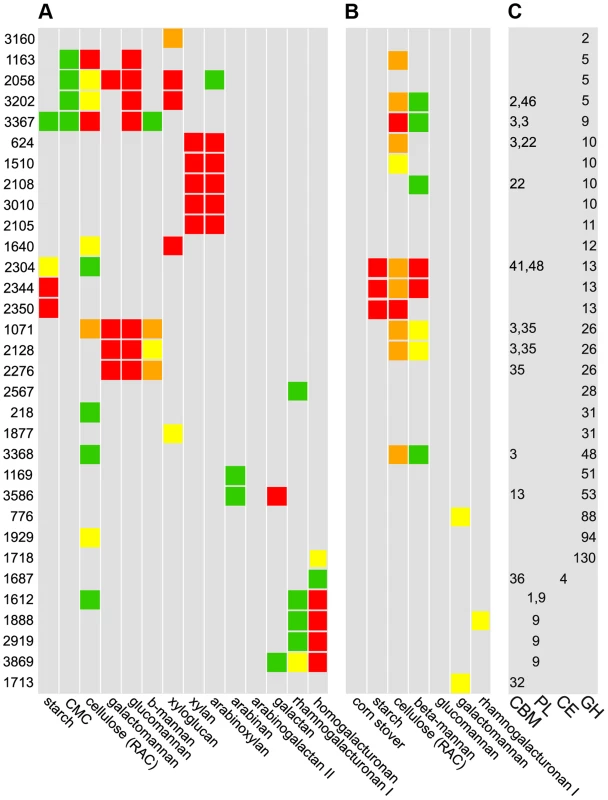
A Polysaccharide cleavage was quantified as nmol reducing sugar released per milligram enzyme per minute: >160 red, 80–160 orange, 40–80 yellow, 20–40 green, <20 gray. B Binding to insoluble polysaccharides was quantified as percentage enzyme bound to substrate: >30% red, 20–30% orange, 15–20% yellow, 10–15% green, <10% gray. C CAZy database classifications: glycoside hydrolases (GH), carbohydrate esterases (CE), polysaccharide lyases (PL), and carbohydrate binding domains (CBM). Among 56 purified CAZymes, only the 32 enzymes for which activities were found are shown. We found 15 CAZymes that bind insoluble polysaccharides, most commonly cellulose and mannan (Fig. 4B). Unexpectedly, no CAZyme bound corn stover, suggesting that partial digestion of raw biomass is needed to facilitate enzyme binding. Nine enzymes that bound substrates have carbohydrate binding modules (CBM), but some enzymes such as the cellulase Cphy1163 can bind their substrate without one. While CBM are known to discriminate between polysaccharides such as cellulose and mannan [30], we observed overlap with cellulase CBMs binding mannan and vice versa. Further, CBM from xylanases can bind cellulose and mannose, but with lower affinity, showing that CBM often bind a range of polysaccharides. Consistent with their cleavage activities, GH13 were the only enzymes to bind starch. Enzymes with CBM usually also have catalytic modules, but Cphy1713, a CAZyme with a CBM32 and no catalytic module, binds galactomannan. CBM32 are known to bind galactose and this protein may function similar to one in Yersinia that is proposed to bind oligosaccharides to prevent them from leaking out of the cell [31].
Thirty-two CAZy families have multiple members, which often have divergent cleavage activities and cellular localizations. Cphy1510 has the highest activity among the four GH10 active on xylan (Fig. 5A). Cphy3010, the GH10 with lowest activity, is the only one lacking a secretion signal, supporting it acts intracellularly on xylo-oligosaccharides while the other GH10 are extracellular. Members of the GH5 family act on a wide range of polysaccharides [32]. C. phytofermentans encodes 3 GH5 enzymes, among which one is active on galactomannan and two on xyloglucan (Fig. 5B). The GH5 Cphy1163 has no activity on either of these substrates, but is the most active on cellulose and glucomannan. The 3 GH26 also vary in substrate specificities (Fig. 5C); all the GH26 are similarly active on β-mannan, but only Cphy1071 has cellulase activity and it has lower activity on gluco - and galactomannan. Sequenced-based families are thus useful to make general substrate predictions for CAZymes, but experiments are needed to determine substrate range and catalytic efficiency.
Fig. 5. Members of the same CAZy family vary in polysaccharide cleavage activities and CAZymes can by potentiated by other enzymes. 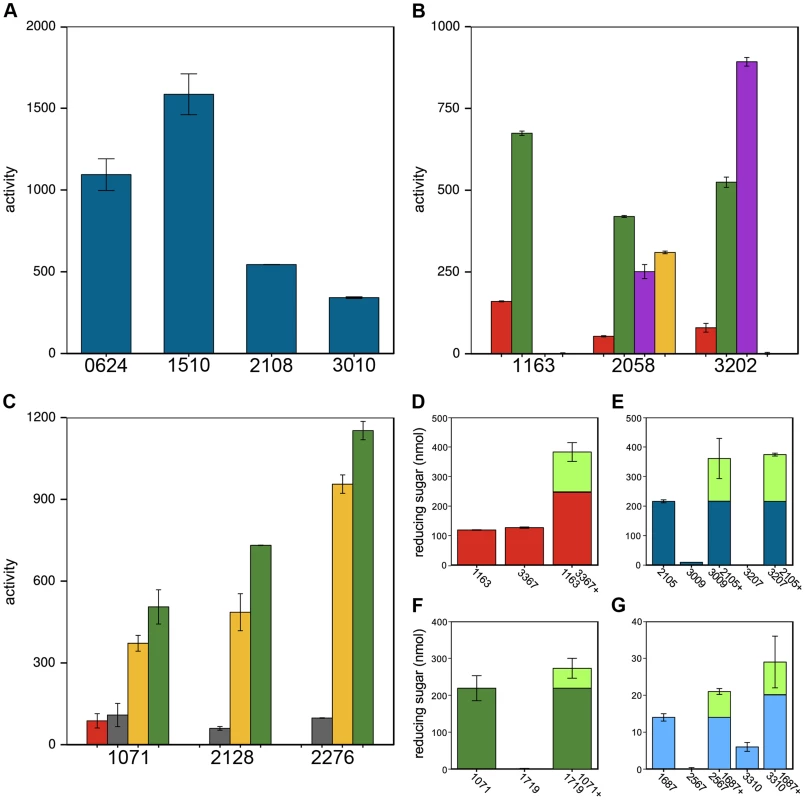
A Variation in cleavage activities of GH10 enzymes on xylan. B GH5 and C GH26 family members differ in their activities and substrate specificities on amorphous cellulose (red), glucomannan (green), xyloglucan (violet), galactomannan (yellow), mannan (gray). Enzyme activities in A–C are nmol reducing sugar released per milligram enzyme per minute. D–G CAZyme mixtures have higher activities than the individual enzymes. D Cphy1163 and Cphy3367 alone and together on amorphous cellulose. E Cphy2105, Cphy3009, and Cphy3207 alone and the latter two enzymes plus Cphy2105 on xylan. F Cphy1719 and Cphy1071 alone and together on glucomannan. G Cphy1687, Cphy2567, and Cphy3310 alone and the latter two enzymes plus Cphy1687 on homogalacturonan. In D–G, enzyme activities are shown as reducing sugar (nmol) produced by individual and combined enzymes. The fraction of the reducing sugar produced by the mixed enzymes that exceeds the sum of the individual enzymes is shown in green. CAZymes mixtures can degrade polysaccharides more efficiently than individual enzymes. We assessed pairwise interactions between each CAZyme and a second enzyme on cellulose (Cphy3367), xylan (Cphy2105), glucomannan (Cphy1071), and homogalacturonan (Cphy1687) (Fig. 5D–G). Similar to results showing synergy between the GH9 Cphy3367 and a B. subtilis GH5 [33], we found that a mix of Cphy3367 and the GH5 Cphy1163 has higher activity on cellulose than either enzyme alone (Fig. 5D), supporting they have complementary roles in cellulolysis. CAZymes can also potentiate other enzymes that have no activity by themselves. For example, the xylanase Cphy2105 activates the putative xylosidases Cphy3009 and Cphy3207 on xylan (Fig. 5E). Similarly, Cphy1071 activates the putative mannosidase Cphy1719 on glucomannan (Fig. 5F). Activities of the GH28 Cphy2567 and Cphy3310 are enhanced by the carbohydrate esterase Cphy1687 (Fig. 5G), supporting this enzyme demethylesterifies homogalacturonan to facilitate its degradation. This carbohydrate esterase did not, however, increase cleavage by the PL9 enzymes that were the most active on homogalacturonan.
Global correlations between CAZyme mRNA expression and cleavage activities were weak for all polysaccharides (Fig. S12), mostly because many CAZyme genes are up-regulated on substrates upon which they have no activity. CAZymes up-regulated on multiple substrates (Fig. 3, cluster A) may act as ‘carbon scouts’ [34] that degrade complex substrates into inducing molecules used to fine-tune the expression of hydrolytic enzymes. As described above, some CAZymes such as xylosidases (Fig. 5D) are inactive on intact xylan, but are potentiated by other xylanases. The GH18 Cphy1799 and Cphy1800 are the most highly-upregulated CAZymes on cellulose (Fig. S12), but are chitinases with no activity on cellulose or other plant substrates [26]. As such, the set of up-regulated CAZymes is useful to identify active enzymes, but strong up-regulation does not necessarily indicate activity on a given substrate.
Conclusions
We assimilated our results into a model of C. phytofermentans polysaccharide catabolism that shows degradation by active CAZymes and uses mRNA expression profiles to predict how these substrates are transported and metabolized (Fig. 6). Unlike other clostridia that transport sugars with numerous phosphotransferase systems (PTS) [35] [36], C. phytofermentans encodes a single, lowly expressed PTS and also lacks the symporters to transport xylose and arabinose [37]. Instead, C. phytofermentans responds to carbon sources by up-regulating between two (galacturonic acid) and twenty-two (arabinan) ABC transporters (Fig. 6). Expression changes support that oligosaccharides and monosaccharides are uptaken by distinct transporters. For example, different ABC transporters are up-regulated on xylose and xylan. Similarly, different transporters respond to glucose, cellobiose, and cellulose. Intracellular cellodextrins are cleaved by at least one cellodextin phosphorylase (GH94); hexoses are phosphorylated, likely by a ROK hexokinase (Cphy0329) and a putative galactokinase (Cphy2237), and fed into glycolysis. While hexokinases may have wide substrate activity [38], poor growth on mannose could be due to inefficient mannose phosphorylation. The pentoses xylose and arabinose are isomerized and metabolized by the pentose phosphate pathway (PPP). Weak growth on arabinose could be due to inefficient transport or the lack of the phosphoketolase in the PPP enabling rapid L-arabinose metabolism by C. acetobutylicum [39].
Fig. 6. Model of polysaccharide degradation and metabolism by C. phytofermentans. CAZymes (shown as the number of enzymes in CAZy families) are based on purified activities and are intra- or extracellular based on putative secretion signals. 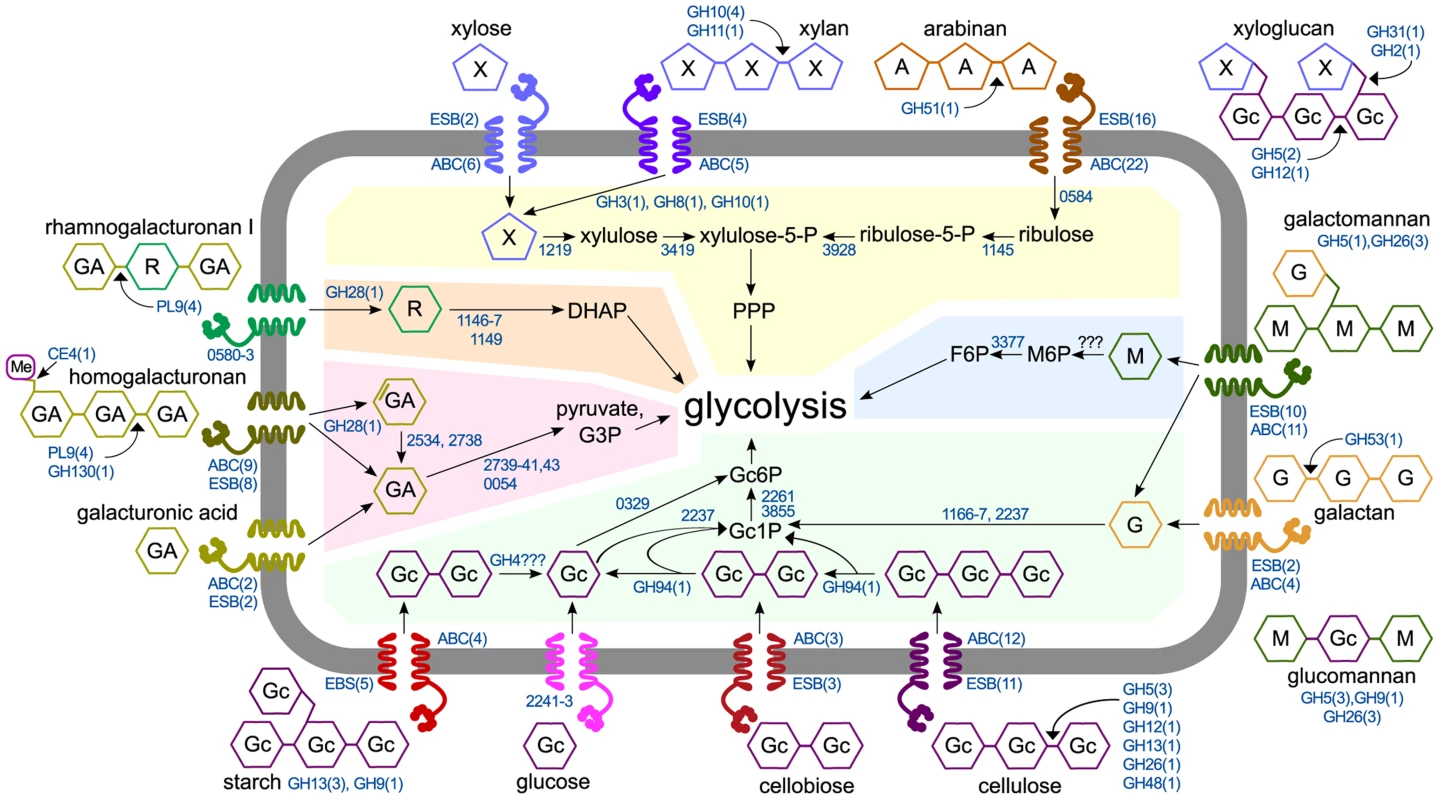
Metabolic enzymes are shown as NCBI numbers and are proposed based on mRNA expression. Rhamnose transport and assimilation is based on pathway from [55]. Abbreviations are D-galacturonic acid (GA), L-rhamnose (R), D-mannose (M), D-glucose (Gc), D-galactose (G), D-xylose (X), L-arabinose (A), fructose (F), phosphate (P), pentose phosphate pathway (PPP), dihydroxyacetone-phosphate (DHAP), glyceraldehyde-3-phosphate (G3P). For each substrate, the number of significantly up-regulated extracellular solute binding proteins (ESB) and ABC transporters (ABC) are shown. Shaded regions show metabolism of glucose (green), mannose (blue), xylose and arabinose (yellow), rhamnose (orange), and galacturonic acid (red). Plant degrading microbes differ widely in their abilities to depolymerize and metabolize polysaccharides, likely reflecting niche differentiation to alleviate resource competition. Among soil clostridia, C. thermocellum ferments cellulose, but not xylan [40]. C. cellulolyticum grows faster on xylose than xylan and faster on cellobiose than glucose [41], both of which differ from C. phytofermentans. Similar specialization exists in the human gut microbiome where microbes catabolize different glycans in dietary fiber [20]. The strategy presented here of high-resolution anaerobic growth measurements, RNA sequencing, and CAZyme assays complements other methods such as proteomics [42] and metagenomics. Elucidating how microbes metabolize polysaccharides is key to understanding the function of plant-degrading microbial communities and to develop improved enzyme mixtures and recombinant microbes for industrial processing of plant biomass.
Materials and Methods
Growth measurements
C. phytofermentans was cultured anaerobically at 30°C in GS2 medium [43]. Growth was quantified in medium containing 3 g l−1 mono-, di-, or polysaccharides (Table S3 for product numbers) in 100-well microtiter plates (Bioscreen 9502550) that were sealed by press-fitting adhesive sheets (Qiagen 1018104) under the lids. Growth was measured every 15 minutes as OD600 at 30°C using a Thermo Scientific Bioscreen C with 30 seconds shaking before each reading. Growth was not measured for ß-mannan and amorphous cellulose cultures because these low solubility substrates occluded optical density measurements. Growth on insoluble substrates (15 g l−1 of 0.5×5 cm strips of filter paper cellulose or raw corn stover) was measured in 10 ml cultures as substrate consumption by collection on 11 µm filters by vacuum filtration, drying overnight at 65°C, and weighing. Growth on solid GS2 medium with 3 g l−1 polysaccharide and 15 g l−1 agar was tested by incubating plates anaerobically for 10 days at 30°C (Fig. S2). Regenerated amorphous cellulose (RAC) for GS2 agar plates and enzyme assays was prepared from Avicel PH-101 by phosphoric acid treatment [44].
RNA-seq
Cultures containing soluble substrates were sampled for RNA-seq in mid-log phase. Filter paper cellulose cultures were sampled under the same conditions as in [45]; corn stover cultures were sampled after 3 days. Cells were arrested with RNA stabilization buffer, collected by centrifugation (5 kg, 5 min, 4°C), and RNA was isolated with an Ambion Ribopure Bacteria Kit (AM1925). Twenty µg total RNA was treated with Turbo DNase (Invitrogen AM1907), phenol-chloroform extracted, ethanol precipitated, and resuspended in 15 µl 10 mM Tris-Cl, pH 8.5. Ribosomal RNA was depleted from 10 µg RNA using a MICROBExpress kit (Invitrogen AM1905), giving a typical yield of 1–2 µg RNA. 500 ng of rRNA-depleted RNA was fragmented with magnesium buffer (NEB E6101) for 2.5 minutes at 94°C, ethanol precipitated, and resuspended in 10 µl water. RNA was resolved on an Agilent Bioanalyzer 2100 to confirm it was 200–800 bp.
Single stranded cDNA was made from 500 ng fragmented RNA by Superscript III first strand synthesis (Invitrogen 18080-051) including 200 ng/µl actinomycin D (Invitrogen 11805017) and 120 ng/µl random hexamers (Invitrogen 48190-011). Single stranded DNA was twice phenol-chloroform extracted, ethanol precipitated, and resuspended in 52 µl water. The second cDNA strand was synthesized by the Invitrogen double stranded cDNA synthesis kit (Invitrogen 11917-010) using 250 µM dNTP (dATP, dCTP, dGTP, dUTP). Double stranded cDNA was purified using 1.8 volumes of Solid Phase Reversible Immobilisation (SPRI) beads [46] prepared from carboxylate-modified microparticles [47] (ThermoScientific 6515-2105-050250) and resuspended in 50 µl 10 mM Tris-Cl, pH 8.5. DNA was resolved on an Agilent Bioanalyzer to confirm cDNA was 100–500 bp.
Sequencing libraries were prepared from ∼250 ng cDNA using the Illumina Truseq DNA Kit (Illumina FC-121-2001). DNA was purified with 1.8 volumes of SPRI beads after end repair. Following adapter ligation, 200–500 bp DNA was selected by Pippin Prep gel extraction (Sage Science). The second cDNA strand (contains dUTP) was then degraded by incubating with 1 unit USER enzyme (NEB NEB M5505S) at 37°C for 15 minutes, followed by 5 minutes at 95°C. Single stranded DNA was PCR amplified and size selected with 1.8 volumes of SPRI beads. Multiplexed libraries were normalized to 2 ng µl−1, pooled, and resolved on an Agilent Bioanalyzer to confirm the peak was at 200–500 bp. Libraries were sequenced on a Illumina HiSeq2000 sequencer. RNA-seq reads were mapped to the C. phytofermentans genome (NCBI NC_010001.1) using Bowtie [48] to report the single, best alignment (see Table S5 for parameters). Gene expression was measured as RPKM (sum of nucleotides in gene per million mapped nucleotides per gene kb) using RSEQtools [49] (Table S6), excluding rRNA reads from RPKM calculations. Differentially expressed genes (Tables S7, S8) were identified by DESeq [25] using the statistical thresholds defined in [50] of >4-fold expression and a p-value<10−5, which corresponds to a p-value<0.05 adjusted for multiple testing of the 3902 genes in C. phytofermentans genome by Bonferroni correction.
Enzyme purification and activities
A set of 56 CAZymes (Table S2) that were up-regulated on plant polysaccharides were His-tagged, overexpressed, and purified. Primers (Table S10) were designed to clone CAZyme genes by Ligation-Independent Cloning [51] into pET-22B(+). Genes were cloned with C-terminal His-tags except for cphy1687 and cphy2105, which were cloned with N-terminal tags to improve yields. The 26 enzymes predicted to be secreted by SignalP [52] were cloned as their mature forms, omitting the N-terminal secretion signals. Forward and reverse sequences of the 56 genes cloned in pET-22B(+) were confirmed by sequencing (Fig. S9). Plasmids were transformed into E. coli BL21(DE3) (Novagen 70235), grown in 50 ml TB medium to OD600 = 1.0, and induced by adding 500 µM IPTG and incubating overnight at 20°C. Cells were pelleted, resuspended in lysis buffer: 50 mM phosphate buffer pH = 8, 0.5M NaCl, 10 mM imidazole, 15% glycerol, 1 mM pefabloc (Sigma 76307). Cells were lysed by sonication (Cole-Parmer Vibracell CV33) in the presence of lysozyme (Novagen 71230). His-tagged proteins were purified from 50 ml culture on Ni-NTA spin columns (Qiagen 31014) and quantified by Bradford assay, giving an average yield of 3 mg protein. Purified proteins were visualized on 12% SDS-PAGE gels (Novex 12% Bis-Tris Gel NP0342BOX) (Fig. S10).
Enzymatic polysaccharide cleavage was quantified by 3,5-dinitrosalicylic acid (DNS) assay [53] in 96 well clear, flat bottom plates (Molecular Devices X6011). Fifty µg enzyme was incubated with 0.25% polysaccharide in 25 mM potassium phosphate buffer pH = 7 for 30 minutes at 37°C, 1 volume DNS reagent was added, incubated at 95°C for 15 minutes, and read at OD600. Enzyme activities (nmol reducing sugar per mg enzyme per min) were calculated from DNS readings by subtracting blanks (+polysaccharide,-enzyme) and calculating sugar produced using standard curves (Fig. S11). Polysaccharide cleavage by enzyme pairs was tested on amorphous cellulose, xylan, glucomannan, and homogalacturonan as the reducing sugar (mM) produced by 25 µg of each enzyme alone and combined with a second enzyme (25 µg Cphy3367 for cellulose, 5 µg Cphy2105 for xylan, 5 µg Cphy1071 for glucomannan, 25 µg Cphy1687 for homogalacturonan). Reducing sugar was measured by DNS assay after 30 minutes incubation as described above. Enzyme binding to insoluble polysaccharides was quantified using a method adapted from [54]. Fifty µg enzyme was suspended in 200 µl 0.1 M phosphate buffer pH = 7 with 6 mg polysaccharide and incubated with rotation for 5 h at 4°C. The polysaccharide was collected by centrifuging at 12 kg for 5 min and the enzyme concentration in the supernatant was calculated before and after incubation by Bradford assay, yielding the percentage of enzyme bound to the polysaccharide.
Supporting Information
Zdroje
1. ShekharC (2011) Future fuel: could biomass be the new petroleum? Chem Biol 18 : 1199–1200 doi:10.1016/j.chembiol.2011.10.010
2. SchwarzWH (2001) The cellulosome and cellulose degradation by anaerobic bacteria. Appl Microbiol Biotechnol 56 : 634–649 doi:10.1007/s002530100710
3. EdwardsJL, SmithDL, ConnollyJ, McDonaldJE, CoxMJ, et al. (2010) Identification of carbohydrate metabolism genes in the metagenome of a marine biofilm community shown to be dominated by gammaproteobacteria and bacteroidetes. Genes 1 : 371–384 doi:10.3390/genes1030371
4. McNeilNI (1984) The contribution of the large intestine to energy supplies in man. Am J Clin Nutr 39 : 338–342.
5. PaulyM, KeegstraK (2008) Cell-wall carbohydrates and their modification as a resource for biofuels. Plant J Cell Mol Biol 54 : 559–568 doi:10.1111/j.1365-313X.2008.03463.x
6. El KaoutariA, ArmougomF, GordonJI, RaoultD, HenrissatB (2013) The abundance and variety of carbohydrate-active enzymes in the human gut microbiota. Nat Rev Microbiol 11 : 497–504 doi:10.1038/nrmicro3050
7. DeutschmannR, DekkerRFH (2012) From plant biomass to bio-based chemicals: latest developments in xylan research. Biotechnol Adv 30 : 1627–1640 doi:10.1016/j.biotechadv.2012.07.001
8. BauerWD, TalmadgeKW, KeegstraK, AlbersheimP (1973) The Structure of Plant Cell Walls: II. The Hemicellulose of the Walls of Suspension-cultured Sycamore Cells. Plant Physiol 51 : 174–187 doi:http://dx.doi.org/10.1104/pp.51.1.174
9. BurkeD, KaufmanP, McNeilM, AlbersheimP (1974) The Structure of Plant Cell Walls: VI. A Survey of the Walls of Suspension-cultured Monocots. Plant Physiol 54 : 109–115 doi:http://dx.doi.org/10.1104/pp.54.1.109
10. CapekP, AlföldiJ, LiskováD (2002) An acetylated galactoglucomannan from Picea abies L. Karst. Carbohydr Res 337 : 1033–1037 doi:10.1016/S0008-6215(02)00090-3
11. BuckeridgeMS (2010) Seed cell wall storage polysaccharides: models to understand cell wall biosynthesis and degradation. Plant Physiol 154 : 1017–1023 doi:10.1104/pp.110.158642
12. CaffallKH, MohnenD (2009) The structure, function, and biosynthesis of plant cell wall pectic polysaccharides. Carbohydr Res 344 : 1879–1900 doi:10.1016/j.carres.2009.05.021
13. RidleyBL, O'NeillMA, MohnenD (2001) Pectins: structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 57 : 929–967 doi:10.1016/S0031-9422(01)00113-3
14. MohnenD (2008) Pectin structure and biosynthesis. Curr Opin Plant Biol 11 : 266–277 doi:10.1016/j.pbi.2008.03.006
15. LombardV, Golaconda RamuluH, DrulaE, CoutinhoPM, HenrissatB (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42: D490–495 doi:10.1093/nar/gkt1178
16. LyndLR, WeimerPJ, van ZylWH, PretoriusIS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol Rev MMBR 66 : 506–577 table of contents. doi:10.1128/MMBR.66.3.506-577.2002
17. Tolonen AC, Petit E, Blanchard JL, Warnick T, Leschine SB (2013) Technologies to Study Plant Biomass Fermentation Using the Model Bacterium Clostridium phytofermentans. In: J. Sun, S.Y. Ding, J.D. Peterson (Eds.), Biological conversion of biomass for fuels and chemicals, Royal Society of Chemistry, Cambridge, pp. 114–139. doi:10.1039/9781849734738-00114
18. WarnickTA, MethéBA, LeschineSB (2002) Clostridium phytofermentans sp. nov., a cellulolytic mesophile from forest soil. Int J Syst Evol Microbiol 52 : 1155–1160 doi:10.1099/ijs.0.02125-0
19. HatfieldRD, WeimerPJ (1995) Degradation characteristics of isolated and in situ cell wall lucerne pectic polysaccharides by mixed ruminal microbes. J Sci Food Agric 69 : 185–196 doi:10.1002/jsfa.2740690208
20. MartensEC, LoweEC, ChiangH, PudloNA, WuM, et al. (2011) Recognition and Degradation of Plant Cell Wall Polysaccharides by Two Human Gut Symbionts. PLoS Biol 9: e1001221 doi:10.1371/journal.pbio.1001221
21. MuirM, WilliamsL, FerenciT (1985) Influence of transport energization on the growth yield of Escherichia coli. J Bacteriol 163 : 1237–1242.
22. ZhangY-HP, LyndLR (2005) Cellulose utilization by Clostridium thermocellum: bioenergetics and hydrolysis product assimilation. Proc Natl Acad Sci U S A 102 : 7321–7325 doi:10.1073/pnas.0408734102
23. NihiraT, NakaiH, ChikuK, KitaokaM (2012) Discovery of nigerose phosphorylase from Clostridium phytofermentans. Appl Microbiol Biotechnol 93 : 1513–1522 doi:10.1007/s00253-011-3515-9
24. NakajimaM, NishimotoM, KitaokaM (2009) Characterization of three beta-galactoside phosphorylases from Clostridium phytofermentans: discovery of d-galactosyl-beta1->4-l-rhamnose phosphorylase. J Biol Chem 284 : 19220–19227 doi:10.1074/jbc.M109.007666
25. AndersS, HuberW (2010) Differential expression analysis for sequence count data. Genome Biol 11: R106 doi:10.1186/gb-2010-11-10-r106
26. TolonenAC, CerisyT, El-SayyedH, BoutardM, SalanoubatM, et al. (2014) Fungal lysis by a soil bacterium fermenting cellulose. Environ Microbiol doi:10.1111/1462-2920.12495
27. TolonenAC, ChilakaAC, ChurchGM (2009) Targeted gene inactivation in Clostridium phytofermentans shows that cellulose degradation requires the family 9 hydrolase Cphy3367. Mol Microbiol 74 : 1300–1313 doi:10.1111/j.1365-2958.2009.06890.x
28. ZhangX-Z, SathitsuksanohN, ZhangY-HP (2010) Glycoside hydrolase family 9 processive endoglucanase from Clostridium phytofermentans: heterologous expression, characterization, and synergy with family 48 cellobiohydrolase. Bioresour Technol 101 : 5534–5538 doi:10.1016/j.biortech.2010.01.152
29. ZhangX-Z, ZhangZ, ZhuZ, SathitsuksanohN, YangY, et al. (2010) The noncellulosomal family 48 cellobiohydrolase from Clostridium phytofermentans ISDg: heterologous expression, characterization, and processivity. Appl Microbiol Biotechnol 86 : 525–533 doi:10.1007/s00253-009-2231-1
30. KnoxJP (2008) Revealing the structural and functional diversity of plant cell walls. Curr Opin Plant Biol 11 : 308–313 doi:10.1016/j.pbi.2008.03.001
31. AbbottDW, HrynuikS, BorastonAB (2007) Identification and characterization of a novel periplasmic polygalacturonic acid binding protein from Yersinia enterolitica. J Mol Biol 367 : 1023–1033 doi:10.1016/j.jmb.2007.01.030
32. AspeborgH, CoutinhoPM, WangY, BrumerH, HenrissatB (2012) Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5). BMC Evol Biol 12 : 186 doi:10.1186/1471-2148-12-186
33. LiaoH, ZhangX-Z, RollinJA, ZhangY-HP (2011) A minimal set of bacterial cellulases for consolidated bioprocessing of lignocellulose. Biotechnol J 6 : 1409–1418 doi:10.1002/biot.201100157
34. BenzJP, ChauBH, ZhengD, BauerS, GlassNL, et al. (2014) A comparative systems analysis of polysaccharide-elicited responses in Neurospora crassa reveals carbon source-specific cellular adaptations. Mol Microbiol 91 : 275–299 doi:10.1111/mmi.12459
35. NöllingJ, BretonG, OmelchenkoMV, MakarovaKS, ZengQ, et al. (2001) Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J Bacteriol 183 : 4823–4838 doi:10.1128/JB.183.16.4823-4838.2001
36. Al MakishahNH, MitchellWJ (2013) Dual substrate specificity of an N-acetylglucosamine phosphotransferase system in Clostridium beijerinckii. Appl Environ Microbiol 79 : 6712–6718 doi:10.1128/AEM.01866-13
37. ServinskyMD, KielJT, DupuyNF, SundCJ (2010) Transcriptional analysis of differential carbohydrate utilization by Clostridium acetobutylicum. Microbiol Read Engl 156 : 3478–3491 doi:10.1099/mic.0.037085-0
38. ChenM, ChenL, ZouY, XueM, LiangM, et al. (2011) Wide sugar substrate specificity of galactokinase from Streptococcus pneumoniae TIGR4. Carbohydr Res 346 : 2421–2425 doi:10.1016/j.carres.2011.08.014
39. ServinskyMD, GermaneKL, LiuS, KielJT, ClarkAM, et al. (2012) Arabinose is metabolized via a phosphoketolase pathway in Clostridium acetobutylicum ATCC 824. J Ind Microbiol Biotechnol 39 : 1859–1867 doi:10.1007/s10295-012-1186-x
40. NgTK, Ben-BassatA, ZeikusJG (1981) Ethanol Production by Thermophilic Bacteria: Fermentation of Cellulosic Substrates by Cocultures of Clostridium thermocellum and Clostridium thermohydrosulfuricum. Appl Environ Microbiol 41 : 1337–1343.
41. XuC, HuangR, TengL, WangD, HemmeCL, et al. (2013) Structure and regulation of the cellulose degradome in Clostridium cellulolyticum. Biotechnol Biofuels 6 : 73 doi:10.1186/1754-6834-6-73
42. TolonenAC, HaasW (2014) Quantitative Proteomics Using Reductive Dimethylation for Stable Isotope Labeling. J Vis Exp (89) e51416 doi:10.3791/51416
43. JohnsonEA, MadiaA, DemainAL (1981) Chemically Defined Minimal Medium for Growth of the Anaerobic Cellulolytic Thermophile Clostridium thermocellum. Appl Environ Microbiol 41 : 1060–1062.
44. HongJ, YeX, WangY, ZhangY-HP (2008) Bioseparation of recombinant cellulose-binding module-proteins by affinity adsorption on an ultra-high-capacity cellulosic adsorbent. Anal Chim Acta 621 : 193–199 doi:10.1016/j.aca.2008.05.041
45. TolonenAC, HaasW, ChilakaAC, AachJ, GygiSP, et al. (2011) Proteome-wide systems analysis of a cellulosic biofuel-producing microbe. Mol Syst Biol 7 : 461 doi:10.1038/msb.2010.116
46. DeAngelisMM, WangDG, HawkinsTL (1995) Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res 23 : 4742–4743 doi:10.1093/nar/23.22.4742
47. RohlandN, ReichD (2012) Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res 22 : 939–946 doi:10.1101/gr.128124.111
48. LangmeadB, TrapnellC, PopM, SalzbergSL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10: R25 doi:10.1186/gb-2009-10-3-r25
49. HabeggerL, SbonerA, GianoulisTA, RozowskyJ, AgarwalA, et al. (2011) RSEQtools: a modular framework to analyze RNA-Seq data using compact, anonymized data summaries. Bioinforma Oxf Engl 27 : 281–283 doi:10.1093/bioinformatics/btq643
50. MandlikA, LivnyJ, RobinsWP, RitchieJM, MekalanosJJ, et al. (2011) RNA-Seq-based monitoring of infection-linked changes in Vibrio cholerae gene expression. Cell Host Microbe 10 : 165–174 doi:10.1016/j.chom.2011.07.007
51. AslanidisC, de JongPJ (1990) Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res 18 : 6069–6074 http://dx.doi.org/10.1093/nar/18.20.6069.
52. PetersenTN, BrunakS, von HeijneG, NielsenH (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8 : 785–786 doi:10.1038/nmeth.1701
53. MillerGL (1959) Use of Dinitrosalicylic Acid Reagent for Determination of Reducing Sugar. Anal Chem 31 : 426–428 doi:10.1021/ac60147a030
54. WatanabeT, ItoY, YamadaT, HashimotoM, SekineS, et al. (1994) The roles of the C-terminal domain and type III domains of chitinase A1 from Bacillus circulans WL-12 in chitin degradation. J Bacteriol 176 : 4465–4472.
55. PetitE, LaToufWG, CoppiMV, WarnickTA, CurrieD, et al. (2013) Involvement of a bacterial microcompartment in the metabolism of fucose and rhamnose by Clostridium phytofermentans. PloS One 8: e54337 doi:10.1371/journal.pone.0054337
56. ZablackisE, HuangJ, MüllerB, DarvillAG, AlbersheimP (1995) Characterization of the cell-wall polysaccharides of Arabidopsis thaliana leaves. Plant Physiol 107 : 1129–1138 doi:http://dx.doi.org/10.1104/pp.107.4.1129
57. PrasadS, SinghA, JoshiHC (2007) Ethanol as an alternative fuel from agricultural, industrial and urban residues. Resour Conserv Recycl 50 : 1–39 doi:10.1016/j.resconrec.2006.05.007
58. ChowPS, LandhäusserSM (2004) A method for routine measurements of total sugar and starch content in woody plant tissues. Tree Physiol 24 : 1129–1136 doi:10.1093/treephys/24.10.1129
59. SchellerHV, UlvskovP (2010) Hemicelluloses. Annu Rev Plant Biol 61 : 263–289 doi:10.1146/annurev-arplant-042809-112315
60. DahalP, NevinsDJ, BradfordKJ (1997) Relationship of Endo-[beta]-D-Mannanase Activity and Cell Wall Hydrolysis in Tomato Endosperm to Germination Rates. Plant Physiol 113 : 1243–1252 doi:http://dx.doi.org/10.1104/pp.113.4.1243
61. PettolinoFA, WalshC, FincherGB, BacicA (2012) Determining the polysaccharide composition of plant cell walls. Nat Protoc 7 : 1590–1607 doi:10.1038/nprot.2012.081
62. Oxenboll SørensenS, PaulyM, BushM, SkjøtM, McCannMC, et al. (2000) Pectin engineering: modification of potato pectin by in vivo expression of an endo-1,4-beta-D-galactanase. Proc Natl Acad Sci U S A 97 : 7639–7644 doi:10.1073/pnas.130568297
63. MooreJP, Nguema-OnaE, ChevalierL, LindseyGG, BrandtWF, et al. (2006) Response of the leaf cell wall to desiccation in the resurrection plant Myrothamnus flabellifolius. Plant Physiol 141 : 651–662 doi:10.1104/pp.106.077701
Štítky
Genetika Reprodukční medicína
Článek The COP9 Signalosome Converts Temporal Hormone Signaling to Spatial Restriction on Neural CompetenceČlánek Coordinate Regulation of Stem Cell Competition by Slit-Robo and JAK-STAT Signaling in the TestisČlánek The CSN/COP9 Signalosome Regulates Synaptonemal Complex Assembly during Meiotic Prophase I ofČlánek GPA: A Statistical Approach to Prioritizing GWAS Results by Integrating Pleiotropy and AnnotationČlánek Heat-Induced Release of Epigenetic Silencing Reveals the Concealed Role of an Imprinted Plant GeneČlánek p53- and ERK7-Dependent Ribosome Surveillance Response Regulates Insulin-Like Peptide SecretionČlánek The Complex I Subunit Selectively Rescues Mutants through a Mechanism Independent of MitophagyČlánek Rad59-Facilitated Acquisition of Y′ Elements by Short Telomeres Delays the Onset of SenescenceČlánek ARTIST: High-Resolution Genome-Wide Assessment of Fitness Using Transposon-Insertion Sequencing
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 11- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- Establishing a Multidisciplinary Context for Modeling 3D Facial Shape from DNA
- RNA Processing Factors Swd2.2 and Sen1 Antagonize RNA Pol III-Dependent Transcription and the Localization of Condensin at Pol III Genes
- Inversion of the Chromosomal Region between Two Mating Type Loci Switches the Mating Type in
- A Thermolabile Aldolase A Mutant Causes Fever-Induced Recurrent Rhabdomyolysis without Hemolytic Anemia
- The Role of Regulatory Evolution in Maize Domestication
- Stress Granule-Defective Mutants Deregulate Stress Responsive Transcripts
- 24-Hour Rhythms of DNA Methylation and Their Relation with Rhythms of RNA Expression in the Human Dorsolateral Prefrontal Cortex
- Pseudoautosomal Region 1 Length Polymorphism in the Human Population
- Fungal Communication Requires the MAK-2 Pathway Elements STE-20 and RAS-2, the NRC-1 Adapter STE-50 and the MAP Kinase Scaffold HAM-5
- The COP9 Signalosome Converts Temporal Hormone Signaling to Spatial Restriction on Neural Competence
- The Protein -glucosyltransferase Rumi Modifies Eyes Shut to Promote Rhabdomere Separation in
- The Talin Head Domain Reinforces Integrin-Mediated Adhesion by Promoting Adhesion Complex Stability and Clustering
- Quantitative Genetics of CTCF Binding Reveal Local Sequence Effects and Different Modes of X-Chromosome Association
- Coordinate Regulation of Stem Cell Competition by Slit-Robo and JAK-STAT Signaling in the Testis
- Genetic Analysis of a Novel Tubulin Mutation That Redirects Synaptic Vesicle Targeting and Causes Neurite Degeneration in
- A Systems Genetics Approach Identifies , , and as Novel Aggressive Prostate Cancer Susceptibility Genes
- Three RNA Binding Proteins Form a Complex to Promote Differentiation of Germline Stem Cell Lineage in
- Approximation to the Distribution of Fitness Effects across Functional Categories in Human Segregating Polymorphisms
- The CSN/COP9 Signalosome Regulates Synaptonemal Complex Assembly during Meiotic Prophase I of
- SAS-1 Is a C2 Domain Protein Critical for Centriole Integrity in
- An RNA-Seq Screen of the Antenna Identifies a Transporter Necessary for Ammonia Detection
- GPA: A Statistical Approach to Prioritizing GWAS Results by Integrating Pleiotropy and Annotation
- Let's Face It—Complex Traits Are Just Not That Simple
- Glutamate Receptor Gene , Coffee, and Parkinson Disease
- The Red Queen Model of Recombination Hotspots Evolution in the Light of Archaic and Modern Human Genomes
- The Ethics of Our Inquiry: An Interview with Hank Greely
- Functional Diversity of Carbohydrate-Active Enzymes Enabling a Bacterium to Ferment Plant Biomass
- Regularized Machine Learning in the Genetic Prediction of Complex Traits
- Phylogenetically Driven Sequencing of Extremely Halophilic Archaea Reveals Strategies for Static and Dynamic Osmo-response
- Lack of Replication of the -by-Coffee Interaction in Parkinson Disease
- Natural Polymorphisms in Human APOBEC3H and HIV-1 Vif Combine in Primary T Lymphocytes to Affect Viral G-to-A Mutation Levels and Infectivity
- A Germline Polymorphism of Thymine DNA Glycosylase Induces Genomic Instability and Cellular Transformation
- Heat-Induced Release of Epigenetic Silencing Reveals the Concealed Role of an Imprinted Plant Gene
- ATPase-Independent Type-III Protein Secretion in
- p53- and ERK7-Dependent Ribosome Surveillance Response Regulates Insulin-Like Peptide Secretion
- The Complex I Subunit Selectively Rescues Mutants through a Mechanism Independent of Mitophagy
- Evolution of DNA Methylation Patterns in the Brassicaceae is Driven by Differences in Genome Organization
- Regulation of mRNA Abundance by Polypyrimidine Tract-Binding Protein-Controlled Alternate 5′ Splice Site Choice
- Systematic Comparison of the Effects of Alpha-synuclein Mutations on Its Oligomerization and Aggregation
- Rad59-Facilitated Acquisition of Y′ Elements by Short Telomeres Delays the Onset of Senescence
- A Functional Portrait of Med7 and the Mediator Complex in
- Systematic Analysis of the Role of RNA-Binding Proteins in the Regulation of RNA Stability
- ARTIST: High-Resolution Genome-Wide Assessment of Fitness Using Transposon-Insertion Sequencing
- Genomic Evidence of Rapid and Stable Adaptive Oscillations over Seasonal Time Scales in Drosophila
- Genome-Wide Associations between Genetic and Epigenetic Variation Influence mRNA Expression and Insulin Secretion in Human Pancreatic Islets
- HAM-5 Functions As a MAP Kinase Scaffold during Cell Fusion in
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- An RNA-Seq Screen of the Antenna Identifies a Transporter Necessary for Ammonia Detection
- Systematic Comparison of the Effects of Alpha-synuclein Mutations on Its Oligomerization and Aggregation
- Functional Diversity of Carbohydrate-Active Enzymes Enabling a Bacterium to Ferment Plant Biomass
- Regularized Machine Learning in the Genetic Prediction of Complex Traits
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



