-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Role of Exonic Variation in Chemokine Receptor Genes on AIDS: Association with Pneumocystis Pneumonia
Chromosome 3p21–22 harbors two clusters of chemokine receptor genes, several of which serve as major or minor coreceptors of HIV-1. Although the genetic association of CCR5 and CCR2 variants with HIV-1 pathogenesis is well known, the role of variation in other nearby chemokine receptor genes remain unresolved. We genotyped exonic single nucleotide polymorphisms (SNPs) in chemokine receptor genes: CCR3, CCRL2, and CXCR6 (at 3p21) and CCR8 and CX3CR1 (at 3p22), the majority of which were non-synonymous. The individual SNPs were tested for their effects on disease progression and outcomes in five treatment-naïve HIV-1/AIDS natural history cohorts. In addition to the known CCR5 and CCR2 associations, significant associations were identified for CCR3, CCR8, and CCRL2 on progression to AIDS. A multivariate survival analysis pointed to a previously undetected association of a non-conservative amino acid change F167Y in CCRL2 with AIDS progression: 167F is associated with accelerated progression to AIDS (RH = 1.90, P = 0.002, corrected). Further analysis indicated that CCRL2-167F was specifically associated with more rapid development of pneumocystis pneumonia (PCP) (RH = 2.84, 95% CI 1.28–6.31) among four major AIDS–defining conditions. Considering the newly defined role of CCRL2 in lung dendritic cell trafficking, this atypical chemokine receptor may affect PCP through immune regulation and inducing inflammation.
Published in the journal: . PLoS Genet 7(10): e32767. doi:10.1371/journal.pgen.1002328
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002328Summary
Chromosome 3p21–22 harbors two clusters of chemokine receptor genes, several of which serve as major or minor coreceptors of HIV-1. Although the genetic association of CCR5 and CCR2 variants with HIV-1 pathogenesis is well known, the role of variation in other nearby chemokine receptor genes remain unresolved. We genotyped exonic single nucleotide polymorphisms (SNPs) in chemokine receptor genes: CCR3, CCRL2, and CXCR6 (at 3p21) and CCR8 and CX3CR1 (at 3p22), the majority of which were non-synonymous. The individual SNPs were tested for their effects on disease progression and outcomes in five treatment-naïve HIV-1/AIDS natural history cohorts. In addition to the known CCR5 and CCR2 associations, significant associations were identified for CCR3, CCR8, and CCRL2 on progression to AIDS. A multivariate survival analysis pointed to a previously undetected association of a non-conservative amino acid change F167Y in CCRL2 with AIDS progression: 167F is associated with accelerated progression to AIDS (RH = 1.90, P = 0.002, corrected). Further analysis indicated that CCRL2-167F was specifically associated with more rapid development of pneumocystis pneumonia (PCP) (RH = 2.84, 95% CI 1.28–6.31) among four major AIDS–defining conditions. Considering the newly defined role of CCRL2 in lung dendritic cell trafficking, this atypical chemokine receptor may affect PCP through immune regulation and inducing inflammation.
Introduction
Functional variation in the human leukocyte antigen (HLA) class I genes and in chemokine receptors affects HIV susceptibility, viral load, and rates of disease progression [1]–[4]. Recent genome-wide association studies (GWAS) performed in HIV-1 cohorts have shown that the HLA region and the chemokine receptor CCR5 gene have major roles in control of HIV-1 replication and disease progression—together they explain approximately 20% of genetic variability [3], [5]–[8] (reviewed in [4], [5]). These findings from GWAS highlighted the leading role of chemokine receptors among non-HLA genes in HIV-1 pathogenesis and prompted us to assess the role of other chemokine receptor genes on HIV disease using a gene-centric approach to identify common or rare functional variants in the region.
The chemokine receptor cluster on chromosome 3 contains at least 12 genes including CCR5, the primary HIV-1 co-receptor [9]–[11]. Multiple genetic variants in chemokine receptors and chemokines have been identified as modifiers of HIV-1 infection or disease progression [12], [13], including CCR5-Δ32 (a 32-bp deletion introduces a premature stop codon) [14] and CCR5 promoter variants [15]–[17] and variants in the CCR5 ligand gene CCL5 [18], [19]. The homozygous CCR5 Δ32/Δ32 genotype and complex heterozygotes with other rare amino acid mutations confers near complete resistance to HIV infection [12], [14], [20]–[22]. Individuals homozygous for a haplotype known as CCR5-P1 [15] or haplogroup HHE [23], a multisite allele of the CCR5 promoter region, progress to AIDS more rapidly than those with other CCR5 promoter haplotypes [15]–[17], [23]–[25]. CCR2 and CXCR6 are minor HIV-1 coreceptors used by a limited number of HIV-1 strains as an entry coreceptor [26], [27]. CCR2-V64I has been associated with delayed progression [12], [25], [28], [29]. Variants in CXCR6 were also associated with disease modification [30], [31].
The chromosome 3 chemokine receptor cluster extends from 3p21 to 3p24, with eight receptors occurring in an 520 kb region of 3p21(Figure 1) [32]. The cluster contains genes for several receptors, CCR3, CCR8, CX3CR1, and CXCR6 that have been shown to bind HIV env or to support varying levels of in vitro replication of HIV-1, HIV-2 or simian immunodeficiency virus (SIV) [26], [33]–[36] (reviewed by [11], [37]). The role played by minor coreceptors in HIV-1 pathogenesis is not clear, but studies have suggested that a broad spectrum of coreceptor usage may be correlated with rapid CD4+ cell depletion and AIDS progression [11], [38], [39]. Primary isolates of HIV-1 have been shown to use a wide spectrum of various chemokine receptors as HIV coreceptors [40]. HIV-1 isolates from a CCR5-Δ32 heterozygous or homozygous individual can use various minor coreceptors such as CCR3, CCR2B, CCR8, CX3CR1 for cell entry [41], [42]; amino acid mutations in the V3 loop of HIV-1 are responsible for utilization of multiple coreceptors [40]. Considering the high mutation rate and sequence heterogeneity of HIV-1, particularly within the env gene, it is plausible that a spectrum of receptors is used in vivo during the course of HIV infection and that genetic variants in the coreceptors may affect usage or binding efficiency by HIV-1. Furthermore, as CCR5 and CXCR4 antagonists blocking these major co-receptors are used therapeutically [37], the potential of HIV-1 to evolve to use other minor coreceptors as alternative cell entry points is expected to increase. Therefore, determining whether HIV-1 minor coreceptor genes, in addition to CCR5 and CXCR4 play a role in HIV pathogenesis is a timely topic.
Fig. 1. Polymorphisms in seven chemokine receptor genes on the Chromosome 3p21–22. 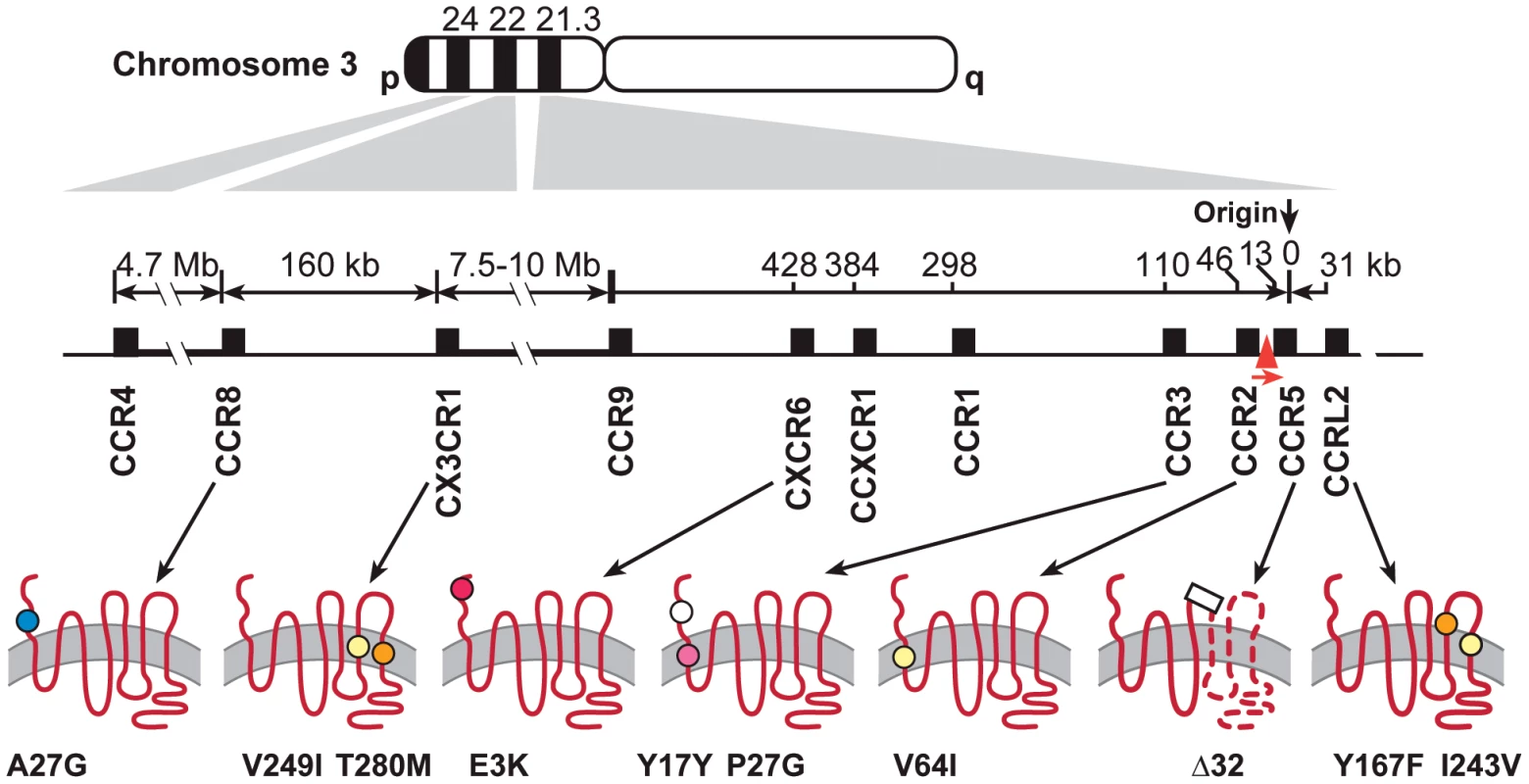
Schematics at bottom show the configuration of the receptors in the cell membrane, with circles representing the position and type of the polymorphisms. White circle indicates a synonymous substitution, colors nonsynonymous substitutions: yellow, conservative; blue, hydrophobic—hydrophilic; orange, hydrophilic—hydrophobic; red, charge changing (acidic—basic), purple, shape changing. Also indicated are Δ32 in CCR5 (rectangle) and the polymorphic CCR5 promoter (red triangle in the gene map). CCR5-2459 promoter allele (rs1799987). In this study, we evaluate the impact of chemokine coreceptors (CCR) on HIV/AIDS using a candidate-gene based population association analysis in five treatment-naive HIV-1 natural history cohorts. Genotypes of exonic polymorphisms in chemokine coreceptor genes CCR3, CCRL2 and CXCR6 on Chromosome 3p21 and in CCR8 and CX3CR1 on 3p22 were tested for their genetic influence on AIDS progression. CCR3, CCR8 and CXCR6 were chosen as they are HIV-1 minor coreceptors [26], [33]–[36] (reviewed by [11], [37]). CCRL2 was selected as a candidate gene because of its homology with CCR5 (45%)—the most of any of the chemokine receptors genes—because of its proximity to CCR5, and because it is an atypical receptor without signal transduction, similar to DARC. Our results suggest that genetic variation in CCR3, CCR8 and CCRL2 may contribute additional genetic regulation of HIV-1 disease in addition to that conferred by the major HIV-1 coreceptor gene CCR5.
Results
Identification of chemokine coreceptor variants
We resequenced selected chemokine receptor genes in the chromosome 3p21–22 region in 72 African Americans and 72 European Americans with three extreme phenotypes (resistance to HIV infection, very rapid or slow progression to AIDS) to assess the extent of exonic variation and to identify rare variants. We observed a total of 6 exonic variants, 5 of which were nonsynonymous, in CCR3, CCR8, CXCR6, and CCRL2 (Figure 1, Table 1). We did not resequence CCR5 and CCR2 since these receptor genes had been previously sequenced in HIV patients [28], [43], nor did we resequence CX3CR1. We previously showed that CX3CR1-V249I and -T280M had no effect on AIDS progression in our group of seroconverter subjects [44], and therefore did not include them in this analysis. No noticeable differences in SNP frequency among three extreme phenotype groups were observed (data not shown). These SNPs were then genotyped in 5 HIV-1 natural cohorts comprising 2594 European Americans (EA). All SNPs were in Hardy-Weinberg Equilibrium (P>0.05). CXCR6-E3K and CCR3-P39L were rare (<1%) in EA and were excluded from the analysis; other SNPs were common (>5%) (Table 1).
Tab. 1. Characteristics and allele frequencies of the chemokine receptor variants. 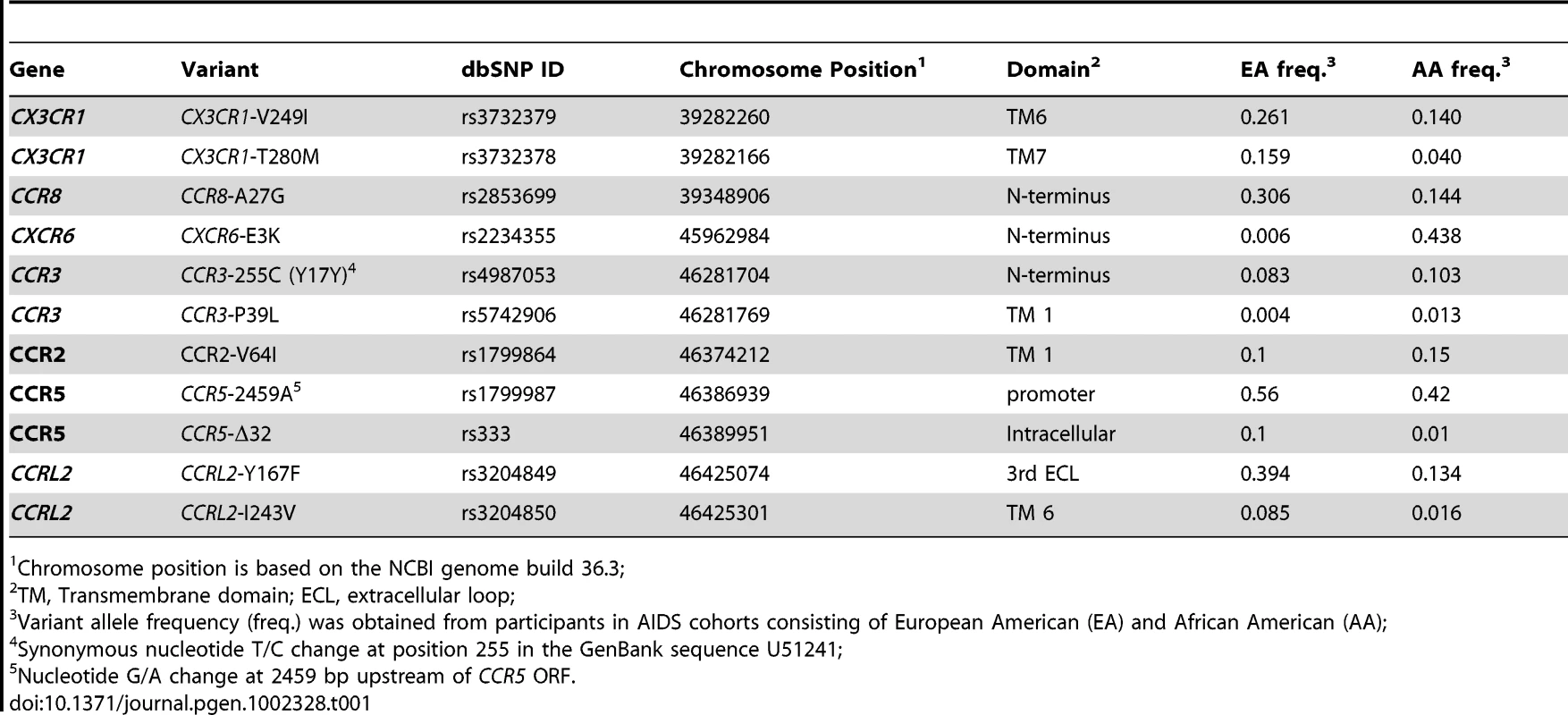
Chromosome position is based on the NCBI genome build 36.3; Linkage disequilibrium (LD) of chemokine receptor variants on 3p21–22
The SNPs considered fell into two distinct haplotype blocks (Figure 1). The larger block in 3p21 comprises CCRL2-F167Y, CCRL2-I243V, CCR5-+/Δ32, the CCR5-promoter SNP CCR5-2459A (rs1799864, previously shown to affect CCR5 expression levels and modify AIDS progression [15], [16]), CCR2-V64I, CCR3-P39L and CCR3-255T/C (Y17Y), spanning 77 Kb, and CXCR6-E3K, 318 Kb teleomeric to CCR3. The second block in 3p22, ∼10 Mb from the first block, consists of CX3CR1-V249I and -T280M and CCR8-A27G (rs2853699), which are separated by ∼160 Kb. There is moderate to substantial LD within the closely spaced CCR3-CCR2-CCR5-CCRL2 group (with pairwise D′ 0.47–1, Figure S1). In the 3p22 block, low levels of LD were observed between CCR8 and CX3CR1 SNPs (D′ ∼0.40). The LD level between 3p21 and 3p22 blocks is minimal.
Genetic associations with progression to AIDS
The identification of independent additional genetic factors in this region, particularly for 3p21, is complicated by a moderate to high level of LD (Figure S1). To detect genes or markers additional to a primary predisposing variant(s) in a genetic region of high LD, stratification analyses or using a restricted dataset are frequently employed [45]. Stepwise regression is also a choice for assessing the relative importance of different variants in the linked region [46]. These approaches are feasible as the SNPs in this region have a low to moderate correlation (for r2, see Figure 2).
Fig. 2. Genetic effect of CCR3-255C and CCRL2-243V on AIDS progression. 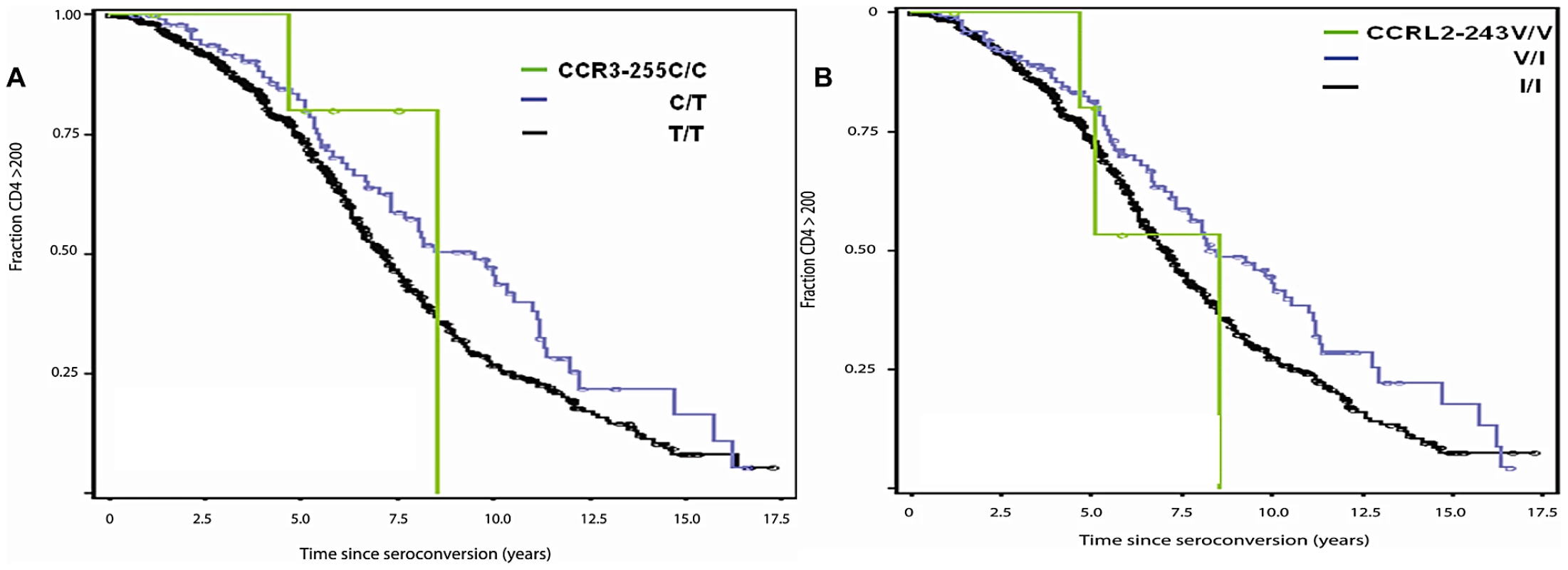
Kaplan-Meier survival curves of time progression since HIV-1 infection to AIDS (1987 CDC definition) were plotted for genotypes of CCR3-255C (A) and CCRL2-243V (B). Circles on the curves indicate those censored. Effects of individual variants on AIDS progression
SNPs with allele frequencies of at least 5% were analyzed by Kaplan-Meier survival curve analysis and Cox proportional hazards model. We present the relative hazards (RH) of the individual variants on survival to AIDS from analysis of 670 EA seroconverters using multivariable Cox regression models in Table 2. Four of the five SNPs in the 3p21 block revealed new significant associations with delayed time progressing to AIDS, after conditioning on HLA alleles and CCR5-Δ32 (Table 2). Significant associations with differential progression to clinical AIDS were observed for CCR3 -255C (RH = 0.62, P = 0.009, Figure 2A), for CCRL2-243V (RH = 0.66, P = 0.03, Figure 2B), and for CCRL2-167F (RH = 1.89, P = 0.0003, Figure 3A) and for CCR8-27G (RH = 1.44, P = 0.004) (Table 2). Adjusting for potential population stratification had little affect on the significance of the associations (Table 2). Correcting for 7 chemokine receptor variants tested using a Bonferroni step-down (Holm) correction [47], the associations for CCRL2-167Y, CCRL2-243V, and CCR3-255C remained significant (P = 0.002, 0.03, and 0.016, respectively), in addition to CCR5-Δ32 and CCR5-2459, while CCR8-27G became non-significant (Table 2).
Fig. 3. Genetic effect of CCRL2-167F on AIDS progression and PCP. 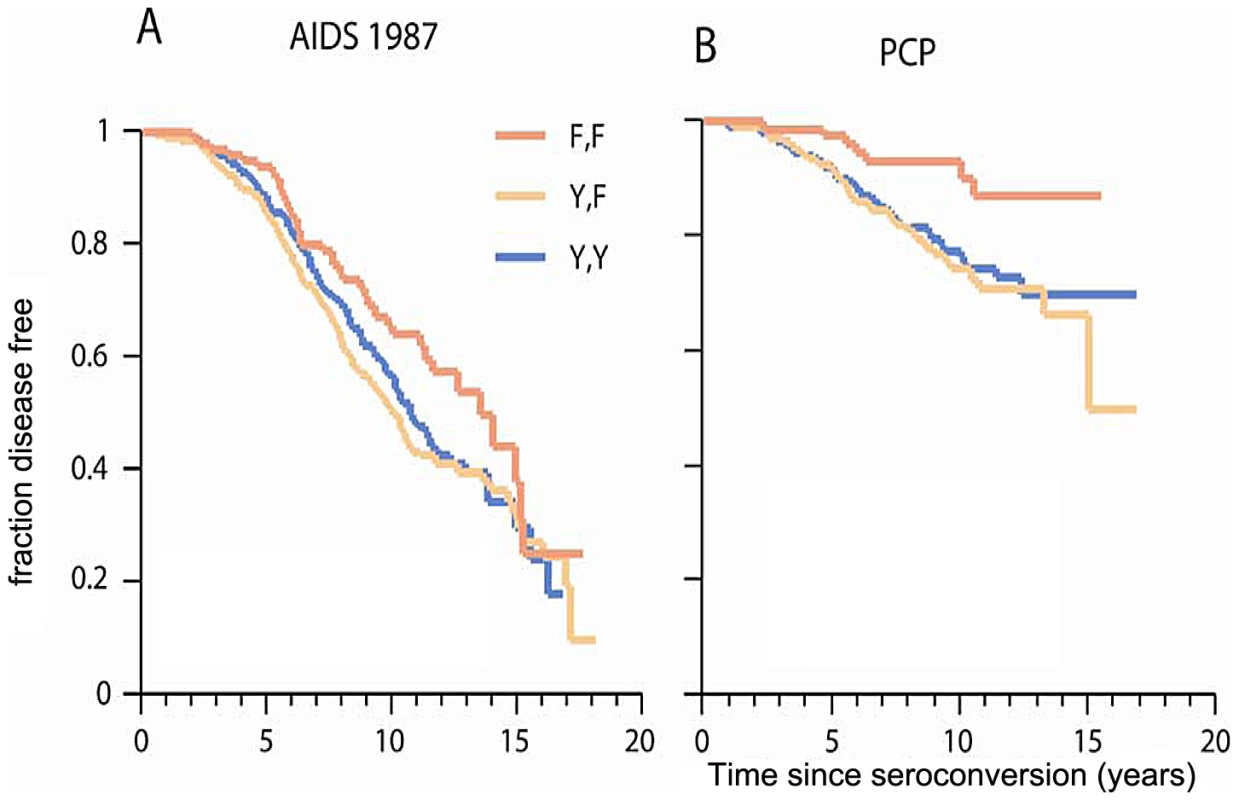
Kaplan-Meier survival curves of time progression since HIV-1 infection to AIDS (1987 CDC definition) (A) or to PCP (B) were plotted for CCRL2-Y167F genotypes. Circles on the curves indicate those censored. Tab. 2. Effects of chemokine receptor variants on AIDS progression. 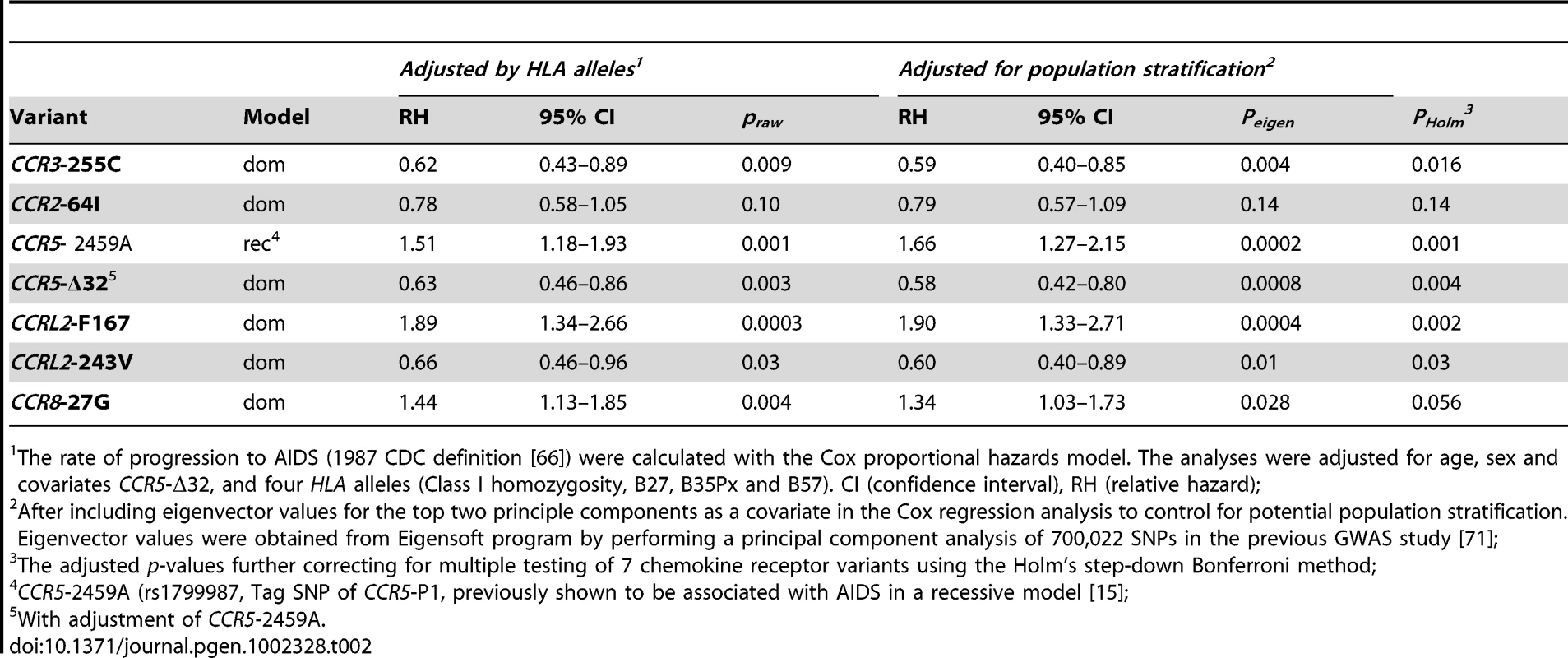
The rate of progression to AIDS (1987 CDC definition [66]) were calculated with the Cox proportional hazards model. The analyses were adjusted for age, sex and covariates CCR5-Δ32, and four HLA alleles (Class I homozygosity, B27, B35Px and B57). CI (confidence interval), RH (relative hazard); Effects of combined chemokine coreceptor variants on AIDS progression
Using stepwise selection determined by the Akaike information criteria (AIC), we built an optimal prediction model with the genetic variants that were associated with AIDS progression in this or previous studies (Table 3). From Table 3, we can see the minimum AIC is achieved at the ninth step with nine covariates. Therefore, the best predictive model accounting for the rate of progression to AIDS (1987 CDC definition) includes the newly identified variants CCRL2-167F and -243V, CCR8-27G, as well as previously known variants: the two locus CCR2-V64I-CCR5-Δ32 composite genotype, CCR5-2459, HLA class 1 homozygosity, HLA-B*35Px, HLA-B*57 and HLA B*27 (without covariates, AIC:1796.32, with 9 covariates, AIC:1737.28). The significance level and RH values in this model represent that of each variant after considering all other linked and unlinked variants (Table 3). The rough shrinkage estimate is 0.88 (104/140), indicating that the model is fairly reliable without further shrinkage or data reduction treatment [48]. We calculated that R2 = 0.11 for the final fitted model, indicating that 11% of the variability in AIDS progression can be explained with the variants in the model.
Tab. 3. Sequential stepwise regression for Cox PH model selection and maximum likelihood estimates of genetic covariates associated with AIDS progression. 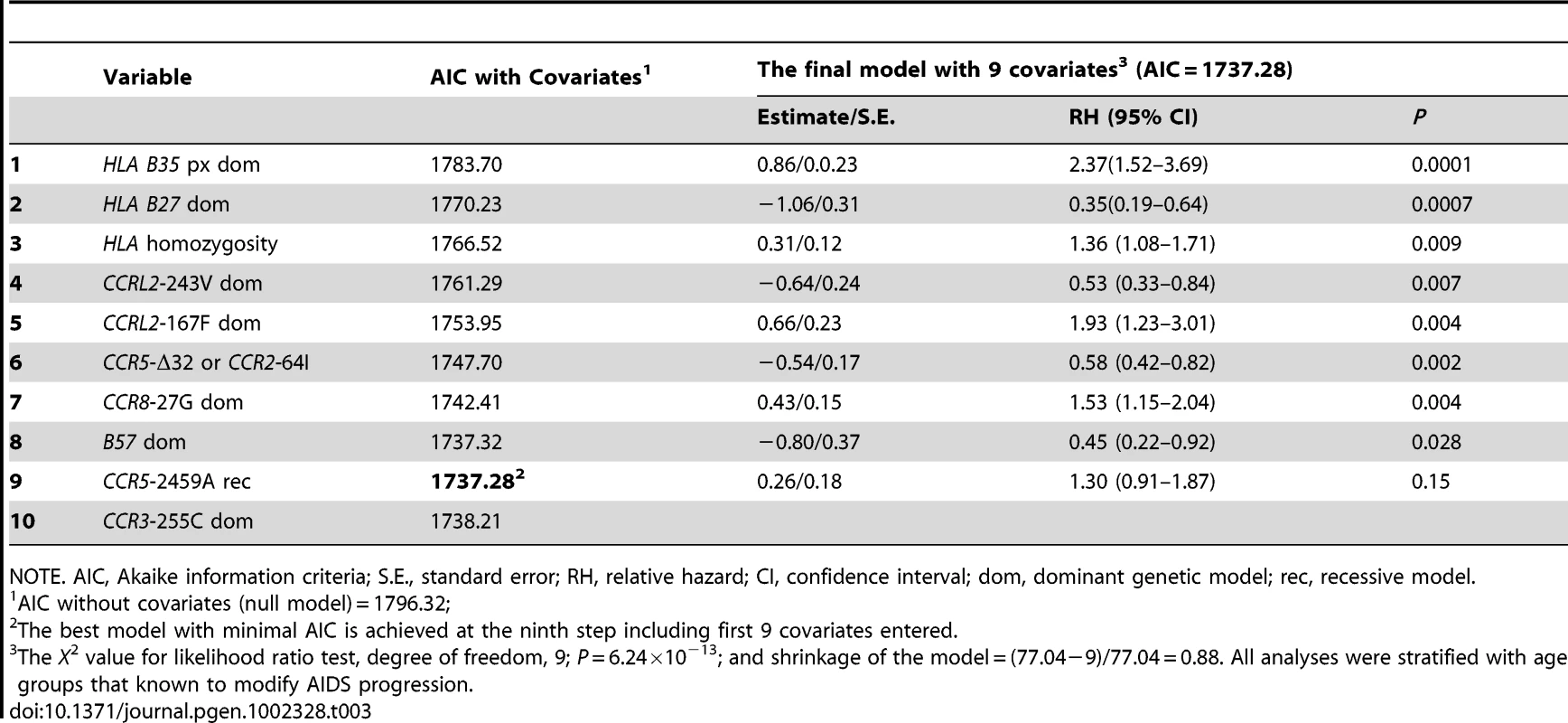
NOTE. AIC, Akaike information criteria; S.E., standard error; RH, relative hazard; CI, confidence interval; dom, dominant genetic model; rec, recessive model. CCRL2 - Y167F is associated with progression to AIDS
The strongest new association was for CCRL2-Y167F with progression to clinical AIDS. The common 167F allele (allele frequency 67% in EA) had a significant dominant detrimental effect on progression to AIDS-defining conditions (CDC 1987 case definition) (RH = 1.89, P = 0.0003, Wald test; P = 0.0001, Likelihood ratio test; n = 670 seroconverters) in the Cox proportional hazards model (Table 2 and Table 3, Figure 3A). When the analysis was restricted to individuals not carrying CCR5-Δ32 (n = 539), the association of CCRL2-167F with AIDS remained significant (RH = 1.69, 95% CI 1.20–2.38), indicating that the CCRL2-167F association is independent of CCR5-Δ32 and not due to its linkage disequilibrium with the latter.
CCRL2 - Y167F and pneumocystis pneumonia (PCP)
In an explanatory analysis, we tested whether CCRL2 - Y167F affects AIDS progression through a specific AIDS-defining illness. A Cox model analysis in the seroconverter group showed significantly faster progression to PCP for carriers of 167F (RH = 2.84, 95% CI 1.28–6.31, P = 0.007; Figure 3B), but no differential effect on Kaposi's sarcoma, microbacterial avium infection, cytomegalovirus, or lymphoma (P>0.05, data not shown).
To assess whether the CCRL2-167F association with PCP was due to LD (D′ = 0.76, r2 = 0.47) with CCR5-2459 that associated with AIDS progression [16], we restricted the analysis in a group of SC (n = 365) that do not carry the homozygous CCR5-2459 genotype; the CCRL2-167F association with PCP remained significant (OR = 2.42, 95% CI 1.09–5.36), in support of its independent role.
Possible function of CCRL2 F167Y
Evolutionary conservation of CCRL2
We assessed the homology of CCRL2 with other C-C chemokine receptor family genes. CCRL2 shares over 40% sequence homology with known chemokine receptors such as CCR1, -2, -3 and -5 but lacks the DRYLAIV motif in the second extracellular loop that is usually required for the signaling capacity of chemokine receptors [49]. The alignment of mammalian CC chemokine coreceptor sequences shows that the 167 site and surrounding region are highly conserved (Figure S2). Phenylalanine (F) at position 167 is apparently ancestral as it occurs in chimpanzee and macaque. Phenylalanine (F) or other nonpolar amino acids occur at the position homologous to human CCRL2-167 in 78 of the 82 chemokine receptors, including all receptors known to be functional, while a polar amino acid occurs at this position in two cases for the non-signaling FY DARC receptor. Tyrosine (Y) never occurs at this position except for human CCRL2. These observations suggest that the F167Y change might have functional consequence.
Impact of F167Y on CCRL2 topology
The impact of F167Y on CCRL2 transmembrane (TM) topology was assessed using ConPred II, [50]. 167F is located in the third extracellular loop adjacent to the 4th TM domain. The F to Y substitution introduces a hydrophilic residue that changes the boundary of the TM domain and the conformation of the nearby domains: TM domains 3, 4 and 5 were formed by residues 104–124, 146–166 and 201–221 with CCRL2 carrying 167F, but were formed by residues 103–123, 145–165 and 202–222 with CCRL2-167Y, respectively (Figure S3).
Three-dimensional protein structure prediction
The effect of the Y167F amino acid substitution on the structure of the CCRL2 receptor was predicted by PROTINFO program. In the structure incorporating 167Y, the hydroxyl (OH) group of the tyrosine makes a hydrogen bond with the nitrogen atom of glycine 108 (G108) (Figure S4). This hydrogen bond is not present with F167, or with the other amino acids commonly observed at this position.
Surrogate migration assay using CCR3
As an indirect test of possible function of CCRL2-F167Y, HEK293 cells expressing human CCR3 containing either phenylalanine (169F) or tyrosine (169Y) at the homologous position were tested for migration in a chemotaxis assay. Mutation of 169F to 169Y in CCR3 markedly reduced (3-fold) the migration of HEK293 cells in response to eotaxin (Figure S5), indicating a role of 169F in CCR3 function and by analogy suggesting that CCRL2-F167 may be a functionally superior receptor than CCRL2-167Y.
Chemotaxis of CCRL2
We tested chemotactic responses of HEK293 cells transfected with either CCRL2-F167 or CCRL2-167Y. We observed no chemotactic responses to a panel of 13 chemokines or peptides including the HIV-1 inhibitory chemokines CCRL3, 4, 5 and CXCL12, suggesting CCRL2 is not a receptor of these chemokines. However, our experiments did not include the recently identified CCRL2 ligands, chemerin and CCL19 [51], [52], due to availability.
Discussion
We investigated the impact of genetic variation of 3p21–22 chemokine receptor genes on HIV/AIDS in this population-based association study. We identified a total of 6 exonic variants in CCR3, CCR8, CXCR6 and CCRL2 in 144 samples with extreme phenotypes. Through Cox regression survival analysis of the variants in HIV-1 natural cohorts, we identified three polymorphisms (CCRL2-Y167F, CCR3-255C and CCR8-27G) as having previously unidentified correlation with AIDS progression that appear to confer additional effect beyond the well-studied AIDS-modifying polymorphisms CCR5-Δ32, CCR5-P1/CCR5-2459A, and CCR2-64I. CCRL2-167F was associated with strong accelerated progression to AIDS, resulted almost entirely from rapid development of the AIDS-defining disease PCP.
The coverage of SNPs in the 3p21–22 region is limited in the GWAS SNP chips. Several exonic SNPs (CCRL2-167F, -243V, CCR8-27G, CCR5-2459A, CCR5 - Δ32) genotyped in this study were not included in the SNP chips that have been used in GWAS for HIV-1 [3], [5]–[8], highlighting the ongoing need of candidate gene analysis in the GWAS era. The newly observed effects of chemokine coreceptor genes thus await further replication.
It must be cautioned that the combination of the presence of multiple AIDS associations in this chemokine receptor complex, and the LD between the receptor genes, makes determining the true source of the associations difficult. Overall, however, the association analysis points to receptor gene variant associations beyond those that can be attributed to the known AIDS affecting receptor gene polymorphisms. The association of additional chemokine receptors, beyond the primary CCR5 and CXCR4, with AIDS progression is plausible as several of these have been shown to bind to HIV in varying degrees [26], [35], [36], [39]–[42], [53]. The complexity of associations in this region makes it essential to identify a functional effect of the genetic variants on disease, before concluding that the associations are real. It should be noted that we may have not detected all existing SNPs in the region and was also underpowered in detecting rare SNPs (1%). We found that another CCRL2 exonic SNP in the public domain (168M, rs6441977) was not associated with progression to AIDS-87 (RH = 0.84, 95% 0.50–1.39) in our seroconverters. We performed haplotype analysis for the three exonic SNPs in CCRL2 and found that only the haplotypes bearing rs3204849 A (CCRL2 Y167F) and rs3204850 (V243I) were associated with AIDS; the haplotype bearing rs6441977A (V168M) had no effect. No additional information was gained by performing a haplotype analysis compared to single SNP analysis.
With these caveats we argue that the association of the CCRL2-167F variant is worthy of interest. First, within the noted limits of the association analysis, the association is quite strong (RH = 1.9, P = 0.002, corrected), remains significant with CCR5-Δ32 taken into account, and in an automatic selection analysis with all known factors taken into account. Second, although there is no direct demonstration of function for this polymorphism, and the F to Y substitution is generally a conservative one, several lines of phylogenetic, chemical modeling, and indirect experimental data suggest that CCRL2-167Y significantly alters the properties of this receptor.
Protein structure modeling data suggest that the risk-associated F to Y substitution could change the boundary of the transmembrane domain, and introduce a hydrogen bond. Of particular interest is the alignment data of the conservation of this residue among CC chemokine receptors. Strikingly, phenylalanine, the ancestral alternate variant to the AIDS risk tyrosine variant, or another nonpolar amino acid, occurs at this position in all receptors known to be functional [54]; a polar amino acid only occurs in the two cases of the nonsignaling DARC receptor. Further, substitution of tyrosine for phenylalanine at this position in the functional receptor CCR3 reduced migration of HEK293 cells in response to eotaxin threefold. We emphasize that all of these tests are in silico or indirect, and direct test of the effect of the F to Y substitution on the function of CCRL2 remains to be done; new knowledge of the functions and ligands of CCRL2 should make this more straightforward.
The association of CCRL2 with AIDS and PCP is unique as CCRL2 has not been shown to serve as a coreceptor for HIV-1. Our chemotaxis assay experiments excluded 11 common chemokines as ligands of CCRL2. The association does not appear to be a general function of increased HIV susceptibility, but instead specifically attributable to an increase in PCP among individuals carrying this receptor variant. PCP, caused by Pneumocystis jirovecii, is the most common opportunistic infection in untreated HIV-1-infected immunosuppressed persons. PCP is mediated by marked inflammatory responses in lung involving macrophages and chemokines and cytokines [55]. The association of CCL2 with PCP might be attributable to two aspects of CCRL2: immune regulation or direct interaction with HIV. CCRL2 may affect PCP through its immune regulating role at local inflammation sites, possibly by concentrating and presenting chemokines [51], [52], [56]. CCRL2 was rapidly upregulated in murine lung macrophages following inflammation induction [57] and deficiency of CCRL2 impaired lung dendritic cell migration [58]. Alternatively, CCRL2 may influence HIV coreceptors entry through interacting or sequestering with CCL5 or anti-HIV chemokines [56], or may serve as a coreceptor for some strains of HIV-1. The mechanism of CCRL2-F167Y effect on PCP remains to be explored.
In summary, this comprehensive study of the chromosome 3 chemokine receptor cluster region identified multiple genetic variants that associated with HIV disease. The strongest new association appears to result from an increased susceptibility to PCP, rather than from a specific effect on HIV. Added to the existing knowledge of the effect of the chromosome 3 chemokine receptors on HIV disease, which has been already been exploited therapeutically, our results affirm this gene complex as a fertile ground for further research, both for HIV and potentially for a broad range of additional diseases.
Materials and Methods
Ethics statement
Institutional review boards (IRB) at National Cancer Institute, National Institutes of Health and participating institutes approved the study protocols. Written informed consent was obtained from all study participants and/or their legal guardians.
Study participants
The study group includes 674 HIV-1 seroconverter European Americans, 669 seronegatives enrolled in the following natural history HIV-1 cohort studies: Multicenter AIDS Cohort Study (MACS) [59], the San Francisco City Clinic Cohort Study (SFCCC) [60], AIDS Link to the Intravenous Experience (ALIVE) [61], Hemophilia Growth and Development Study (HGDS) [62], and the Multicenter Hemophilia Cohort Study (MHCS) [63]. Seroconversion date was estimated as the midpoint between the last seronegative and the first seropositive HIV-1 antibody test date (mean interval 0.79 years, range 0.07–3.0 years). The censoring date was the earliest of the date of the last recorded visit, or July 31, 1997 for the ALIVE cohort, or December 31, 1995 for all other cohorts, to avoid the confounding effect of highly active anti-retroviral therapy (HAART). A later censoring date was used for ALIVE cohort because few ALIVE participants received HAART prior to July 31, 1997 [64].
Resequencing and SNP identification
A panel of 72 EA and 72 AA samples representing extreme phenotypes for infection and progression (rapid progression, long term non-progression, and infection resistance) were used for SNP identification. The 5′ and 3′ untranslated (UTR) and coding regions of the CCR3, CCR8, CCRL2, and CXCR6 genes were PCR amplified by overlapping primer sets (Table S2). PCR products were resequenced by BigDye terminator (Applied Biosystems, Foster City, CA). We did not sequence or genotyping SNPs in CCR1, CCXCR1, CCR9 and CCR4 as they are not recognized as HIV-1 coreceptors [26], [33]–[36] (reviewed by [11], [37]).
Genotyping of genetic variants
Genotyping was done using PCR-restriction fragment length polymorphism (RFLP) or TaqMan assays. PCR primer sequences, TaqMan probes and primers, PCR conditions, and restriction enzymes used to genotype each variant are listed in Table S1. Briefly, PCR was carried out with 35 cycles of denaturing at 94°C for 30 s, annealing at 54–60°C for 30 s and extension at 72°C for 45 s. TaqMan assays were performed according to the manufacturer's manual (Applied Biosystems, Foster City, CA). The CX3CR1 variants V249I and T280M were typed as previously reported [65].
Statistical analysis
Kaplan-Meier survival statistics and the Cox proportional hazards (PH) model (Cox PH model) were used to assess the effects of genetic variants on the time of progression from HIV-1 infection to AIDS (1987 CDC definition) [66], using PROC PHREG and LIFETEST of SAS version 9.13 (SAS, Cary, North Carolina). For SNPs that showed significant association with AIDS development, explanatory analyses were performed for their specific impact on the AIDS-defining diseases Pneumocystis pneumonia (PCP), Kaposi's sarcoma, microbacterial avium infection, and cytomegalovirus. The relative hazard (RH) and significance of associations were determined using a Cox PH model without or with adjustment for confounding genetic factors not on chromosome 3: for EA HLA-B*27 and HLA-B*57, HLA-B*35Px, and HLA Class I homozygosity [1], [67], [68]; for AA CCRL5-In1.1, HLA-B*57 and HLA Class I homozygosity [1], [19], [67]. CCR2-64I, CCR5-Δ32 and CCR5-2459 were also included as covariates in the adjusted multivariable regression analysis. CCR5 promoter haplotypes (P1, P2, and P4) are tagged by SNPs CCR5-2459 (rs1799987) and rs2734648 [43]. A visual inspection of the data with Kaplan-Meier survival curves was performed to determine the genetic models to be used in the Cox PH regression model. A dominant genetic model was tested for all genetic factors in this study, except for CCR5-2459 (recessive) [15], [16]. Participants were stratified by sex and by age at seroconversion: 0–20, >20–40, and over 40 years.
To determine the best explanatory set of genetic variants, while minimizing the number of comparisons in model selection, We used StepAIC procedure to build Cox proportional hazards models for the AIDS-1987 phenotype based on stepwise regression, Akaike information criteria (AIC), and the best subset selection [69]. Here we used PROC PHREG in SAS software with SLENTRY = 0.99 and SLSTAY = 0.995 (values chosen close to 1 to generate a sequence of models from null models to full model ordered by AIC) [48]. Model uncertainty caused by including large number of variables can be estimated by shrinkage (LR-p)/LR, where LR denotes the likelihood ratio χ2 and p denotes the number of the predictors in the final model. A shrinkage below 0.85 raises concern of overfitting [48]. It is recommended that no more than m (the number of uncensored event)/10 predictor degree of freedom p (number of parameters) should be examined to fit a multiple regression model. As in our sample, there were 194 events without censoring, we expect that fitting with <19 variables would be appropriate [48].
To control for potential population stratification, we adjusted the regression analysis with eigenvector values [70]. Eigenvector values were obtained by performing principal component analysis of 700,022 SNPs from a previous GWAS study carried out in the same samples [71]. In the Cox regression analysis, eigenvector values for the top two principle components were included as covariates. The genomic inflation factor in this seroconverter population showed minimal systematic overall bias due to population structure in regards to disease progression phenotype as it was quite close to 1.0 (λ = 1.01) (expected under no population stratification) [71].
We further corrected for multiple comparisons by counting 7 chemokine receptor variants tested using a Bonferroni step-down (Holm) correction method [47], as implemented in the MULTITEST procedure in SAS.
The “generalized” R2 statistic in Cox model is based on the likelihood-ratio statistic (LRT) for testing the global null hypothesis [72]. The formula is given as: R2 = 1−e−(LRT/n), where LRT = −2logL(0)−[−2logL(p)], n is the total sample size, logL(0) is the log-likelihood for a null model with no covariates, and logL(p) is the log-likelihood for the fitted model with p covariates.
We quantified LD between all pairs of biallelic SNPs using the absolute unsigned value of Lewontin's D′ statistic [73]. P values represent significance of departure from the null hypothesis that the pair is in equilibrium. All P values are two-tailed. Haploview was used for the LD plots.
Prediction of CCRL2 protein structures
Three-dimensional models of the two CCRL2 proteins with 167F or 167Y were constructed using the PROTINFO structure prediction server (http://www.protinfo.compbio.washington.edu), using the comparative modeling protocol [74], [75]. The detailed modeling method is presented in Text S1. The CCRL2 TM topology was established using ConPred II (http://bioinfo.si.hirosaki-u.ac.jp/~ConPred2/), a predication program based on consensus results of several prediction methods including TMpred, TMAP,TMHMM, HMTOP and MEMSAT [50].
Expression of human CCRL2 in HEK293 cells
cDNAs coding for the full-length open reading frame (ORF) of human CCRL2 carrying 167Y or 167F were PCR amplified from two individuals with the respective homozygous genotype with proofreading DNA polymerase pfu (Stratage, La Jolla, CA). After confirmation of sequence accuracy, they were ligated into the pcDNA3 (Invitrogen, Gaithersburg, MD). Human embryonic epithelial cells line HEK293 was transfected with the constructs of CCRL2-167F or CCRL2-167Y. Stable transfectants were selected by culturing the cells in 800 µg/ml G418. Once the stable cell lines were established, they were examined for chemotactic responses to chemokines.
Chemotaxis assays
Migration of CCRL2-167F and CCRL2-167Y transfected HEK293 cells was assessed using a 48-well microchemotaxis chamber technique. They were examined for chemotactic responses to the following chemokines: CCL5, CCL3, CCL4, CCL2, CCL8, CCL7, CCL11, CXCL12, CCL21, CCL20, and CXCL10. The cells were also tested for migration in response to chemotactic peptides using formayl peptide receptors, including W peptide and MMK-1 (Text S1).
Expression of human CCR3 in HEK293 cells
A cDNA coding for 169Y-CCR3 was created from the wild-type human 169F-CCR3 ligated into the pcDNA3, which were used to transfect HEK293 cells (Text S1).
Supporting Information
Zdroje
1. CarringtonMNelsonGWMartinMPKissnerTVlahovD 1999 HLA and HIV-1: heterozygote advantage and B*35-Cw*04 disadvantage. Science 283 1748 1752
2. O'BrienSJNelsonGW 2004 Human genes that limit AIDS. Nat Genet 36 565 574
3. FellayJShiannaKVGeDColomboSLedergerberB 2007 A whole-genome association study of major determinants for host control of HIV-1. Science 317 944 947
4. AnPWinklerCA 2010 Host genes associated with HIV/AIDS: advances in gene discovery. Trends Genet 26 119 131
5. FellayJGeDShiannaKVColomboSLedergerberB 2009 Common genetic variation and the control of HIV-1 in humans. PLoS Genet 5 e1000791 doi:10.1371/journal.pgen.1000791
6. LimouSLe ClercSCoulongesCCarpentierWDinaC 2009 Genomewide association study of an AIDS-nonprogression cohort emphasizes the role played by HLA genes (ANRS Genomewide Association Study 02). J Infect Dis 199 419 426
7. HerbeckJTGottliebGSWinklerCANelsonGWAnP 2010 Multistage genomewide association study identifies a locus at 1q41 associated with rate of HIV-1 disease progression to clinical AIDS. J Infect Dis 201 618 626
8. PereyraFJiaXMcLarenPJTelentiAde BakkerPI 2010 The major genetic determinants of HIV-1 control affect HLA class I peptide presentation. Science 330 1551 1557
9. DengHLiuREllmeierWChoeSUnutmazD 1996 Identification of a major co-receptor for primary isolates of HIV-1. Nature 381 661 666
10. AlkhatibGCombadiereCBroderCCFengYKennedyPE 1996 CC CKR5: a RANTES, MIP-1alpha, MIP-1beta receptor as a fusion cofactor for macrophage-tropic HIV-1. Science 272 1955 1958
11. BergerEAMurphyPMFarberJM 1999 Chemokine receptors as HIV-1 coreceptors: roles in viral entry, tropism, and disease. Annu Rev Immunol 17 657 700
12. IoannidisJPRosenbergPSGoedertJJAshtonLJBenfieldTL 2001 Effects of CCR5-Delta32, CCR2-64I, and SDF-1 3′A alleles on HIV-1 disease progression: An international meta-analysis of individual-patient data. Ann Intern Med 135 782 795
13. O'BrienSJMooreJP 2000 The effect of genetic variation in chemokines and their receptors on HIV transmission and progression to AIDS. Immunol Rev 177 99 111
14. DeanMCarringtonMWinklerCHuttleyGASmithMW 1996 Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. Hemophilia Growth and Development Study, Multicenter AIDS Cohort Study, Multicenter Hemophilia Cohort Study, San Francisco City Cohort, ALIVE Study. Science 273 1856 1862
15. MartinMPDeanMSmithMWWinklerCGerrardB 1998 Genetic acceleration of AIDS progression by a promoter variant of CCR5. Science 282 1907 1911
16. McDermottDHZimmermanPAGuignardFKleebergerCALeitmanSF 1998 CCR5 promoter polymorphism and HIV-1 disease progression. Multicenter AIDS Cohort Study (MACS). Lancet 352 866 870
17. AnPMartinMPNelsonGWCarringtonMSmithMW 2000 Influence of CCR5 promoter haplotypes on AIDS progression in African-Americans. AIDS 14 2117 2122
18. LiuHChaoDNakayamaEETaguchiHGotoM 1999 Polymorphism in RANTES chemokine promoter affects HIV-1 disease progression. Proc Natl Acad Sci U S A 96 4581 4585
19. AnPNelsonGWWangLDonfieldSGoedertJJ 2002 Modulating influence on HIV/AIDS by interacting RANTES gene variants. Proc Natl Acad Sci U S A 99 10002 10007
20. HuangYPaxtonWAWolinskySMNeumannAUZhangL 1996 The role of a mutant CCR5 allele in HIV-1 transmission and disease progression. Nat Med 2 1240 1243
21. SamsonMLibertFDoranzBJRuckerJLiesnardC 1996 Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature 382 722 725
22. LiuRPaxtonWAChoeSCeradiniDMartinSR 1996 Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 86 367 377
23. GonzalezEBamshadMSatoNMummidiSDhandaR 1999 Race-specific HIV-1 disease-modifying effects associated with CCR5 haplotypes. Proc Natl Acad Sci U S A 96 12004 12009
24. WinklerCAHendelHCarringtonMSmithMWNelsonGW 2004 Dominant effects of CCR2-CCR5 haplotypes in HIV-1 disease progression. J Acquir Immune Defic Syndr 37 1534 1538
25. KostrikisLGNeumannAUThomsonBKorberBTMcHardyP 1999 A polymorphism in the regulatory region of the CC-chemokine receptor 5 gene influences perinatal transmission of human immunodeficiency virus type 1 to African-American infants. J Virol 73 10264 10271
26. RuckerJEdingerALSharronMSamsonMLeeB 1997 Utilization of chemokine receptors, orphan receptors, and herpesvirus-encoded receptors by diverse human and simian immunodeficiency viruses. J Virol 71 8999 9007
27. AlkhatibGLiaoFBergerEAFarberJMPedenKW 1997 A new SIV co-receptor, STRL33. Nature 388 238
28. SmithMWDeanMCarringtonMWinklerCHuttleyGA 1997 Contrasting genetic influence of CCR2 and CCR5 variants on HIV-1 infection and disease progression. Hemophilia Growth and Development Study (HGDS), Multicenter AIDS Cohort Study (MACS), Multicenter Hemophilia Cohort Study (MHCS), San Francisco City Cohort (SFCC), ALIVE Study. Science 277 959 965
29. MummidiSAhujaSSGonzalezEAndersonSASantiagoEN 1998 Genealogy of the CCR5 locus and chemokine system gene variants associated with altered rates of HIV-1 disease progression. Nat Med 4 786 793
30. DuggalPAnPBeatyTHStrathdeeSAFarzadeganH 2003 Genetic influence of CXCR6 chemokine receptor alleles on PCP-mediated AIDS progression among African Americans. Genes Immun 4 245 250
31. LimouSCoulongesCHerbeckJTvan ManenDAnP 2010 Multiple-cohort genetic association study reveals CXCR6 as a new chemokine receptor involved in long-term nonprogression to AIDS. J Infect Dis 202 908 915
32. DaughertyBLSpringerMS 1997 The beta-chemokine receptor genes CCR1 (CMKBR1), CCR2 (CMKBR2), and CCR3 (CMKBR3) cluster within 285 kb on human chromosome 3p21. Genomics 41 294 295
33. ChoeHFarzanMSunYSullivanNRollinsB 1996 The beta-chemokine receptors CCR3 and CCR5 facilitate infection by primary HIV-1 isolates. Cell 85 1135 1148
34. ChoeHFarzanMKonkelMMartinKSunY 1998 The orphan seven-transmembrane receptor apj supports the entry of primary T-cell-line-tropic and dualtropic human immunodeficiency virus type 1. J Virol 72 6113 6118
35. ScarlattiGTresoldiEBjorndalAFredrikssonRColognesiC 1997 In vivo evolution of HIV-1 co-receptor usage and sensitivity to chemokine-mediated suppression. Nat Med 3 1259 1265
36. NedellecRCoetzerMShimizuNHoshinoHPolonisVR 2009 Virus entry via the alternative coreceptors CCR3 and FPRL1 differs by human immunodeficiency virus type 1 subtype. J Virol 83 8353 8363
37. LussoP 2006 HIV and the chemokine system: 10 years later. EMBO J 25 447 456
38. XiaoLRudolphDLOwenSMSpiraTJLalRB 1998 Adaptation to promiscuous usage of CC and CXC-chemokine coreceptors in vivo correlates with HIV-1 disease progression. AIDS 12 F137 143
39. ConnorRISheridanKECeradiniDChoeSLandauNR 1997 Change in coreceptor use coreceptor use correlates with disease progression in HIV-1-infected individuals. J Exp Med 185 621 628
40. ShimizuNTanakaAOueAMoriTOhtsukiT 2009 Broad usage spectrum of G protein-coupled receptors as coreceptors by primary isolates of HIV. AIDS 23 761 769
41. GorryPRDunfeeRLMeffordMEKunstmanKMorganT 2007 Changes in the V3 region of gp120 contribute to unusually broad coreceptor usage of an HIV-1 isolate from a CCR5 Delta32 heterozygote. Virology 362 163 178
42. CilliersTWilleySSullivanWMPatienceTPugachP 2005 Use of alternate coreceptors on primary cells by two HIV-1 isolates. Virology 339 136 144
43. CarringtonMDeanMMartinMPO'BrienSJ 1999 Genetics of HIV-1 infection: chemokine receptor CCR5 polymorphism and its consequences. Hum Mol Genet 8 1939 1945
44. HendelHWinklerCAnPRoemer-BinnsENelsonG 2001 Validation of genetic case-control studies in AIDS and application to the CX3CR1 polymorphism. J Acquir Immune Defic Syndr 26 507 511
45. ThomsonGBarcellosLFValdesAM 2008 Searching for additional disease loci in a genomic region. Adv Genet 60 253 292
46. CordellHJClaytonDG 2002 A unified stepwise regression procedure for evaluating the relative effects of polymorphisms within a gene using case/control or family data: application to HLA in type 1 diabetes. Am J Hum Genet 70 124 141
47. HolmS 1979 A Simple Sequentially Rejective Bonferroni Test Procedure. Scand J Stat 6 65 70
48. HarrellFEJrLeeKLMarkDB 1996 Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat Med 15 361 387
49. YoshimuraTOppenheimJJ 2008 Chemerin reveals its chimeric nature. J Exp Med 205 2187 2190
50. AraiMMitsukeHIkedaMXiaJXKikuchiT 2004 ConPred II: a consensus prediction method for obtaining transmembrane topology models with high reliability. Nucleic Acids Res 32 W390 393
51. LeickMCatusseJFolloMNibbsRJHartmannTN 2009 CCL19 is a specific ligand of the constitutively recycling atypical human chemokine receptor CRAM-B. Immunology 129 536 546
52. ZabelBANakaeSZunigaLKimJYOhyamaT 2008 Mast cell-expressed orphan receptor CCRL2 binds chemerin and is required for optimal induction of IgE-mediated passive cutaneous anaphylaxis. J Exp Med 205 2207 2220
53. LeeSTiffanyHLKingLMurphyPMGoldingH 2000 CCR8 on human thymocytes functions as a human immunodeficiency virus type 1 coreceptor. J Virol 74 6946 6952
54. GalliganCLMatsuyamaWMatsukawaAMizutaHHodgeDR 2004 Up-regulated expression and activation of the orphan chemokine receptor, CCRL2, in rheumatoid arthritis. Arthritis Rheum 50 1806 1814
55. ThomasCFJrLimperAH 2004 Pneumocystis Pneumonia. N Engl J Med 350 2487 2498
56. HartmannTNLeickMEwersSDiefenbacherASchraufstatterI 2008 Human B cells express the orphan chemokine receptor CRAM-A/B in a maturation-stage-dependent and CCL5-modulated manner. Immunology 125 252 262
57. OostendorpJHylkemaMNLuingeMGeerlingsMMeursH 2004 Localization and enhanced mRNA expression of the orphan chemokine receptor L-CCR in the lung in a murine model of ovalbumin-induced airway inflammation. J Histochem Cytochem 52 401 410
58. OteroKVecchiAHirschEKearleyJVermiW 2010 Non-redundant role of CCRL2 in lung dendritic cell trafficking. Blood 116 2942 2949
59. PhairJJacobsonLDetelsRRinaldoCSaahA 1992 Acquired immune deficiency syndrome occurring within 5 years of infection with human immunodeficiency virus type-1: the Multicenter AIDS Cohort Study. J Acquir Immune Defic Syndr 5 490 496
60. BuchbinderSPKatzMHHessolNAO'MalleyPMHolmbergSD 1994 Long-term HIV-1 infection without immunologic progression. AIDS 8 1123 1128
61. VlahovDGrahamNHooverDFlynnCBartlettJG 1998 Prognostic indicators for AIDS and infectious disease death in HIV-infected injection drug users: plasma viral load and CD4+ cell count. JAMA 279 35 40
62. HilgartnerMWDonfieldSMWilloughbyAContantCFJrEvattBL 1993 Hemophilia growth and development study. Design, methods, and entry data. Am J Pediatr Hematol Oncol 15 208 218
63. GoedertJJKesslerCMAledortLMBiggarRJAndesWA 1989 A prospective study of human immunodeficiency virus type 1 infection and the development of AIDS in subjects with hemophilia. N Engl J Med 321 1141 1148
64. CelentanoDDVlahovDCohnSShadleVMObasanjoO 1998 Self-reported antiretroviral therapy in injection drug users. JAMA 280 544 546
65. FaureSMeyerLCostagliolaDVaneensbergheCGeninE 2000 Rapid progression to AIDS in HIV+ individuals with a structural variant of the chemokine receptor CX3CR1. Science 287 2274 2277
66. Centers for Disease Control 1987 Revision of the CDC surveillance case definition for acquired immunodeficiency syndrome. Council of State and Territorial Epidemiologists; AIDS Program, Center for Infectious Diseases. MMWR Morb Mortal Wkly Rep 36 Suppl 1 1S 15S
67. KaslowRACarringtonMAppleRParkLMunozA 1996 Influence of combinations of human major histocompatibility complex genes on the course of HIV-1 infection. Nat Med 2 405 411
68. GaoXNelsonGWKarackiPMartinMPPhairJ 2001 Effect of a single amino acid change in MHC class I molecules on the rate of progression to AIDS. N Engl J Med 344 1668 1675
69. ShtatlandESKleinmanKCainEM 2005 Model Building in PROC PHREG with Automatic Variable Selection and Information Criteria. 1 10 Proceedings of the 30th Annual SAS Users Group International Conference Paper 206-30
70. PriceALPattersonNJPlengeRMWeinblattMEShadickNA 2006 Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38 904 909
71. TroyerJLNelsonGWLautenbergerJAChinnLMcIntoshC 2011 Genome-wide association study implicates PARD3B-based AIDS restriction. J Infect Dis 203 1491 1502
72. AllisonPD 1995 Survival Analysis Using the SAS System: A Practical Guide Cary, NC SAS Institute Inc
73. LewontinRC 1964 The Interaction of Selection and Linkage. II. Optimum Models. Genetics 50 757 782
74. HungLHNganSCLiuTSamudralaR 2005 PROTINFO: new algorithms for enhanced protein structure predictions. Nucleic Acids Res 33 W77 80
75. HungLHSamudralaR 2003 PROTINFO: Secondary and tertiary protein structure prediction. Nucleic Acids Res 31 3296 3299
Štítky
Genetika Reprodukční medicína
Článek Macroautophagy Is Regulated by the UPR–Mediator CHOP and Accentuates the Phenotype of SBMA MiceČlánek Dynamic Replacement of Histone H3 Variants Reprograms Epigenetic Marks in Early Mouse EmbryosČlánek Mutations in a Guanylate Cyclase GCY-35/GCY-36 Modify Bardet-Biedl Syndrome–Associated Phenotypes in
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2011 Číslo 10- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- Transcriptional Robustness Complements Nonsense-Mediated Decay in Humans
- Identification, Replication, and Fine-Mapping of Loci Associated with Adult Height in Individuals of African Ancestry
- Genetic Determinants of Serum Testosterone Concentrations in Men
- A One Base Pair Deletion in the Canine Gene Causes Exon Skipping and Late-Onset Neuronal Ceroid Lipofuscinosis in the Tibetan Terrier
- Three Structure-Selective Endonucleases Are Essential in the Absence of BLM Helicase in
- Identification of Widespread Ultra-Edited Human RNAs
- Multiple Wnts Redundantly Control Polarity Orientation in Epithelial Stem Cells
- The Bicoid Stability Factor Controls Polyadenylation and Expression of Specific Mitochondrial mRNAs in
- Transcriptome-Wide Binding Sites for Components of the Non-Poly(A) Termination Pathway: Nrd1, Nab3, and Sen1
- Macroautophagy Is Regulated by the UPR–Mediator CHOP and Accentuates the Phenotype of SBMA Mice
- Genetic Rearrangements Can Modify Chromatin Features at Epialleles
- Novel Function of as a Gap Gene during Spider Segmentation
- A Genome-Wide Screen for Interactions Reveals a New Locus on 4p15 Modifying the Effect of Waist-to-Hip Ratio on Total Cholesterol
- Comparative Genomic Analysis of Human Fungal Pathogens Causing Paracoccidioidomycosis
- Genetic Diversity in Cytokines Associated with Immune Variation and Resistance to Multiple Pathogens in a Natural Rodent Population
- Mutator Suppression and Escape from Replication Error–Induced Extinction in Yeast
- Dynamic Replacement of Histone H3 Variants Reprograms Epigenetic Marks in Early Mouse Embryos
- A Barcode Screen for Epigenetic Regulators Reveals a Role for the NuB4/HAT-B Histone Acetyltransferase Complex in Histone Turnover
- HIF–VEGF Pathways Are Critical for Chronic Otitis Media in and Mouse Mutants
- A Conserved Developmental Patterning Network Produces Quantitatively Different Output in Multiple Species of Drosophila
- Role of Exonic Variation in Chemokine Receptor Genes on AIDS: Association with Pneumocystis Pneumonia
- Whole-Exome Sequencing Identifies Homozygous Mutations in a Spastic Ataxia-Neuropathy Syndrome Linked to Mitochondrial -AAA Proteases
- Von Hippel-Lindau () Inactivation in Sporadic Clear Cell Renal Cancer: Associations with Germline Polymorphisms and Etiologic Risk Factors
- A Systems Biology Approach Reveals the Role of a Novel Methyltransferase in Response to Chemical Stress and Lipid Homeostasis
- Identification of Genomic Regions Associated with Phenotypic Variation between Dog Breeds using Selection Mapping
- Global Mapping of Cell Type–Specific Open Chromatin by FAIRE-seq Reveals the Regulatory Role of the NFI Family in Adipocyte Differentiation
- Natural Selection Affects Multiple Aspects of Genetic Variation at Putatively Neutral Sites across the Human Genome
- MicroRNA Expression and Regulation in Human, Chimpanzee, and Macaque Brains
- An Adaptive Allelic Series Featuring Complex Gene Rearrangements
- Feed-Forward Microprocessing and Splicing Activities at a MicroRNA–Containing Intron
- Developmental Stability: A Major Role for in
- A Phenomics-Based Strategy Identifies Loci on , , and Associated with Metabolic Syndrome Phenotype Domains
- Association of , , , , and with Systemic Lupus Erythematosus
- Small RNAs Prevent Transcription-Coupled Loss of Histone H3 Lysine 9 Methylation in
- Successive Increases in the Resistance of to Viral Infection through a Transposon Insertion Followed by a Duplication
- Mutations in a Guanylate Cyclase GCY-35/GCY-36 Modify Bardet-Biedl Syndrome–Associated Phenotypes in
- The Glycobiome Reveals Mechanisms of Pentose and Hexose Co-Utilization in Bacteria
- Insights into Hox Protein Function from a Large Scale Combinatorial Analysis of Protein Domains
- Mutations Cause Seckel and Jawad Syndromes
- Zelda Binding in the Early Embryo Marks Regions Subsequently Activated at the Maternal-to-Zygotic Transition
- Temporal Coordination of Gene Networks by Zelda in the Early Embryo
- Genetic Interaction between MTMR2 and FIG4 Phospholipid Phosphatases Involved in Charcot-Marie-Tooth Neuropathies
- Oxr1 Is Essential for Protection against Oxidative Stress-Induced Neurodegeneration
- Transforming Growth Factor β Receptor Type 1 Is Essential for Female Reproductive Tract Integrity and Function
- Positional Cloning of a Type 2 Diabetes Quantitative Trait Locus; , a Negative Regulator of Insulin Secretion
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- The Glycobiome Reveals Mechanisms of Pentose and Hexose Co-Utilization in Bacteria
- Global Mapping of Cell Type–Specific Open Chromatin by FAIRE-seq Reveals the Regulatory Role of the NFI Family in Adipocyte Differentiation
- Genetic Determinants of Serum Testosterone Concentrations in Men
- MicroRNA Expression and Regulation in Human, Chimpanzee, and Macaque Brains
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



