-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
The Analysis of Mutant Alleles of Different Strength Reveals Multiple Functions of Topoisomerase 2 in Regulation of Chromosome Structure
Type II topoisomerases (Topo II) are enzymes that disentangle DNA molecules during essential cellular processes such as DNA replication, chromosome condensation and mitotic cell division. Topo II is a major component of mitotic chromosomes and it is a well known target for cancer chemotherapy. Topo II inhibitors block the Topo II enzymatic activity leading to extensive DNA damage, which ultimately kills the cancer cell. Thus, investigating the role of Topo II in the assembly and structural maintenance of chromosomes is not only relevant to understand chromosome biology but might also have a translational impact on cancer therapy. Here we used Drosophila as model system to analyze the effect of Topo II depletion on chromosome stability. We show that the chromosomal phenotypes of mutant flies vary with the amount of residual Topo II, ranging from site-specific chromosome breaks, variations in chromosome number (aneuploidy and poliploidy) and dramatic defects in chromosome morphology. The chromosomal phenotypes observed in flies recapitulate all phenotypes seen in Topo II-depleted vertebrate chromosomes, reconciling the phenotypic discrepancies reported in previous studies. In addition, our finding that the Topo II dependent phenotypes vary with the residual amount of the enzyme provides useful information on the possible outcome of cancer therapy with Topo II inhibitors.
Published in the journal: . PLoS Genet 10(10): e32767. doi:10.1371/journal.pgen.1004739
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004739Summary
Type II topoisomerases (Topo II) are enzymes that disentangle DNA molecules during essential cellular processes such as DNA replication, chromosome condensation and mitotic cell division. Topo II is a major component of mitotic chromosomes and it is a well known target for cancer chemotherapy. Topo II inhibitors block the Topo II enzymatic activity leading to extensive DNA damage, which ultimately kills the cancer cell. Thus, investigating the role of Topo II in the assembly and structural maintenance of chromosomes is not only relevant to understand chromosome biology but might also have a translational impact on cancer therapy. Here we used Drosophila as model system to analyze the effect of Topo II depletion on chromosome stability. We show that the chromosomal phenotypes of mutant flies vary with the amount of residual Topo II, ranging from site-specific chromosome breaks, variations in chromosome number (aneuploidy and poliploidy) and dramatic defects in chromosome morphology. The chromosomal phenotypes observed in flies recapitulate all phenotypes seen in Topo II-depleted vertebrate chromosomes, reconciling the phenotypic discrepancies reported in previous studies. In addition, our finding that the Topo II dependent phenotypes vary with the residual amount of the enzyme provides useful information on the possible outcome of cancer therapy with Topo II inhibitors.
Introduction
Type II topoisomerases are large ATP-dependent homodimeric enzymes that transiently cleave double stranded DNA, pass a second DNA double helix through the break, and then reseal the break [1], [2]. In this way, Topo II enzymes solve a variety of topological problems that normally arise in double stranded DNA during processes such as replication, transcription, recombination and sister chromatid segregation [1], [2]. Topo II enzymes are structurally and functionally conserved, and the genomes of all eukaryotes harbor at least one Topo II enzyme. Vertebrates have two Topo II isoforms, alpha and beta [3]; these enzymes have identical catalytic activities but distinct localization patterns during mitosis. The beta isoform is primarily cytoplasmic, while most of Topo II alpha is concentrated in mitotic chromosomes [4]. In contrast, yeast and Drosophila have a single Topoisomerase II (Top2) gene. Notably, both the Drosophila Top2 and each of the human Topo II genes can rescue the phenotype of yeast Top2 mutants, highlighting the strong functional conservation of type II topoisomerases [5]–[7].
Topo II alpha is a major component of vertebrate mitotic chromosomes [8]–[10]. In vivo studies have shown that Topo II alpha has a dynamic behavior and that chromosome-associated Topo II alpha is rapidly exchanged with the cytoplasmic pool [4], [11], [12]. In fixed mitotic chromosomes, Topo II alpha exhibits a discontinuous localization pattern with Topo II alpha alternating with cohesin along chromatid axes [13], [14]. There is also evidence that in some systems Topo II alpha accumulates at centromeres in prometaphase and metaphase, suggesting a role of this enzyme in the regulation of centromere structure and/or cohesion [4], [12], [15]–[17].
Studies in yeast have shown that Top2 is not required for completion of DNA synthesis but plays essential roles in mitotic chromosome condensation and sister chromatid segregation. Failure to decatenate sister chromatids results in anaphase chromatin bridges that cause chromosome breakage during anaphase or cytokinesis [18]–[22]. Loss of Topo II activity does not affect S phase progression and disrupts sister chromatid separation also in vertebrate cells [2], [23]–[25]. However, the role of Topo II in vertebrate chromosome structure is rather controversial, possibly due to species-specific differences in chromosome organization and/or the different methods used to inhibit Topo II function (chemical inhibitors, immunodepletion, mutations or RNAi). For example, treatment of Indian muntjac cells with the Topo II inhibitor ICRF-193 caused frequent failures in sister chromatid individualization, the process by which duplicated DNA is resolved into two distinct chromatids [26]. In contrast, Topo II inhibition with ICRF-193 did not affect sister chromatid resolution in both Chinese hamster ovary cells (CHO) and baby hamster kidney (BHK) cells [27]. Moreover, Topo II alpha inhibition in different vertebrate systems resulted in a variety of chromosome morphologies ranging from relatively mild effects on the axial compaction of chromosomes to severe defects in chromosome condensation [14], [24], [28]–[32].
Although Topo II beta is not normally able to compensate for Topo II alpha loss, overexpression of Topo II beta in human cells can correct the defects caused by Topo II alpha depletion [33]. In addition, it has been shown that DT40 avian cells and human cells depleted of both Topo II alpha and Topo II beta display chromosomal defects more severe than those observed in cells lacking Topo II alpha alone [17], [24], [25], [32]. These results suggest that the two Topo II isoforms are partially redundant. Thus, studies of cells depleted of both Topo II alpha and Topo II beta are particularly relevant to define the role of Topo II in the maintenance of proper chromosomal architecture. An analysis of DT40 avian cells conditionally depleted of both Topo II isoforms showed that they exhibit extensive anaphase chromatin bridges, defective cytokinesis and polyploid cells [24]. Similar defects were observed in human cells lacking both Topo II alpha and Topo II beta [17], [25], [32]. In addition, cytological analysis showed that the latter cells exhibit severe defects in chromosome structure ranging from chromosome entangling to disrupted chromosome morphology [25].
Another controversial issue is the existence of a decatenation checkpoint triggered by loss of Topo II activity. The existence of such a checkpoint was suggested by studies in human cells showing that catalytic inhibitors of Topo II such as ICRF-187 or ICFR-193 are able to induce a caffeine-sensitive G2 delay that is dependent on ATR and BRCA1, but apparently independent of the DNA damage checkpoint [34]–[36]. However, a decatenation checkpoint is not present in both S. cerevisiae and S. pombe, in which Top 2 mutations cause minimal cell cycle delays [2], . Recent RNAi-based studies have also shown that depletion of Topo II alpha alone or both Topo II alpha and II beta does not trigger a decatenation checkpoint in vertebrate cells [24], [25]. In addition, no G2 delay was observed in Topo II-depleted vertebrate cells that were also treated with ICRF-193 [24], [38]. In contrast, expression of certain mutant forms of Top2 resulted in a G2 arrest in budding yeast cells [22], [37]. Collectively, these results indicate that loss of Topo II, and thus DNA catenation per se, is not able to induce a cell cycle delay and that a G2 checkpoint is instead activated by specific DNA lesions caused by catalytically inactive forms of Topo II.
Several studies have addressed the role of Top2 in Drosophila. Early work showed that injection of anti-Top2 antibodies or Top2 inhibitors into live Drosophila embryos result in strong defects in chromosome condensation and sister chromatid segregation at anaphase [39]. Two RNAi-based studies on S2 tissue culture cells showed that Top2-depleted fixed cells exhibit defects in longitudinal compaction of chromosomes, defective sister chromatid segregation and extensive anaphase bridges [40], [41]. Another study performed on live S2 cells, in which Top2 activity was reduced by either RNAi or chemical inhibition, did not detect defects in chromosome condensation and suggested that Top2 is required for centromere resolution and to prevent incorrect microtubule-kinetochore attachment [42]. Remarkably, downregulation of Top2 did not affect the mitotic index, indicating that Top2 deficiency does not activate cell cycle checkpoints in S2 cells [40], [42]. Other studies suggested that Drosophila Top2 is involved in homolog pairing in cell cultures [43], modulation of insulator function [44], and regulation of polytene chromosome structure [45].
Surprisingly, the role of Top2 in the maintenance of mitotic and meiotic chromosome structure in living flies has never been investigated. In a previous study we identified viable mutants in the solofuso (suo) gene. suo1 and suo2 mutant spermatocytes exhibit severely defective ana-telophases with extensive chromatin bridges; these telophases give rise to achromosomal secondary spermatocytes that are able to assemble bipolar spindles and divide in the complete absence of chromosomes [46]. Here we show that suo1 and suo2 are weak mutant alleles of the Top2 gene and describe the isolation and characterization of a stronger Top2 mutation. We show that modulation of Top2 function by means of these mutant alleles and in vivo RNAi results in multiple cytological phenotypes including site-specific chromosome aberrations, heterochromatin undercondensation, polyploidy, and complete disruption of chromosome morphology accompanied by a cell cycle arrest. These phenotypes recapitulate most of the phenotypes observed in vertebrate cells and indicate that Drosophila chromosomes are exquisitely sensitive to the residual level of Top2 in the cell.
Results
Isolation and characterization of Top2 mutations
In a previous study we identified and characterized suo1 and suo2, two viable but sterile mutants that exhibit many chromatin bridges and lagging chromosomes in male meiosis [46]. suo was originally mapped to the polytene interval 37C-37F5 uncovered by Df(2L)VA17 [46]. Using the Exelis kit of deficiencies, we restricted this interval to the polytene band 37E1, uncovered by Df(2L)Exel9043, which contains four genes: CG10237, RanGAP, Hs2st and Topoisomerase II (Top2). Complementation tests revealed that suo mutations are not allelic to either CG10237 or RanGAP. However, sequencing of Hs2st and Top2 in suo mutants did not reveal alterations in the protein coding exons of these genes with respect to those of another stock of the “Zuker collection” from which the suo mutants have been originally isolated (see Materials and Methods for detailed explanation). We thus screened a collection of 1,500 lethal mutations for those that fail to complement suo1 (see Materials and Methods). We isolated a lethal mutation that in combination with suo1 is semi-lethal and elicits the same meiotic phenotype observed in suo mutants (see below); this mutation was initially named suo3. DNA analysis of suo3 mutants revealed a G-A transition at nucleotide 1,040 of the Top2 coding sequence, corresponding to the 5′ splicing site of the second intron of the gene. This substitution results in a premature stop codon that would lead to a truncated form of the protein. These results indicate that suo3 is allelic to Top2. Thus, we renamed suo1, suo2 and suo3 as Top2suo1, Top2suo2 and Top2suo3, respectively. We note that we analyzed only the protein coding sequences of the Top2suo1 and Top2suo2 mutants but not the introns the UTRs or the 5′ regulatory sequences. Thus, the molecular lesions resulting in these mutations remain to be determined.
As previously reported, both Top2suo1 and Top2suo2 homozygous flies are viable but sterile in both sexes [46]. Top2suo3 homozygous individuals die during embryonic development, but this early lethal phenotype is due to a second site mutation, as Top2suo3/Df(2L)Exel9043 (henceforth Top2suo3/Df) animals survive until an early third instar larval stage. Top2suo3/Df larvae are devoid of imaginal discs like strong mitotic mutants [47] and exhibit small brains compared to controls at the same stage. Top2suo2/Df individuals are viable but male and female sterile; Top2suo1/Df and Top2suo1/Top2suo3 mutant flies are semi-lethal, with a few escapers. Together these results allow the ordering of our Top2 mutations in an allelic series with Top2suo3>Top2suo1>Top2suo2.
To assess the amount of the Top2 protein in different mutant combinations, we performed a Western blot analysis on larval extracts using a polyclonal antibody directed against Drosophila Top2 [48]. In wild type extracts, this antibody recognized a single band of the expected size (170 kDa). The intensity of this band was reduced to approximately 80% of the control level in extracts from Top2suo1 homozygous brains, showed further reduction in extracts from both Top2suo1/Df and Top2suo1/Top2suo3 larvae, and was virtually undetectable in Top2suo3/Df extracts (Figure 1A, B). Immunostaining with an anti-Top2 antibody [49] showed that Top2 is enriched in interphase nuclei and mitotic chromosomes. In particularly favorable preparations we could observe a discontinuous Top2 distribution on chromosomes with alternating stained and unstained regions and no specific centromeric enrichments (Figure 1C–E). This staining pattern is consistent with that observed in vertebrate chromosomes in which the Topo II negative regions are more enriched in condensins compared with the Topo II positive regions [13], [14]. An alternating condensin/Top2 immunostaining pattern has been previously observed also in Drosophila S2 cells, although it was less sharply defined than in vertebrate chromosomes [42], [50]. In Top2suo1/Df and Top2suo1/Top2suo3 brains immunostained for Top2, both mitotic chromosomes and interphase nuclei showed a weaker fluorescence compared with wild type (Figure 1F and G); the nuclei of Top2suo3/Df did not show any Top2 signal. These results agree with the genetic data and indicate that Top2suo1 and Top2suo2 are hypomorph alleles, while Top2suo3 is a very strong and potentially null allele.
Fig. 1. Top2 localization in wild type and Top2 mutant larval brain cells. 
(A) Western blotting of brain extracts showing that the anti-Top2 antibody recognizes a 170 kDa band in control cells; this band is reduced in Top2suo1/Top2suo1, Top2suo1/Df and Top2suo1/Top2suo3 mutants and undetectable in Top2suo3/Df. Tubulin was used as loading control. (B) Quantification of the relative intensities of the bands (± SD) from 4 different Western blotting experiments; the columns represent the relative ratios between the Top2 and the tubulin band intensities determined by densitometric analysis, with the wild type ratio set to 1. (C–E) Wild type male late prophase (C/C′), and female (D/D′) and male (E/E′) metaphases stained with the anti-Top2 antibody; note the discontinuous staining and the absence of Top2 accumulation at the kinetochores in D′ (arrowheads) and C′. In E′, the discontinuous staining is not visible but the centromeric regions of the major autosomes are clearly less stained than the other chromosomal regions. In Top2suo1/Df mutant brains (F/F′) and Top2suo1/Top2suo3 (G/G′) chromosome immunostaining is reduced compared to wild type. Scale Bar, 5 µm. The availability of flies expressing different levels of Top2 allowed us to examine the roles of this protein in a physiological context. We focused on the phenotypes elicited by Top2 mutants in three different Drosophila tissues: testes, brain and salivary glands.
Top2 is required for proper chromatin organization within the nuclei of prophase I spermatocytes
During prophase I, Drosophila spermatocytes organize their chromatin in 3 main distinct clusters that localize at the periphery of the nucleus. Each of these clusters corresponds to one of the major Drosophila bivalents (X-Y, 2-2 and 3-3); the small fourth chromosome bivalent sometimes is separated from the three major chromatin clumps but more often associates with the X-Y bivalent. Before meiosis, chromosomes become progressively individualized within each chromatin territory but bivalents remain well separated from each other until metaphase I, when they congress at the center of the cell [51]–[54]. Previous work has shown that chromatin clumps morphology is affected by mutations in the condensin II coding genes and in genes that specify proteins required for homologous paring of the achiasmatic male chromosomes [55]–[58].
We thus analyzed chromatin organization within the prophase nuclei of Top2suo1/Df primary spermatocytes. To compare mutants and wild type spermatocytes, we stained cells for DNA (with DAPI), tubulin and the centriole marker DSpd-2 [59], and used centriole length as an additional criterion for stage identification (Figure 2). Top2 spermatocytes at stages S2 and S3 were not very different from those of wild type (see ref [52] for stage description). However, at stage S4 they showed a higher number of distinct chromatin masses than wild type controls. This increase in chromatin masses was more evident in S5 spermatocytes; Top2suo1/Df spermatocyte nuclei displayed an average number of 5.6 chromatin masses per cell (n = 60) while control cells showed an average of 3.4 masses/nucleus (n = 60). Notably, in Top2 mutant spermatocytes at stages S4 and S5 chromatin clumps of similar size were often closely apposed, suggesting that homologs were unpaired but remained in the vicinity of one another (Figure 2). In the subsequent stages of Top2/Df spermatocyte growth, the number of chromatin clumps progressively reduced. At prometaphase (stage M1), mutant spermatocytes displayed 3 compact clumps like wild type cells (Figure 2, and ref [46]); at metaphase (stage M3) they showed a central chromatin mass like wild type controls [46]. Collectively, these results indicate that during the early stages of spermatocyte growth Top2 is required for homolog co-mingling in a single nuclear territory. However, at later stages of spermatocyte development the homologs appear to pair through a Top2-independent mechanism.
Fig. 2. Top2suo1/Df spermatocytes exhibit an abnormal chromatin distribution within the prophase nuclei. 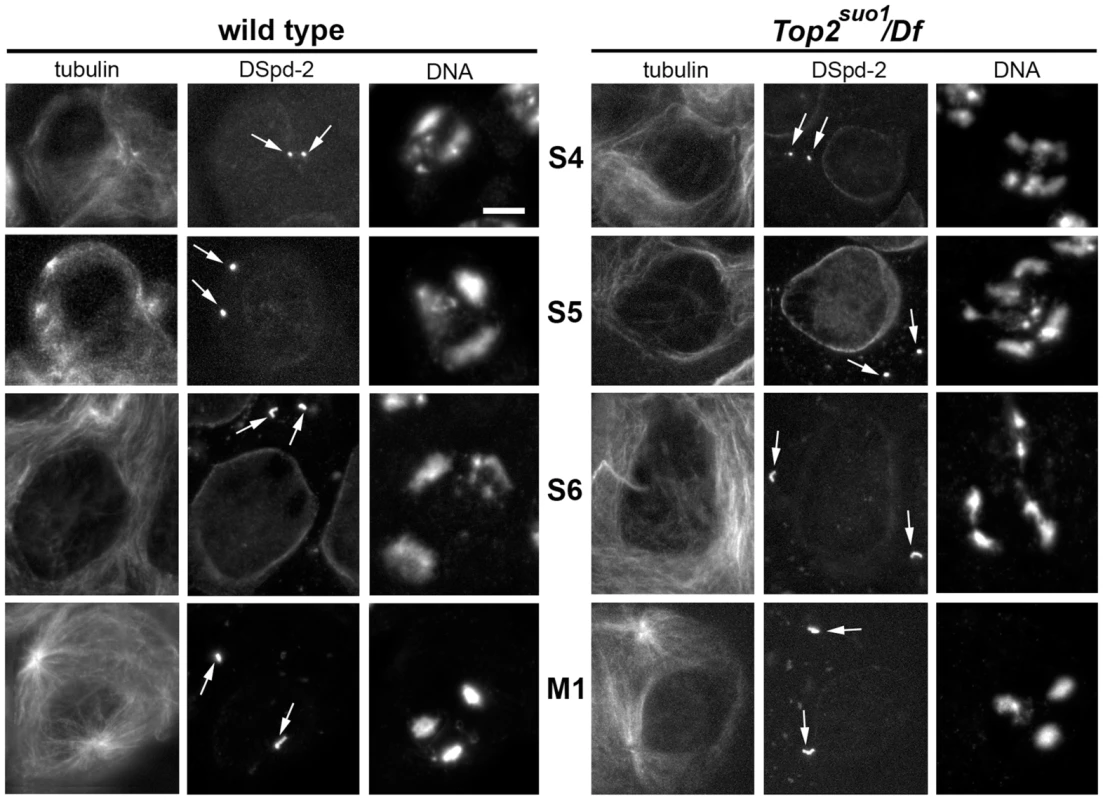
Primary spermatocytes were stained for DNA, tubulin and the centriole marker DSpd-2 (arrows). Centriole size and morphology facilitated stage (see ref [52]) recognition and comparison between Top2suo1/Df and wild type cells. Scale Bar, 10 µm. Top2 mutant males exhibit severe defects in meiotic chromosome structure and segregation
Previous studies characterized the meiotic phenotype of mutant males homozygous or hemizygous for Top2suo1 or Top2suo2 [46]. Here we examined the meiotic phenotype of flies bearing the Top2suo3 allele after staining for tubulin, DSpd-2 and DNA. In Top2suo3/Df testes, no meiotic divisions were found due a severe defect in germ cell proliferation. The meiotic phenotype of Top2suo1/Top2suo3 males was similar to that of Top2suo1/Df males, with all ana-telophase I figures characterized by highly defective chromosome segregation and chromatin bridges (Figure 3A, B). As previously described, in approximately one half of these aberrant ana-telophases, all chromosomes segregated to a single pole, giving rise to secondary spermatocytes that divided in the complete absence of chromosomes. Secondary spermatocytes that received some chromosomes also displayed extensive defects in chromosome segregation with frequent chromatin bridges (Figure 3A; see also ref [46]). These results demonstrate that Top2 is required for chromosome segregation during both meiotic divisions of Drosophila males, highlighting a crucial role of the protein not only in sister chromatid separation but also in bivalent segregation.
Fig. 3. Top2suo1/Top2suo3 mutant males display strong defects in meiotic chromosome morphology and segregation. 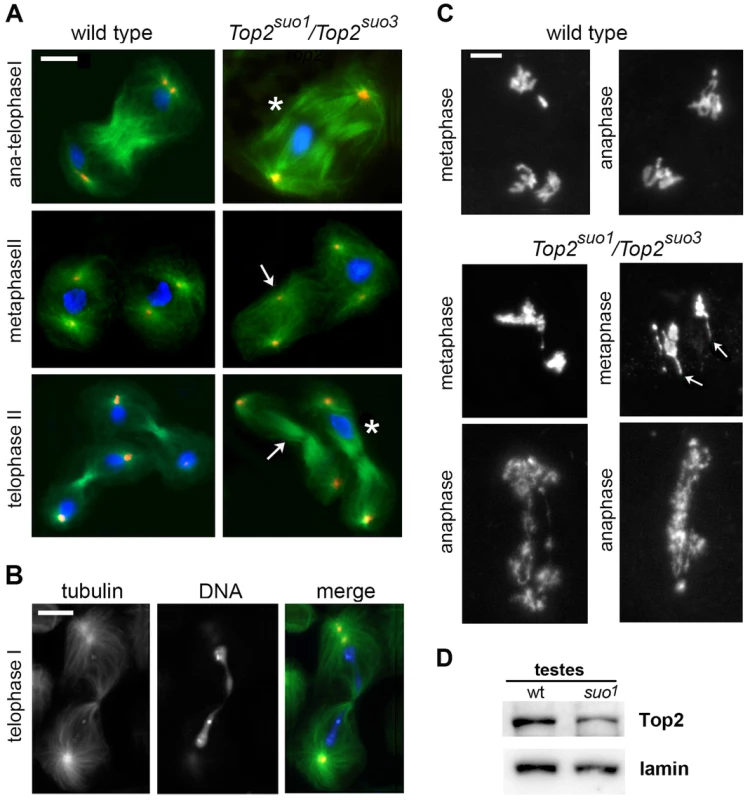
(A, B) Cells were fixed with methanol/acetone (see materials and methods) and stained for tubulin (green), Dspd-2 (red), and DNA (DAPI, blue). In the mutant telophases I and II marked with an asterisk (A), all chromosomes are segregating to one of the daughter cells; in another telophase I (B), a long chromatin bridge connects the two sets of segregating chromosomes. As a consequence of the aberrant chromosome segregation during the first meiotic division, many spindles of the second division are chromosome-free (arrows). Scale Bars, 10 µm. (C) Wild type and Top2suo1/Top2suo3 meiosis I figures fixed with 45% acetic acid and stained with DAPI. The Top2suo1/Top2suo3 metaphase chromosomes appear as chromatin masses in which the individual chromatids are no longer recognizable. Note the thin chromatin protrusions (arrows) in mutant metaphases, and the aberrant chromosome structure and segregation in mutant anaphases. Scale Bars, 5 µm. (D) Western blot showing that testes from Top2suo1/Top2suo1 males exhibit a reduction in the Top2 level with respect to wild type. Densitometric analysis of the bands from 3 different blots revealed that this reduction is of 50±5%. Lamin was used as loading control. Because the methanol-based fixation technique used to preserve spindle structure results in poorly defined chromosomes, we fixed larval testes in 45% acetic acid to maintain proper chromosome morphology (see Materials and Methods). The chromosomes of all metaphase and early anaphase I figures of Top2suo1/Df and Top2suo1/Top2suo3 spermatocytes displayed very severe structural defects. In metaphase figures, bivalents appeared as chromatin masses in which individual sister chromatids were no longer discernible (Figure 3C). Interestingly, approximately 60% of these aberrant metaphase figures (n = 18) showed one or more long chromatin protrusions that were not observed in wild type spermatocytes (Figure 3C). Similar protrusions have been observed in Top2-depleted meiotic cells of Drosophila females; these protrusions emanated from the chromatin mass of metaphase I figures and contained the centromeres at their tips, suggesting that they were generated by the pulling forces exerted by the spindle (Hughes and Hawley; cosubmitted). In anaphase-like figures, individual chromosomes were also no longer recognizable, and the two presumptive daughter cells were almost invariably connected by a number of entangled and irregularly condensed chromatin threads (Figure 3C). These entangled threads resulted in long ana-telophase chromatin bridges suggesting that the kinetochores of the entangled chromosomes are pulled away by the spindle microtubules (Figure 3B and 3C). The different appearance of metaphase and anaphase chromosomes is a likely consequence of the spindle pulling forces, which would stretch and partially disentangle the metaphase chromosomes. However, in approximately 50% of the cases the chromosome tangles generated by loss of Top2 function were not resolved and gave rise to telophase figures in which all chromosomes remained in one of the daughter cells (Figure 3A).
Top2 is required for mitotic chromosome integrity
To address the controversial role of Topo II in the maintenance of proper mitotic chromosome architecture, we first analyzed Top2suo1/Top2suo1, Top2suo1/Df and Top2suo1/Top2suo3 brains fixed without hypotonic and colchicine pre-treatments. In contrast with the spermatocyte chromosomes, the chromosomes of Top2suo1/Top2suo1 mutant brains were morphologically normal. Chromosome morphology was substantially normal also in Top2suo1/Df and Top2suo1/Top2suo3 mutant brains; we observed only a few cells (∼5%) showing a mild undercondensation of the heterochromatic regions of the major autosomes but the appearance of euchromatic arms was always normal (Figure 4). To determine whether the chromosomal phenotypes observed in spermatocytes and brain cells were due to different levels of Top2 we prepared testis extracts from Top2suo1/Top2suo1 mutants. Western blotting analysis showed these extracts exhibit a ∼50 reduction in the Top2 abundance. This reduction is comparable to that observed for Top2suo1/Df brains (∼44%; Figure 1A) in which the chromosome morphology is essentially normal. This suggests that spermatocyte and brain chromosomes require different levels of Top2 activity to achieve a regular structure, and that spermatocyte chromosomes are particularly sensitive to Top2 depletion.
Fig. 4. The site-specific CABs observed in Top2suo1/Top2suo3 mutant brains are generated by bridge severing during anaphase. 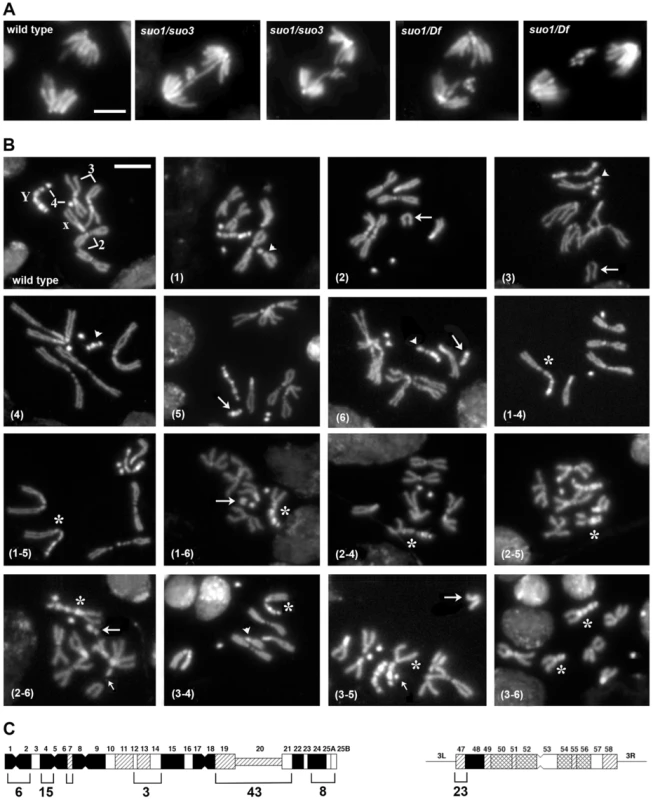
(A) Examples of wild type and Top2 mutant anaphase figures with bridges and lagging acentric fragments. Scale Bar, 5 µm. (B) Examples of aberrations generated by chromosome breaks that occurred during the anaphase of the previous cells cycle. Panels 1–3 show metaphases carrying a broken third chromosome without the corresponding acentric fragment (1), a complete chromosome complement with an extra acentric fragment (2), and a complete isochromatid deletion with a breakpoint in region h47 of 3L heterochromatin; note that at one of the ends of the acentric fragments shown in panels 2 and 3 the sister chromatids are closely apposed as occurs in the heterochromatic regions. The aberrant metaphases shown in panels 1–3 are likely to be generated by anaphase events leading to the G1 configurations depicted in Figure 5. Panels 4–6 show Y chromosome aberrations resulting from duplication of the broken G1 Ys depicted in Figure 5. The remaining panels show chromosome exchanges between the Y chromosome and the 3L heterochromatin generated by interactions between broken G1 chromosomes, as illustrated in Figure 5. For example the Y-3 dicentric chromosome shown in panel (1–4) is generated by fusion of the broken ends of the G1 chromosomes in the 1 and 4 “cells” of Figure 4. In all panels, arrowheads and arrows point to centric and acentric chromosome fragments, respectively; the asterisks indicate dicentric and translocation chromosomes involving the Y and 3L heterochromatin. The small arrows in panels (2–6) and (3–5) point to an additional break in 3L heterochromatin and an extra Y fragment, respectively. Scale Bar, 5 µm. (C) Distribution of 77 Y chromosome breakpoints and 23 third chromosome breakpoints of selected dicentrics and translocations. The heterochromatin maps are from Gatti and Pimpinelli [60]. The numbers above and below the maps indicate the different heterochromatin regions and the numbers of breakpoints found in these regions, respectively. In Top2suo1/Top2suo1 mutant brains fixed without colchicine and hypotonic pre-treatments, the mitotic index (MI, the proportion of cells engaged in mitosis; see Materials and Methods) and the frequency of anaphases were comparable to those of wild type brains (Table 1). Top2suo1/Df brains also showed a normal MI but displayed a significant increase in the frequency of anaphases with respect to both control and Top2suo1/Top2suo1 brains. This finding is likely to reflect a delay in progression through anaphase of mutant cells. In addition, 37% of the anaphases observed in Top2suo1/Df brains displayed chromatin bridges and/or acentric fragments that failed to correctly segregate (Figure 4 A). However, we never observed entire lagging chromosomes with unseparated sister chromatids. Thus, in Top2suo1/Df brains centromere resolution does not appear to be affected as occurs in Top2-depleted S2 cells [42].
Tab. 1. Mitotic parameters in <i>Top2</i> mutants. 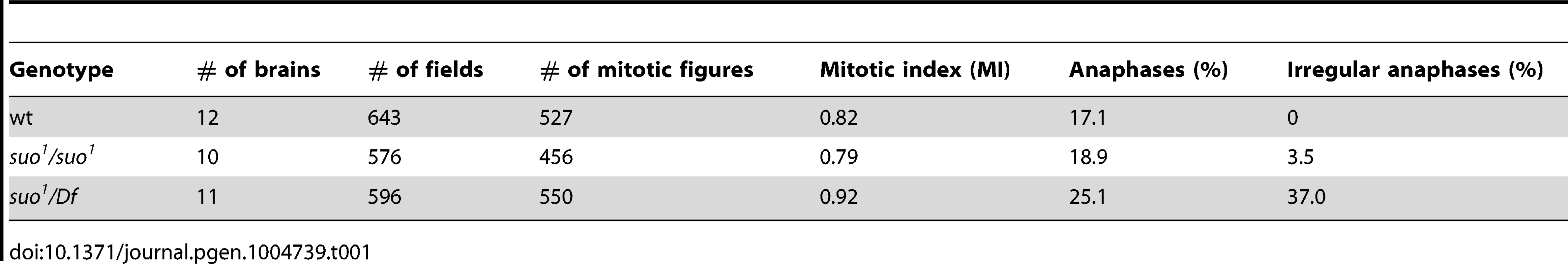
Top2 mutant brains fixed without colchicine and hypotonic pre-treatments also displayed several chromosome aberrations (CABs). To define the type and frequency of CABs elicited by the different Top2 mutant alleles we incubated dissected brains in saline with 10−5 M colchicine for 1 hour and treated them with hypotonic solution before fixation. Colchicine arrests mitotic cells in metaphase and hypotonic treatment results in chromosome spreading facilitating CAB scoring. In Top2suo2/Top2suo2 brains the CAB frequency was not significantly higher than that of wild type controls, which exhibit 0.008 CABs per cell (Table 2). Top2suo1/Top2suo1 mutant brains showed approximately 0.04 CABs/cell, while brains of both Top2suo1/Df and Top2suo1/Top2suo3 larvae displayed CAB frequencies ranging from 0.25 to 0.53 per cell (Figure 4B; Table 2). An analysis of the CABs observed in Top2suo1/Df and Top2suo1/Top2suo3 brain metaphases reveled a striking specificity. In mutant males, most CABs were isochromatid breaks that preferentially involved the entirely heterochromatic Y chromosome (48.3% of isochromatid breaks) and the third chromosome heterochromatin (46.1%). In addition, mutant males displayed 18–19% chromosome exchanges, most of which (∼98%) were dicentrics or translocations generated by breaks in the Y chromosome and 3L heterochromatin (Figure 4B). A similar CAB pattern was observed in XXY females in which 76% of the aberrations involved the Y chromosome, the third chromosome heterochromatin or both (Table 2). Examination of selected rearrangements displaying particularly clear heterochromatin banding revealed that the Y and third chromosome breakpoints are non-randomly distributed; the third chromosome breakpoints were clustered in region h47 of 3L heterochromatin, while the Y breakpoints were localized in several regions with a preference for regions h4-h5 and h19-h22 (Figure 4B and C; see ref [60] for a cytological map of Drosophila heterochromatin). In Top2suo1/Df and Top2suo1/Top2suo3 XX females most of the CABs were isochromatid breaks in region h47 of 3L heterochromatin. In contrast with males, these females did not display chromosome exchanges (Table 2). However, should these exchanges occur between the third chromosomes and involve the heterochromatic regions, they would be hard to detect as they would appear as intact chromosomes 3.
Tab. 2. Types and frequencies of chromosome aberrations (CABs) observed in Top2 mutant brains. 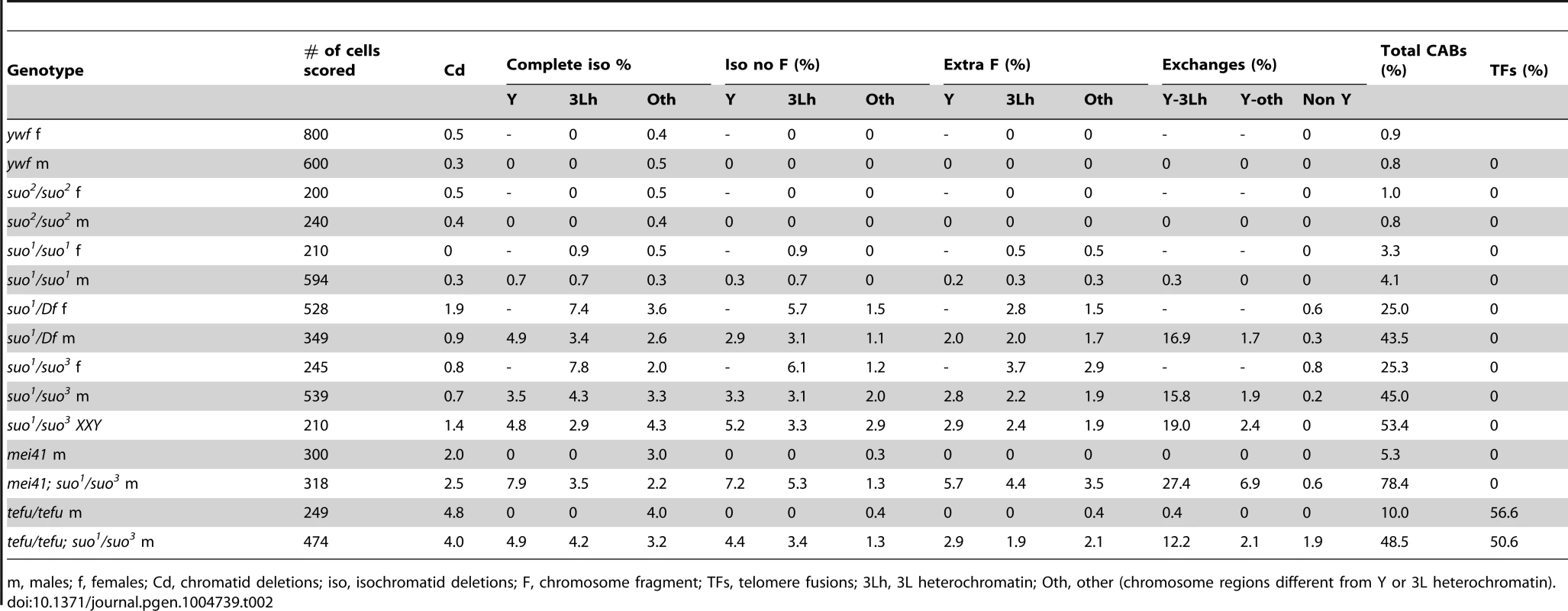
m, males; f, females; Cd, chromatid deletions; iso, isochromatid deletions; F, chromosome fragment; TFs, telomere fusions; 3Lh, 3L heterochromatin; Oth, other (chromosome regions different from Y or 3L heterochromatin). Importantly, in 65% of the mutant metaphases with a broken Y chromosome the acentric fragment was either missing or present as an additional element to a normal chromosome complement (the Y is easily recognized for its brightly fluorescent bands; see Figure 4). Similarly, 35% of the cells with a broken third chromosome showed only the centric portion of the chromosome. We also observed many cells (20%) with a normal chromosome complement and an extra acentric fragment of the size and the appearance of an entire 3L euchromatic arm. We thus assume that this acentric fragment is the exact complement of the centric element broken within 3L heterochromatin. Cells carrying either an isocromatid break lacking the acentric fragment or a normal chromosome complement plus an extra acentric fragment are likely to be generated during the anaphase of the previous cell cycle by the breakage of bridges between entangled chromatids.
The bridges observed in Top2 mutant anaphases are likely to involve sister rather than nonsister chromatids because the analysis of 100 anaphase figures revealed that the bridges, or the acentric fragments generated by their rupture, never involve a banded Y chromatid and a uniformly stained autosomal or XL chromatid (Figure 4A and S1). Thus, although we cannot formally exclude that some of the evenly stained bridges/fragments are generated by entanglements between nonsister chromatids, we assume most bridges observed in Top2 mutant anaphases are due to aberrant associations between the sister chromatids of either the Y or the third chromosome (Figures 4 and 5). As shown in Figure 4B (panels 1–6) and depicted in Figure 5, the acentric fragment resulting from the rupture of a bridge is subject to three different fates: it could be lost, co-segregate with its intact sister chromatid, or co-segregate with its complementary centric fragment.
Fig. 5. Schematic representation of the events leading to the chromosome rearrangements shown in Figure 3. 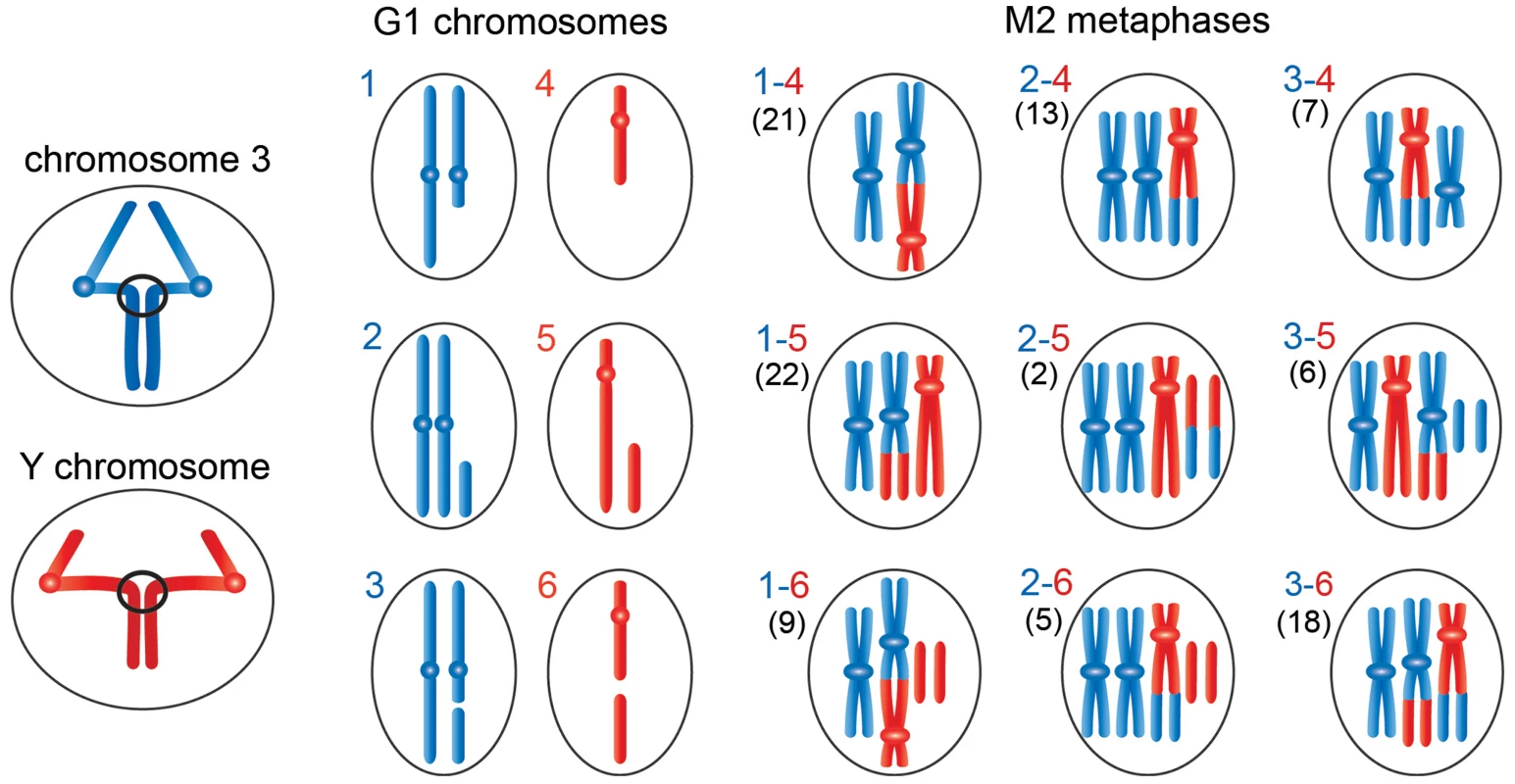
The panels on the left show third (blue) and Y (red) chromosomes undergoing anaphase with entangled sister chromatids. The G1 panels show the results of ruptures of the chromatin bridges generated by the anaphase entanglements; we considered breaks involving only one of the two chromosomes 3. During anaphase the acentric fragment could be lost (outcome in schemes 1 and 4), segregate at the cell pole that contains an intact chromosome complement (outcome in schemes 2 and 5), or segregate with the centric element of the broken chromosome (outcome in schemes 3 and 6). The other panels show the types of Y-3 exchanges that could be generated by the chromosome breaks depicted in panels 1–6. The exchanges generated by 1–4, 1–5, 2–4 or 2–5 interactions have unique configurations (see Figure 4); 1–6, 2–6, 3–4, 3–5 or 3–6 interactions can give rise to different type of exchanges. For example, 3–4 interactions can produce the half translocation depicted here and shown in Figure 4 or the dicentric chromosome shown Figure S2. Similarly, 3–6 interactions can produce the translocation shown here and in Figure 3 or the dicentric chromosome shown in figure S2. We classified 103 Y-3 exchanges; the numbers of the different types of rearrangements observed are reported between parentheses on the left of each diagram. The hypothesis that incomplete isochromatid breaks are generated during the anaphase of the previous cell cycle is supported by the analysis of 103 chromosome exchanges (selected on the basis of their cytological quality) involving the Y and 3L heterochromatin. Indeed, as shown in Figures 4 and 5, the majority (82%) of these rearrangements involve at least one element that can only be generated by the rupture of an anaphase bridge: either a centric fragment devoid of its acentric companion (panels 1 and 4 of Figures 4B and 5) or an acentric fragment accompanied by its intact homologue (panels 2 and 5 of Figures 4B and 5). Interactions between these elements and between them and the two complementary fragments of a broken chromosome (which are also likely to be generated during anaphase; see panels 3 and 6 of Figures 4B and 5) would result in the 9 types of chromosome rearrangements shown in figure 4B and depicted in Figure 5. In Figure 5, we depicted only the outcomes of single breaks in the Y and/or in one of the two third chromosomes. We did not consider the rearrangements generated by breaks in both chromosomes 3 or by two breaks in the Y chromosome. However, these complex CABs are rather rare and represent only 3% of the rearrangements involving the Y and/or the third chromosome; some examples of these complex rearrangements are shown in figure S2.
An analysis of the frequencies of the various types of rearrangements permits us to pinpoint the anaphase events that led to their formation. As shown in Figure 5, the rearrangements that include centric Y and third chromosome elements with no complementary acentric fragments are 41 and 52, respectively; those that contain intact Ys or third chromosomes plus extra fragments are 30 and 20, respectively; and those including centric Y and third chromosome elements plus the respective complementary fragments are 32 and 31, respectively. Given that the cells contain two chromosomes 3 and a single Y, this analysis establishes that the Y is more frequently involved in anaphase breaks than the third chromosome, consistent with the fact that the Y is breakable in multiple sites and the third chromosome in a single site. In addition, it appears that the most frequent anaphase event that will subsequently generate chromosome exchanges is the transmission to one of the two daughter cells of a centric element not accompanied by its complementary fragment. These findings are consistent with the observed frequencies of isochromatid breaks (Table 2). However, the relative frequencies of complete isochromatid breaks (centric element plus complementary fragment; Table 2) are slightly higher than those derived from the analysis of the exchanges. The simplest explanation for this discrepancy is that the anaphase events leading to the chromosomal configurations depicted in Figure 4 are not completely independent, so that the frequencies of double events is not the one expected based on independency of single events. In summary, our findings strongly suggest that most CABs observed in Top2suo1/Df and Top2suo1/Top2suo3 mutants derive from site-specific breaks generated during the anaphase of the previous cell cycle.
Top2 is required for cell cycle progression
While Top2suo1/Df and Top2suo1/Top2suo3 mutant cells appear to progress almost normally through mitosis despite the presence of CABs, Top2suo3/Df brains displayed a dramatic drop in the MI. In brains stained only with DAPI we were unable to observe clear mitotic figures. We thus analyzed Top2suo3/Df brains stained with DAPI and immunostained for both tubulin and the mitotic H3 phospho-histone, which marks mitotic chromatin [61]. In 25 brains examined, we observed only 10 metaphase-like figures (a single wild type brain usually contains 50–100 mitotic figures). The chromosomes of these rare metaphases appeared as chromatin masses in which individual chromosomes and sister chromatids were no longer recognizable (Figure 6). However, despite these severe chromosomal defects, all metaphases showed bipolar spindles with relatively normal microtubule densities. Growing evidence indicates that Drosophila somatic cells require kinetochore-driven MT growth for correct bipolar spindle formation [62], [63]. Thus, the observation that Top2suo3/Df cells can assemble a normal bipolar spindle suggests that Top2-depleted chromosomes/kinetochores retain the ability to drive MT nucleation.
Fig. 6. Examples of the rare mitotic metaphases observed in Topsuo3/Df mutants. 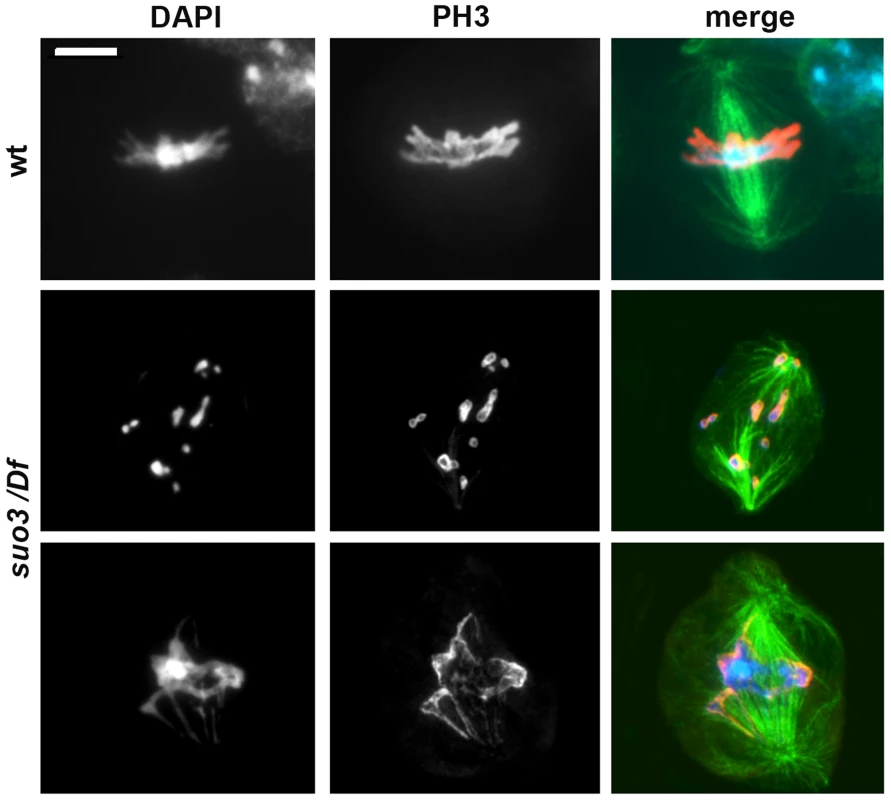
Cells were stained for DNA (DAPI, blue in merges), mitotic phosphohistone (red in merges) and tubulin (green in merges); wt, wild type. Scale Bar, 5 µm. The extremely low MI observed in Top2suo3/Df brain preparations might be the consequence of a checkpoint that prevents cells to progress through the cell cycle and enter mitosis. To ask whether the interphase arrest of Top2-deficient cells is caused by checkpoint activation we examined mei-41; Top2suo3/Df and Top2suo3/Df; tefu/tefu double mutants. mei-41 and tefu are the Drosophila orthologues of ATR and ATM, respectively; the kinases encoded by these genes are involved in the signaling pathway of the S and G2-M DNA damage checkpoints [64], [65], and ATR is thought to mediate the DNA decatenation checkpoint [35]. We found that neither double mutant displays a MI higher than that seen in Top2suo3/Df mutants (in all cases we examined more than 20 brains and observed less than one mitotic cell per brain). Thus, neither ATR nor ATM loss was able to rescue the block in mitotic progression in Top2suo3/Df brains.
To obtain further insight into the type of DNA lesions caused by Top2 deficiency we examined brains from mei-41; Top2suo1/Top2suo3 and Top2suo1/Top2suo3; tefu/tefu larvae. mei-41; Top2suo1/Top2suo3 brains showed a substantial increase in the frequency of CABs compared to either single mutant (Table 2). In doubly mutant males, the relative frequencies (with respect to the total number of CABs) of isochromatid deletions involving the Y chromosome or the third chromosome heterochromatin and the relative frequencies of Y-3 exchanges were comparable to those found in Top2suo1/Top2suo3 mutants. This suggests that downregulation of mei-41 (ATR) function might either increase the frequency of anaphase entanglements generated by Top2 deficiency or inhibit repair of the site-specific chromosome breaks they produce. Previous studies have shown that mutations in tefu (ATM) cause both telomeric fusions (TFs) and CABs [66]. Top2suo1/Top2suo3; tefu/tefu brains displayed frequencies of TFs and CABs that appear to be the sum of those observed in Top2suo1/Top2suo3 and tefu/tefu mutants. Thus, we conclude that tefu (ATM) and Top2 function in different pathways that mediate maintenance of chromosome integrity.
Analysis of the mitotic phenotype induced by in vivo Top2 RNA interference
The Top2 mutants analyzed here showed different mitotic phenotypes. Top2suo1/Df and Top2suo1/Top2suo3 brain cells showed a normal MI and frequent CABs that preferentially involve the Y and the 3L heterochromatin. In contrast, Top2suo3/Df brains showed a drastically reduced MI and rare metaphases with collapsed and/or shattered chromosomes. Top2suo1/Df and Top2suo1/Top2suo3 brains contain ∼60% less Top2 than wild type brains, and Top2 is undetectable in Top2suo3/Df brains (Figure 1A). This suggests that larval brains with intermediate Top2 levels would produce intermediate phenotypes that would provide additional information on Top2 function. Thus, we analyzed the phenotype produced by RNAi in flies bearing an inducible UAS-Top2 RNAi construct. Animals bearing this construct and an Actin-GAL4 driver died as third instar larvae; the brains of these larvae displayed a Top2 content that was barely detectable in Western blots but definitely higher than that of Top2suo3/Df larvae (Figure S3). Consistent with this finding, the mitotic phenotype of Top2 RNAi brains was milder than that observed in Top2suo3/Df brains. The frequency of mitotic figures in brains stained for DNA, mitotic phosphohistone and tubulin was still very low (an average of 7 mitotic figures per brain; 23 brains examined) but basic chromosome morphology was maintained. The analysis of chromosome preparations from colchicine-treated and acetic acid-fixed brains revealed a phenotype that was not previously observed in Top2suo1/Df or Top2suo3/Df mutants. Most metaphases were either polyploid (31%) or hyperploid (27%) and showed extensive chromosome breakage (Figure 7A and B). These metaphases showed many free autosomal arms generated either by breakage or drastic undercondensation of centric heterochromatin (Figure 7A). We estimated that the frequency of these free arms was approximately three-fold higher than the frequency of breaks in euchromatin. This finding, together with the fact that the Y chromosome was invariably broken in multiple fragments or rearranged, suggests that in Top2 RNAi cells most CABs involve heterochromatin.
Fig. 7. RNAi against Top2 disrupts mitotic division in larval brains. 
(A) Examples of hyperploid and polyploid cells found in Top2 RNAi brains. The panel in the upper left shows a hyperploid metaphase with tightly paired homologs; the other panels show polyploid metaphases with many broken chromosomes. Note that several breaks appear to involve the centric heterochromatin of the major autosomes; they could be either real interruptions in chromosome continuity or drastic failures in heterochromatin condensation. Scale Bar, 5 µm. (B) Frequencies of the various types of aberrant mitotic cells observed in Top2 RNAi brains (n = 164). The presence of hyperploid and polyploid metaphases in Top2-depleted brains suggests that these aberrant cells experienced nondisjunction events or complete cell division failures in the previous cell cycles. We also observed several prometaphase figures in which the homologous chromosomes were as tightly paired as in control cells (Figure 7A), suggesting that Top2 activity is not essential for somatic pairing in larval brain mitotic cells.
Analysis of polytene chromosomes in Top2 mutants
Examination of salivary glands revealed a role of Top2 in the control of polytene chromosome structure. In polytene preparations from Top2suo1/Df, Top2suo1/Top2suo3 and Top2suo3/Df males we consistently observed a specific alteration in the morphology of the X chromosome, which appeared bloated and often detached from the chromocenter (Figure 8 and Figure S4). In Top2suo1/Df and Top2suo1/Top2suo3 males, this deformed X chromosome conserved a banded appearance but its bands were less sharp than those of either the autosomes of the same nucleus or an X chromosome from wild type males. In general, the X chromosome condensation phenotype observed in Top2suo1/Df and Top2suo1/Top2suo3 mutants was rather variable but clearly visible in all polytene nuclei. A more severe defect was consistently observed in Top2suo3/Df and Top2 RNAi males, which showed a specific bloating and shortening of the X chromosome accompanied by a partial or total loss of its typical banding pattern (Figure 8 and Figure S4). In contrast, Top2suo1/Df, Top2suo1/Top2suo3 and Top2suo3/Df females displayed morphologically normal polytene chromosomes. These results indicate that Top2 is required for proper chromatin organization of the dosage-compensated X chromosome of males. We note that the defect in X chromosome organization and condensation observed in Top2suo3/Df and Top2 RNAi males is quite different from that caused by Cap-H2 (condensin) overexpression; an excess of Cap-H2 caused a tremendous axial compaction of all arms of polytene chromosomes [67], which was never observed in the X chromosome of Top2 mutants.
Fig. 8. Mutations in Top2 affect the X chromosome structure in male polytene nuclei. 
The polytene chromosomes of Top2suo1/Df males exhibit an X chromosome that maintains its banded appearance but is slightly bloated compared to wild type (wt). The polytene chromosomes of Top2suo3/Df and Top2 RNAi males display very similar defects; they show a specific bloating and shortening of the X chromosome accompanied by an almost total loss of its banded structure. Note that the X chromosome morphology in Iswi mutants is very different from that seen in Top2 mutants. Defects in the X chromosome condensation were previously observed in male polytene nuclei of Imitation switch (ISWI), Su(var)2-5 (HP1) and Su(var)3-7 mutants [68]–[70]. HP1 and Su(var)3-7 are interacting proteins with multiple roles in Drosophila chromatin regulation [71], [72]; ISWI is the ATPase subunit of several chromatin remodeling complexes including CHRAC, ACF and NURF [73]. Dosage compensation and the architecture of compensated chromatin depend on the Male Specific Lethal (MSL) complex, which includes Msl1, Msl2, Msl3, Mle, the Mof acetyltransferase and the roX1 and roX2 noncoding RNAs (see refs [74] and [75] for review). Previous studies have shown that blocking H4k16 acetylation completely rescues the X chromosome condensation defects caused by ISWI, Su(var)2-5 and Su(var)3-7 mutations [70], [76], suggesting that histone acetylation modulates chromatin compaction of the male X chromosome. Based on these results, we investigated the relationships between Top2 and the dosage compensation system. Previous work has shown that the X chromosome of Top2 mutants binds the Msl1 and Mle proteins [45]. Our immunostaining experiments showed that the bloated X chromosome of Top2 mutants also binds the dosage compensation factor Mof (Figure 9). In addition, we found that the bloated appearance of the X chromosome of Top2 mutants is completely rescued in mof; Top2 double mutants (Figure S5).
Fig. 9. The X chromosome of polytene nuclei from Top2 mutant males binds Mof. 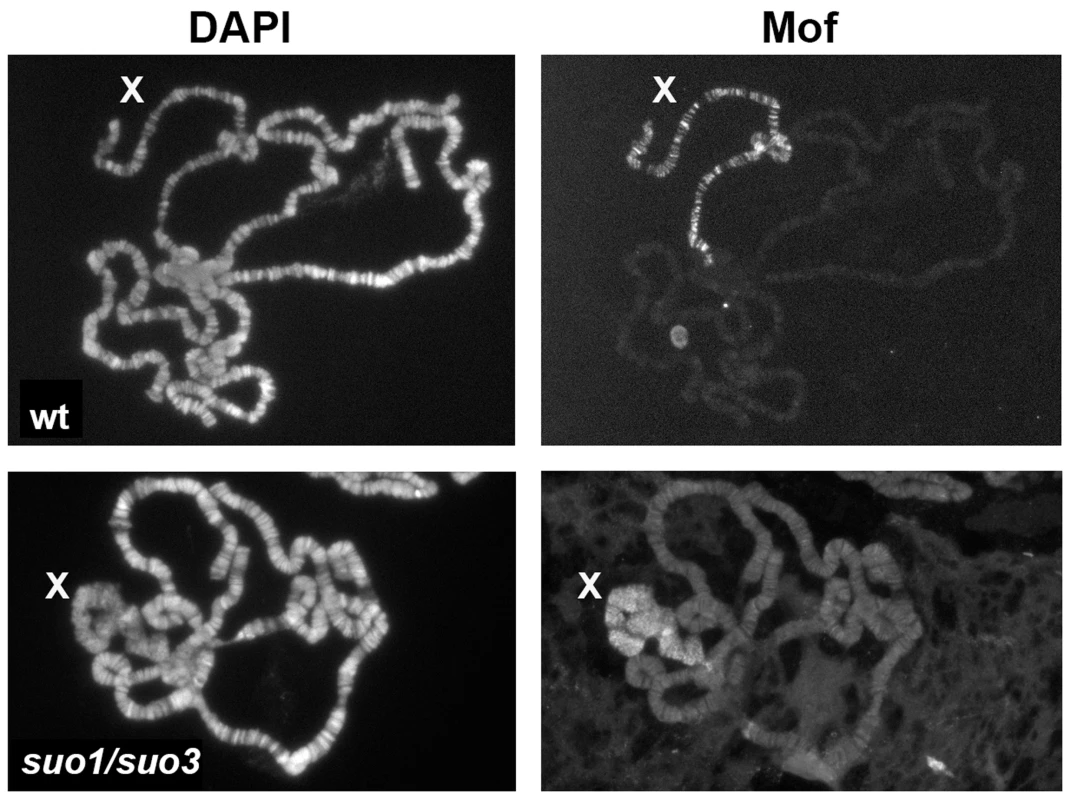
Note that Mof specifically decorates the X chromosome of both wild type and Top2suo1/Top2suo3 males; wt, wild type. Although mutations in ISWI and Top2 both affect X chromosome structure in male polytene nuclei, they do not result in identical phenotypes. A direct comparison between the polytene chromosomes of the mutants revealed that the X chromosome of ISWI mutants is shorter and more compact than those of either wild type or Top2 mutant larvae (Figure 8). Because previous studies have suggested that Top2 and ISWI are both contained in the CHRAC remodeling complex [77], we asked whether the male X chromosome deformation caused by Top2 mutations is a consequence of a failure to recruit ISWI at polytene chromosomes. Immunostaining of salivary glands from Top2suo1/Top2suo3 males showed that an anti-ISWI antibody decorates the X chromosome, although with a diffuse pattern reflecting the altered architecture of the chromosome (Figure 9). However, immunostaining of salivary glands from mof; Top2suo1/Top2suo3 doubly mutant males showed that the X with rescued morphology recruits a normal amount of ISWI despite the absence of Top2 (Figure S5). Thus, Top2 requirement for compensated chromatin modeling appears to be independent of ISWI localization.
Discussion
Top2 is required for chromatin organization within prophase nuclei of primary spermatocytes
We have shown that mutations in Top2 affect chromatin organization within the primary spermatocyte nuclei. During prophase I, wild type spermatocytes exhibit 3 main distinct chromatin clusters, which correspond to the major Drosophila bivalents (X-Y, 2-2 and 3-3); the small fourth chromosome bivalent is either separated from these chromatin masses or associated with the X-Y bivalent [51]–[54]. We found that at early growth stages (S1 and S2) the chromatin distribution within the spermatocyte nuclei of Top2suo1/Df males was not substantially different from wild type, suggesting that spermatogonial divisions are not severely affected. However, at stages S4 and S5 mutant spermatocytes displayed approximately twice as many chromatin masses as their wild type counterparts. In addition, in most mutant nuclei masses of similar size were closely apposed, suggesting a separation of the homologs within each chromatin territory. In the subsequent stages of spermatocyte growth, the number of chromatin masses in mutant nuclei progressively decreased, so that at prometaphase they displayed 3 compact chromatin clumps like their wild type counterparts.
Chromosome behavior during spermatocyte growth and male meiosis has been investigated in previous studies, which revealed a complex pairing mechanism [53]–[58],[78]. The X and the Y pair through their rDNA regions, while no specific euchromatic or heterochromatic pairing sites have been identified for the major autosomes, which are thus likely to exploit a homology-based pairing mechanism [53], [54], [78]. Tagging of allelic chromosome sites using the GFP-Lac repressor/lacO system or fluorescent in situ hybridization (FISH) showed that the major autosomes are tightly paired during the S1 and S2 stages. However, pairing is suddenly lost at the S2/S3 transition; the chromosomes remain then unpaired throughout the rest of meiosis but are included in a common nuclear territory until they condense prior to meiotic division [53], [54], [78]. One open problem is the mechanism underlying homolog co-mingling within the territories. Such co-mingling is unlikely to be the result of a canonical meiotic pairing, as the homologs remain uncondensed throughout prophase. It has been thus postulated that during early prophase the homologs might be held together in a single territory by chromatin entanglements [53], [54]. Our results are consistent with this idea and lead us to hypothesize that Topo II plays an active role in generating the entanglements that mediate homolog association. However, it is equally possible that Topo II is required for some kind of chromatin modifications that are important for homolog conjunction within the territories.
The chromatin organization defects within the prophase nuclei of Top2suo1/Df spermatocytes are very different from those previously observed in Cap-H2 and Cap-D3 mutants. In these mutants the chromatin remains diffuse within the spermatocyte nuclei from stage S4 through S6, indicating that the Cap-H2 and Cap-D3 condensin II subunits are required for the formation of the intranuclear territories that comprise the homologous chromosomes [58]. The chromatin organization defect in Top2suo1/Df spermatocytes it is also different from that caused by mutations in genes mediating achiasmate homolog pairing in Drosophila males (teflon, MNM and SNM). Spermatocytes of mutants in these genes display diffuse and slightly expanded chromatin territories during stages S4–S6; at prometaphase they show up to eight distinct chromatin clumps corresponding to unpaired univalents [54]–[57].
Collectively, the available results suggest that condensins (Cap-H2 and Cap-D3), the proteins required for homolog conjunction (teflon, MNM and SNM) and Top2 play distinct roles in chromatin organization during spermatocyte growth. As previously suggested, condensins are essential for territory formation and appear to function in opposition to homolog conjunction [58]. Top2, Teflon, MNM and SNM are all required for proper territory formation and homolog pairing. Top2 is primarily required for homolog conjunction and correct territory organization during stages S4–S6 of spermatocyte growth, whereas Teflon, MNM and SNM are primarily required for meiotic chromosome pairing during prometaphase and metaphase. However, Top2 might have a redundant role in metaphase chromosome pairing that would be masked by the activity of Teflon, MNM and SNM, which would be able to mediate homolog pairing even when Top2 is reduced. The finding that Top2 mediates an aspect of homologous chromosome pairing in males is intriguing, as this enzyme ensures proper biorientation of achiasmatic homologs in females (Hughes and Hawley; cosubmitted). Thus, despite the profound differences between Drosophila male and female meiosis, both types of meiotic divisions share a common Top2-depedent mechanism to facilitate achiasmate chromosome pairing.
Top2 is required for proper chromosome organization and segregation during Drosophila male meiosis
Previous studies have shown that loss of Topo II function results in species-specific meiotic defects. In top2 mutant cells of S. cerevisiae, premeiotic DNA synthesis, recombination and chromosome condensation are not affected but cells arrest at metaphase I and do not undergo the first meiotic division. However, top2 rad50 double mutants, in which recombination and synaptonemal complex formation are suppressed, perform the first meiotic division but not the second [79], [80]. A similar meiotic phenotype has been observed in Top2 mutant cells of S. pombe, which exhibit only a mild defect in the final steps of meiotic chromosome condensation and arrest at metaphase. This arrest is relieved by mutations in rec7 that strongly reduce recombination [81]. Thus, in both budding and fission yeast, Topo II has little or no role in chromosome condensation but its activity is required for segregation of recombinant chromosomes at meiosis I and, at least in S. cerevisiae, for sister chromatid separation at meiosis II.
Studies on meiotic cells from mouse and Chinese hamster injected with Topo II inhibitors did not reveal gross defects in chromosome condensation at the doses used in the experiments. However, the inhibitors induced a substantial meiotic delay and resulted in anaphase bridges and lagging chromosomes at both the first and the second meiotic anaphase [82]–[85]. It has been also suggested that the defect in homolog separation at meiosis I was due to a primary defect in chiasmata resolution [82]. In contrast, studies on mouse pachytene spermatocyte cultured in vitro showed that treatments with the Topo II inhibitors ICRF-193 and teniposide cause drastic defects in chromosome condensation. In teniposide-treated spermatocytes, both chromatin condensation and sister chromatid individualization were strongly affected. The effects of ICRF-193 were milder and some chromosomes managed to condense reaching a diplotene-like configuration [86].
We showed that weak Drosophila Top2 mutants (Top2suo1/Top2suo1 and Top2suo2/Df) with virtually no defects in brain cell mitoses exhibit strong defects in chromosome segregation during both meiotic divisions of males (this report and ref [46]). In addition, we have shown that in Top2suo1/Df and Top2suo1/Top2suo3 testes all meiotic divisions exhibit severe defects in chromosome structure and segregation. In most cells, the chromosomes formed amorphous metaphase I masses where the sister chromatids were no longer discernible. In addition, these chromatin masses often emanated protrusions that are likely to correspond to stretched pericentric regions. Despite the strong defect in chromosome structure, Drosophila spermatocytes did not arrest at metaphase like yeast cells [79] or mouse spermatocytes treated with teniposide [86]. This finding is consistent with the limited delay caused by the spindle checkpoint in Drosophila male meiosis and by the inability of this checkpoint to prevent spermatid formation and differentiation [87], [88]. Given that Drosophila male meiosis is achiasmatic, our observations strongly suggest that the aberrant meiotic chromosome segregation observed in Top2 mutant males is the consequence of a primary defect in chromatin folding within the chromosomes. As shown in Figure 3, this defect is particularly evident when the chromosomes are pulled away by the meiotic spindle. In the anaphase-like figures of Top2 mutants, the chromatids are not individualized and the spindle poles are connected by an irregular network of chromatin threads. This suggests that the chromatin fibers of both the homologous and the heterologous chromosomes as well as those of the sister chromatids remain trapped by multiple entanglements, which prevent correct chromosome segregation during both meiotic divisions.
The observation that Top2 is required for homolog conjunction during early meiotic prophase and then for correct chromosome segregation during anaphase is intriguing. We favor the hypothesis that Top2 has two independent activities, one required for catenation of the homologs within each chromosome territory and one required for proper chromatin folding within the metaphase chromosomes. However we cannot exclude the possibility that Top 2 activity during early prophase results in aberrant chromatin configurations that interfere with the chromatin folding processes leading to proper chromosome assembly.
Frequent anaphase bridges have been also observed in spermatocytes from mutants in the Cap-H2 and Cap-D3 genes. However, the morphology of the meiotic chromosomes of these mutants is not severely disrupted as occurs in Top2 mutants. Judging from the published micrographs, the chromatids of Cap-H2 and Cap-D3 mutants are clearly individualized and their appearance is not very different from that of their wild type counterparts [58]. Our data do not provide an explanation of why the meiotic chromosomes of Drosophila males are much more sensitive to Top2 depletion than brain chromosomes. We can only envisage that the two types of chromosomes contain different proteins and/or different numbers of high affinity Top2 binding sites. If these binding sites were more frequent in meiotic than in mitotic chromosomes, then a ∼50% reduction of the Top2 protein would be sufficient to disrupt meiotic chromosome organization in males, but would have only a limited effect on mitotic chromosomes of larval brains.
A meiotic phenotype reminiscent of that seen in Top2 mutant males has been observed after RNAi-mediated depletion of Top2 in females (Hughes and Hawley; cosubmitted). In wild type female meiosis, the heterochromatic regions of the homologous chromosomes remain connected during prometaphase I by chromatin threads that ensure proper biorientation of achiasmatic homologues; these homologous connections are then resolved at later stages of meiosis allowing chromosome segregation. In Top2 - depleted oocytes, heterochromatic regions of chromosomes usually fail to separate during prometaphase and metaphase I, and are often stretched into long protrusions with centromeres at their tips (Hughes and Hawley and references therein). These findings indicate that Top2 is required for resolution of the DNA entanglements that normally connect homologous heterochromatic regions during female meiosis and suggest that the pulling forces exerted by the spindle generate chromatin protrusions. However, the meiotic phenotypes elicited by Top2 depletion in males and females are similar but not identical. While in female meiosis only the heterochromatic regions appear to be affected, in male meiosis both euchromatin and heterochromatin are affected. A higher sensitivity of heterochromatin to Top2 depletion is consistent with our observations on Top2suo1/Df and Top2suo1/Top2suo3 mitotic cells, which exhibit chromosome breaks that preferentially involve the 3L and the Y heterochromatin.
Drosophila Top2 is required for maintenance of chromosome integrity
We have shown that relatively weak mutant combinations of Top2 alleles (Top2suo1/Df and Top2suo1/Top2suo3) only exhibit chromosome aberrations (CABs), most of which involve specific regions of the Y and third chromosome heterochromatin. Severe RNAi-mediated Top2 depletion results in extensive chromosome breakage involving all chromosome regions with a preference for heterochromatin. Previous studies with pharmacological inhibitors of Topo II have also shown that treatments with these drugs cause CABs, but it is currently unclear to which extent these drugs directly induce DNA lesions, cause DNA damage via Topo II inhibition, or affect DNA stability through other off target effects [89]. However, CABs have been also observed in Top2 mutants of budding and fission yeasts [18], [20],[21],[90] and in vertebrate cells depleted of Topo II by RNAi [17], [24], [25], [32], [91]. In both yeast and vertebrate systems, most CABs induced by Topo II deficiency are thought do be produced by breakage of the anaphase chromatin bridges generated by failure to decatenate sister chromatids [17], [25], [32], [90], [91].
We have shown that a relatively modest reduction of the Top2 level results in many isochromatid breaks and chromosome exchanges (translocations and dicentric chromosomes) that primarily involve 4 regions of the entirely heterochromatic Y chromosome (regions h1-2; h4-6, h19-21 and h24-25) and a specific region of the 3L heterochromatin (region h47). To the best of our knowledge, previous studies did not detect site-specific chromosome aberrations after inhibition of Topo II function. What is then the mechanism underlying the chromosome damage specificity in weak Top2 Drosophila mutants? Two observations help answering this question. First, in mutant brain cells not treated with colchicine, 37% of the anaphases displayed chromatin bridges or lagging chromosome fragments generated by severing of the bridges. Second, most CABs observed in colchicine-treated cells were “incomplete” chromosome type aberrations (i.e involving both sister chromatids). Namely, they consisted in broken centric chromosomes not accompanied by a corresponding acentric fragment, in normal chromosome complements with an additional acentric fragment, in Y-3 translocations lacking the reciprocal element, or Y-3 dicentric chromosomes lacking the acentric fragment. As illustrated in Figure 5, these aberrations are likely to be the consequence of chromosome breaks generated during the anaphase of the previous cell cycle. We propose that these breaks preferentially occur in chromosomal sites whose stability is particularly dependent on Top2-mediated DNA decatenation. In Top2 deficient cells, these sites would not be properly untangled and would break when the sister chromatids are pulled apart by the mitotic spindle. It has been shown that a prominent Top2 cleavage target is the 359 bp Drosophila satellite DNA, which is mainly found in the centric heterochromatin of the X chromosome [92]. We examined the extant maps of satellite DNA and transposable element distribution along Drosophila heterochromatin [93]–[95] but did not find any sequence that uniquely maps to the Top2-sensitive regions. Thus these regions might share an as yet unidentified DNA or might correspond to junctions between different DNA sequences (e. g. satellite-satellite; satellite transposon, or transposon-transposon).
Top2 RNAi brain cells contain very small amounts of Top2 and exhibit only a few divisions, most of which are hyperploid or polyploid. The few scorable diploid figures almost invariably displayed incomplete aberrations involving the Y or third chromosome heterochromatin, often accompanied by breaks in other chromosomes. We could not assess the presence of incomplete aberrations in polyploid metaphases, most of which displayed many apparent breaks of the centric heterochromatin of the major autosomes. These discontinuities in chromosome structure could be either due to drastic failures of heterochromatin condensation or to real breaks generated by the rupture of chromatin bridges during anaphase. Our results do not permit us to discriminate between these possibilities, but we favor the first.
The role of Top 2 in the control of larval brain cell cycle
Our observations on different Top2 mutant combinations and Top2 RNAi cells revealed different and apparently contradictory effects on cell cycle progression. In Top2suo1/Df brains that exhibit a ∼60% reduction in the wild type Top2 level, the MI was comparable to that of wild type controls, but mutant brains displayed an increase in the frequency of anaphases. These data are consistent with previous studies on Drosophila S2 cells showing that RNAi-mediated depletion of Top2 does not affect the MI and causes only a small increase in the anaphase frequency [40], [42]. The MI was not substantially affected also in DT40 and human cells depleted of both Topo II alpha and Topo II beta [24], [25], [32].
In contrast, in Top2 RNAi brains and Topsuo3/Df brains the MI was reduced by one and two orders of magnitude, respectively. Top2 RNAi brains also displayed many aneuploid and polyploid cells. Polyploidy has been also observed in chicken and human cells lacking Topo II activity, and was attributed either to defects in cytokinesis or to a reentry into interphase following a mitotic arrest (restitution) [24], [25]. The low MI and the extensive chromosome damage in RNAi brains did not allow us to reliably pinpoint the mechanism of polyploid cells formation. Polyploidy in Drosophila brains can be generated by either restitution or cytokinesis failure (see for example refs [47] and [63]). It is thus possible that the polyploid cells of Top2 RNAi brains were generated through both mechanisms.
The observations on weak Top2 mutants (Top2suo1/Df and Top2suo1/Top2suo3; this study) and Top2 RNAi S2 cells [40], [42] strongly suggest that Drosophila does not have a decatenation checkpoint that arrests cell cycle in response to loss of Top2 function (see Introduction). This conclusion agrees with recent data indicating that Topo II depletion and the resulting excess of DNA catenation does not trigger a G2 arrest in vertebrate cells [24], [25], [38], [96]. However, a catenation-independent but Topo II-dependent checkpoint is activated by interruptions of the decatenation process caused by catalytically inactive forms of Topo II [22], [37], [96].
We found that in Top2suo3/Df and Top2 RNAi brains the MI is drastically reduced, indicating that cells are blocked in interphase. We also showed that this block is not relieved by mutations in either mei-41 (ATR) or tefu (ATM). Because these kinases are involved in the signaling pathways that mediate most cell cycle checkpoints [64], [65], and because ATR has been previously implicated in the decatenation checkpoint [35], we believe that the interphase block observed in the nearly complete absence of Top2 is not due to the activation of a checkpoint. We instead believe that this block could depend on the failure to remove supercoils during DNA replication, which would cause extensive DNA damage and make the cell unable to sustain cell cycle progression.
Top2 is required for chromatin organization of the dosage-compensated X chromosome
We first reported that mutations in Top2 cause a specific alteration of the X chromosome morphology in male polytene chromosomes (Bonaccorsi et al., 50th Drosophila Research Conference; abstract 350b, 2009). This observation was confirmed and extended by Hohl and coworkers [45], who showed that in polytene nuclei of Top2 mutants the X chromosome retains the ability to recruit the MSL dosage compensation complex. In agreement with this study, we found that the bloated X chromosomes of Top2 mutants are decorated by anti-Mle, anti-Msl3 and anti-Mof antibodies. However, we have been unable to assess whether the staining intensity is the same as that of a normally condensed wild type X. Thus, it is quite possible that a reduction in Top2 expression partially affects the association of the MSL complex with the male X chromosome as recently suggested [97]. Regardless of the role of Top2 in recruitment and/or stabilization of the MSL complex, the observation that loss or inhibition of Top2 activity specifically disrupts the X chromosome morphology in males is fully consistent with the ChIP/Mass Spec experiments indicating that Top2 is the major MSL interactor [98] and with the co-IP assays showing that Top2 interacts with MSL through its Mle component [97].
Previous studies have shown that the X chromosome of polytene nuclei from Top2 mutants is decorated by antibodies against histone H4 acetylated at lysine 16 (H4K16ac). This post-translational modification is mediated by the Mof histone acetyltransferase, whose association with the X chromosome depends on Mle; a mutation in mle or blocking H4k16 acetylation rescues the X chromosome condensation defects caused by mutations in ISWI [76]. We found that the X chromosome phenotype elicited by mutations in Top2 is rescued in mof; Top2 double mutants, and that both the bloated X of Top2 mutants and the reconstituted X of mof; Top2 double mutants normally recruit the ISWI protein. These results suggest that loss of Top2 does not affect condensation of the dosage compensated chromatin by inhibiting ISWI recruitment. However, in the absence of Mof-mediated H4k16 acetylation the chromatin compaction functions of Top2 and ISWI are both dispensable. The genetic interaction between Top2 and mof is consistent with the previously shown physical and functional interactions between topoisomerase II and histone deacetylases (HDACs) [2], and with the synergistic cytotoxic effects caused by simultaneous inhibition of HDAC and Topo II [99], [100].
Conclusions
We have shown that meiotic chromosomes are extremely sensitive to Top2 depletion and exhibit drastic defects in chromosome morphology even in weak Top2 mutants. Top2 downregulation in brain mitotic cells by either mutations or in vivo RNAi produced different cytological phenotypes. Moderate Top2 depletion (Top2suo1/Df) did not affect chromosome structure, and produced site-specific chromosome aberrations generated by the rupture of anaphase bridges. Severe Top2 depletion (Top2 RNAi) strongly reduced the MI, and induced heterochromatin undercondensation, extensive chromosome breakage, aneuploidy and polyploidy. Finally, complete (or nearly complete) Top2 deficiency (Top2suo3/Df) caused an interphase block and disrupted chromatid individualization in the rare diving cells. These phenotypes indicate that Drosophila chromosomes are exquisitely sensitive to the residual level of Top2 in the cell. In addition, they recapitulate most, if not all, phenotypes previously observed in vertebrate cells exposed to Topo II inhibitors or RNAi against Topo II (see above). Thus, our results suggest that the previously observed discrepancies in vertebrate chromosome phenotypes elicited by Topo II downregulation might depend on the type of chromosomes examined (e.g. mitotic vs meiotic), slight differences in Topo II activity, or both.
Materials and Methods
Fly stocks and genetics
Top2suo1 and Top2suo2 mutant alleles were previously isolated by a cytological screen of a collection of male sterile mutants induced by EMS in C. Zuker laboratory [46], [101]. Top2suo3 was isolated from a collection of about 1,500 lines carrying lethal mutations on chromosome 2, arisen in the Zucker collection of viable mutants [101]. All mutations were kept in stock over the second chromosome balancer CyOTbA, bearing the Tubby1 (Tb1) dominant transgene [102]. Df(2L)Exel9043 was obtained from the Bloomington Drosophila Stock Center. mei-4129D [103] was kept in stock over an FM7-GFP balancer. The mei-4129D; Top2suo1/Top2suo3 and mei-4129D; Top2suo3/Df(2L)Exel9043 double mutants were obtained by crossing mei-4129D/FM7-GFP; Top2suo3/CyOTbA females to FM7-GFP/Y; Top2suo1/CyOTbA and to FM7-GFP/Y; Df(2L)Exel9043/CyOTbA males, respectively. Male larvae carrying both mutations were identified based on their non-GFP non-Tb phenotype. tefu3 (or atm3, a gift of S. D. Campbell; ref [104]) was kept in stock over the TM6C balancer carrying the Stubble (Sb) and Tb dominant markers. The Top2suo1/Top2suo3; tefu3/tefu3 double mutant was obtained by crossing Top2suo1/CyOGFP; tefu3/TM6C females to Top2suo3/CyOGFP; tefu3/TM6C males. Doubly mutant larvae were identified on the basis of their non-GFP, non-Tb phenotype. The mof1 mutant stock was kindly provided by J. Lucchesi. To construct mof; Top2 double mutants we crossed mof1/FM7-GFP; Top2suo1/CyOTbA females to FMT-GFP/Y; Top2suo3/CyOTbA males; non-GFP and non-Tb male larvae were then selected for cytological examination of polytene chromosomes. Iswi mutant larvae were generated by crossing y w; Iswi2 sp; +/T(2;3)B3 CyO, TM6B females to Iswi1 Bc/SM5, Cy sp males and recognized for the Bc non-Tb phenotype (Iswi mutant stocks are a gift of D. Corona; see ref [76]).
For in vivo RNAi-experiments, flies carrying a Top2 RNAi construct (line 4570; VDRC collection) were crossed to males carrying the Actin-Gal4 driver. The Oregon R laboratory strain was used as wild type control. All the stocks were maintained at 25°C on a standard medium. For markers, balancers and special chromosomes details see FlyBase (http://www.flybase.org).
Western blotting
Extract preparation and Western blotting were performed according to ref. [105] using a rabbit antiserum directed against aa 534–950 of Top2 (gift of P. Fisher; see ref [48]) diluted 1∶2,500. The anti-alpha tubulin (SIGMA) and the anti-lamin (Dm0, Hybridoma Bank) antibodies used for loading control were diluted 1∶20,000 and 1∶2,500, respectively. Bands were detected with Chemi Doc XRS+ (BIORAD laboratories), and densitometric analysis was performed using the Image Lab 4.0.1 software.
Mitotic and meiotic chromosome cytology
To analyze the morphology and the integrity of metaphase chromosomes, brains from third-instar larvae were dissected in saline (NaCl 0.7%). After incubation for 1 h with colchicine (10−5 M in saline), brains were treated for 8 min with hypotonic solution (0.5% Na Citrate), squashed in 45% acetic acid under a 20×20 mm coverslip, and immediately frozen in liquid nitrogen. To analyze anaphases and assess mitotic parameters, larval brains were disssected in saline, directly squashed without colchicine and hypothonic pretreatment and immediately frozen. The mitotic index (MI) was calculated by determining the average number of mitotic figures per optic field as described previously [47]. To visualize meiotic chromosome morphology (Figure 3C), pupal testes isolated in saline were squashed in 45% acetic acid and frozen in liquid nitrogen. For polytene chromosome analysis, salivary glands were dissected in saline, squashed in 45% acetic acid and then frozen in liquid nitrogen. After removal of the coverslip, slides were air dried and mounted in Vectashield H-200 (Vector Laboratories) containing the DNA dye DAPI.
Immunostaining
To analyze nuclear organization of primary spermatocytes larval and pupal testes were fixed according to ref [52]. To analyze mitosis, brains from third instar larvae were dissected and fixed according to ref [106]. After several rinses in phosphate-buffered saline (PBS) slides were incubated overnight at 4°C with a monoclonal anti-alpha tubulin antibody (Sigma Aldrich) diluted 1∶1,000 in PBS and either a rabbit anti-DSpd-2 (1∶5,000; ref [59]) or a rabbit anti-phosphorylated histone H3 (1∶1000 Millipore), also diluted in PBS. Primary antibodies were detected by 1 hour incubation at room temperature with FITC-conjugated anti-mouse IgG (1∶20; Jackson Laboratories) and CY3-conjugated anti-rabbit IgG (1∶300; Invitrogen), both diluted in PBS. For anti-Top2 immunostaining, larval brains fixed according to ref [107] were incubated overnight with a rabbit anti-Top2 antibody (gift of Donna J. Ardnt-Jovin; ref [49]) diluted 1∶100 in PBS. After rinsing in PBS, slides were incubated for 1 hour at room temperature with a CY3-conjugated anti-rabbit IgG (1∶300).
For polytene chromosome immunostaining, salivary glands were dissected in saline, fixed as described in ref. [108] and incubated overnight with either of the following polyclonal antibodies diluted in PBS: rabbit anti-ISWI (1 : 100; gift of D. Corona) and rabbit anti-Mof, (1 : 100; gift of J. Lucchesi). CY3-conjugated anti-rabbit IgG (1 : 300; Invitrogen) used as secondary antibodies. Immunostained preparations were mounted in Vectashield H-200 (Vector Laboratories) with DAPI.
Microscopy
All cytological preparations were examined with a Zeiss Axioplan fluorescence microscope, equipped with a cooled charged–coupled device (CCD camera; Photometrics CoolSnap HQ). Grayscale images were collected separately, pseudocolored and merged.
Supporting Information
Zdroje
1. WangJC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3 : 430–440.
2. NitissJL (2009) DNA topoisomerase II and its growing repertoire of biological functions. Nat. Rev Cancer 9 : 327–337.
3. DrakeFH, HofmannGA, BartusHF, MatternMR, CrookeST, et al. (1989) Biochemical and pharmacological properties of p170 and p180 forms of topoisomerase II. Biochemistry 28 : 8154–8160.
4. ChristensenMO, LarsenMK, BarthelmesHU, HockR, AndersenCL, et al. (2002) Dynamics of human topoisomerase II alpha and II beta in living cells. J Cell Biol 157 : 31–44.
5. WyckoffE, HsiehTS (1988) Functional expression of a Drosophila gene in yeast: genetic complementation of DNA topoisomerase II. Proc Natl Acad Sci USA 85 : 6272–6276.
6. AdachiN, MiyaikeM, IkedaH, KikuchiA (1992) Characterization of cDNA encoding the mouse DNA topoisomerase II that can complement the budding yeast top2 mutation. Nucleic Acids Res 20 : 5297–5303.
7. JensenS, RedwoodCS, JenkinsJR, AndersenAH, HicksonID (1996) Human DNA topoisomerases II alpha and II beta can functionally substitute for yeast TOP2 in chromosome segregation and recombination. Mol Gen Genet 252 : 79–86.
8. EarnshawWC, HalliganB, CookeCA, HeckMM, LiuLF (1985) Topoisomerase II is a structural component of mitotic chromosome scaffolds. J Cell Biol 100 : 1706–1715.
9. GasserSM, LarocheT, FalquetJ, Boy de la TourE, LaemmliUK (1986) Metaphase chromosome structure: involvement of topoisomerase II. J Mol Biol 188 : 613–629.
10. OhtaS, Bukowski-WillsJC, Sanchez-PulidoL, Alves FdeL, WoodL, et al. (2010) The protein composition of mitotic chromosomes determined using multiclassifier combinatorial proteomics. Cell 142 : 810–821.
11. SwedlowJR, AgardDA, SedatJW (1993) Multiple populations of topoisomerase II detected in vivo by time-lapse, threedimensional wide-field microscopy. Cell 73 : 97–108.
12. TavorminaPA, ComeM-G, HudsonJR, MoY-Y, BeckWT, et al. (2002) Rapid exchange of mammalian topoisomerase II alpha at kinetochores and chromosome arms in mitosis. J Cell Biol 158 : 23–29.
13. MaeshimaK, LaemmliUK (2003) A two-step scaffolding model for mitotic chromosome assembly. Dev Cell 4 : 467–480.
14. SamejimaK, SamejimaI, VagnarelliP, OgawaH, VargiuG, et al. (2012) Mitotic chromosomes are compacted laterally by KIF4 and condensin and axially by topoisomerase II alpha. J Cell Biol 199 : 755–770.
15. RattnerJB, HendzelMJ, FurbeeCS, MullerMT, Bazett-JonesDP (1996) Topoisomerase II alpha is associated with the mamammalian centromere in a cell cycle - and species-specific manner and is required for proper centromere/kinetochore structure. J Cell Biol 134 : 1097–1107.
16. BachantJ, AlcasabasA, BlatY, KlecknerN, ElledgeSJ (2002) The SUMO-1 isopeptidase Smt4 is linked to centromeric cohesion through SUMO-1 modification of DNA topoisomerase II. Mol Cell 9 : 1169–1182.
17. SpenceJM, PhuaHH, MillsW, CarpenterAJ, PorterAC, et al. (2007) Depletion of topoisomerase II alpha leads to shortening of the metaphase interkinetochore distance and abnormal persistence of PICH-coated anaphase threads. J Cell Sci 120 : 3952–3964.
18. DiNardoS, VoelkelK, SternglanzR (1984) DNA topoisomerase II mutant of Saccharomyces cerevisiae: topoisomerase II is required for segregation of daughter molecules at the termination of DNA replication. Proc Natl Acad Sci USA 81 : 2616–2620.
19. UemuraT, YanagidaM (1984) Isolation of type I and II DNA topoisomerase mutants from fission yeast: single and double mutants show different phenotypes in cell growth and chromatin organization. EMBO J 3 : 1737–1744.
20. HolmC, GotoT, WangJC, BotsteinD (1985) DNA topoisomerase II is required at the timing of mitosis in yeast. Cell 41 : 553–563.
21. UemuraT, OhkuraH, AdachiY, MorinoK, ShiozakiK, et al. (1987) DNA topoisomerase II is required for condensation and separation of mitotic chromosomes in S. pombe. Cell 50 : 917–925.
22. BaxterJ, DiffleyJF (2008) Topoisomerase II inactivation prevents the completion of DNA replication in budding yeast. Mol Cell 30 : 790–802.
23. BaxterJ, AragonL (2012) A model for chromosome condensation based on the interplay between condensin and topoisomerase II. Trends Genet 28 : 110–117.
24. JohnsonM, PhuaHH, BennettSC, SpenceJM, FarrCJ (2009) Studying vertebrate topoisomerase 2 function using a conditional knockdown system in DT40 cells. Nucleic Acids Res 37: e98.
25. GonzalezRE, LimCU, ColeK, BianchiniCH, SchoolsGP, et al. (2011) Effects of conditional depletion of topoisomerase II on cell cycle progression in mammalian cells. Cell Cycle 10 : 3505–3514.
26. Gimenez-AbianJF, ClarkeDJ, MullingerAM, DownesCS, JohnsonRT (1995) A postprophase topoisomerase II-dependent chromatid core separation step in the formation of metaphase chromosomes. J Cell Biol 131 : 7–17.
27. AndreassenPR, LacroixFB, MargolisRL (1997) Chromosomes with two intact axial cores are induced by G2 checkpoint override: evidence that DNA decatenation is not required to template the chromosome structure. J Cell Biol 136 : 29–43.
28. AdachiY, LukeM, LaemmliUK (1991) Chromosome assembly in vitro: topoisomerase II is required for condensation. Cell 64 : 137–148.
29. HiranoT, MitchisonTJ (1993) Topoisomerase II does not play a scaffolding role in the organization of mitotic chromosomes assembled in Xenopus egg extracts. J Cell Biol 120 : 601–612.
30. Gimenez-AbianJF, ClarkeDJ, DevlinJ, Gimenez-AbianMI, De la TorreC, et al. (2000) Premitotic chromosome individualization in mammalian cells depends on topoisomerase II activity. Chromosoma 109 : 235–344.
31. CarpenterAJ, PorterAC (2004) Construction, characterization, and complementation of a conditional-lethal DNA topoisomerase II alpha mutant human cell line. Mol Biol Cell 15 : 5700–5711.
32. SakaguchiA, KikuchiA (2004) Functional compatibility between isoform alpha and beta of type II DNA topoisomerase. J Cell Sci 117 : 1047–1054.
33. LinkaRM, PorterAC, VolkovA, MielkeC, BoegeF, et al. (2007) C-terminal regions of topoisomearase II alpha and II beta determine isoform-specific functioning of the enzymes in vivo. Nucleic Acids Res 35 : 3810–3822.
34. DownesCS, ClarkeDJ, MullingerAM, Gimenez-AbianJF, CreightonAM, et al. (1994) A topoisomerase II dependent G2 cycle checkpoint in mammalian cells. Nature 372 : 467–470.
35. DemingPB, CistulliCA, ZhaoH, GravesPR, Piwnica-WormsH, et al. (2001) The human decatenation checkpoint. Proc Natl Acad Sci USA 98 : 12044–12049.
36. SkoufiasDA, LacroixFB, AndreassenPR, WilsonL, MargolisRL (2004) Inhibition of DNA decatenation, bu not DNA damage, arrests cells at metaphase. Mol Cell 15 : 977–990.
37. FurnissKL, TsaiH-J, BylJA, LaneAB, VasAC, et al. (2013) Direct Monitoring of the strand passage reaction of DNA Topoisomerase II triggers checkpoint activation. PLoS Genet 9: e1003832.
38. BowerJJ, KaracaGF, ZhouY, SimpsonDA, Cordeiro-StoneM, et al. (2010) Topoisomerase II alpha maintains genomic stability through decatenation G(2) checkpoint signaling. Oncogene 29 : 4787–4799.
39. BuchenauP, SaumweberH, Arndt-JovinDJ (1993) Consequences of topoisomerase II inhibition in early embryogenesis of Drosophila revealed by in vivo confocal laser scanning microscopy. J Cell Sci 104 : 1175–1185.
40. ChangCJ, GouldingS, EarnshawWC, CarmenaM (2003) RNAi analysis reveals an unexpected role for topoisomerase II in chromosome arm congression to a metaphase plate. J Cell Sci 116 : 4715–4726.
41. SommaMP, CepraniF, BucciarelliE, NaimV, De ArcangelisV, et al. (2008) Identification of Drosophila mitotic genes by combining co-expression analysis and RNA interference. PloS Genetics 4: e1000126.
42. CoelhoPA, Queiroz-MachadoJ, CarmoAM, Moutinho-PereiraS, MaiatoH, et al. (2008) Dual role of topoisomerase II in centromere resolution and aurora B activity. PLoS Biol 6: e207.
43. WilliamsB, BatemanJR, NovikovND, WuCT (2007) Disruption of topoisomerase II perturbs pairing in Drosophila cell culture. Genetics 177 : 31–46.
44. RamosE, TorreEA, BusheyAM, GurudattaBV, CorcesVG (2011) DNA topoisomerase II modulates insulator function in Drosophila. PLOS ONE 6: e16562.
45. HohlAM, ThompsonM, SoshnevAA, WuJ, MorrisJ, et al. (2012) Restoration of topoisomerase 2 function by complementation of defective monomers in Drosophila. Genetics 192 : 843–856.
46. BucciarelliE, GiansantiMG, BonaccorsiS, GattiM (2003) Spindle assembly and cytokinesis in the absence of chromosomes during Drosophila male meiosis. J Cell Biol 160 : 993–999.
47. GattiM, BakerBS (1989) Genes controlling essential cell-cycle functions in Drosophila melanogaster. Genes Dev 3 : 438–453.
48. MellerVH, FisherPA, BerriosM (1995) Intranuclear distribution of DNA topoisomerase II and chromatin. Chromosome Res 4 : 255–260.
49. GemkowMJ, DichterJ, Arndt-JovinDJ (2001) Developmental Regulation of DNA-Topoisomerases during Drosophila Embryogenesis. Exp Cell Res 262 : 114–121.
50. SavvidouE, CobbeN, SteffensenS, CotterillS, HeckMM (2005) Drosophila CAP-D2 is required for condensin complex stability and resolution of sister chromatids. J Cell Sci 118 : 2529–2543.
51. Cooper KW (1950) Normal spermatogenesis in Drosophila. In: Demerec M, ed. Biology of Drosophila. New York: Wiley. pp 1–61.
52. CenciG, BonaccorsiS, PisanoC, VerniF, GattiM (1994) Chromatin and microtubule organization during premeiotic, meiotic and early postmeiotic stages of Drosophila melanogaster spermatogenesis. J Cell Sci. 107 : 3521–3534.
53. VazquezJ, BelmontAS, SedatJW (2002) The dynamics of homologous chromosome pairing during male Drosophila meiosis. Curr Biol 12 : 1473–1483.
54. McKeeBD, YanR, TsaiJH (2012) Meiosis in male Drosophila. Spermatogenesis 2 : 167–184.
55. TomkielJE, WakimotoBT, BriscoeAJr (2001) The teflon gene is required for maintenance of autosomal homolog pairing at meiosis I in male Drosophila melanogaster. Genetics 157 : 273–281.
56. ThomasSE, Soltani-BejnoodM, RothP, DornR, LogsdonJMJr, McKeeBD (2005) Identification of two proteins required for conjunction and regular segregation of achiasmate homologs in Drosophila male meiosis. Cell 123 : 555–568.
57. AryaGH, LodicoMJ, AhmadOI, AminR, TomkielJE (2006) Molecular characterization of teflon, a gene required for meiotic autosome segregation in male Drosophila melanogaster. Genetics 174 : 125–134.
58. HartlTA, SweeneySJ, KneplerPJ, BoscoG (2008) Condensin II resolves chromosomal associations to enable anaphase I segregation in Drosophila male meiosis. PLoS Genet 4: e1000228.
59. GiansantiMG, BucciarelliE, BonaccorsiS, GattiM (2008) Drosophila SPD-2 is an essential centriole component required for PCM recruitment and astral-microtubule nucleation. Curr Biol 18 : 303–309.
60. GattiM, PimpinelliS (1992) Functional elements in Drosophila melanogaster heterochromatin. Annu Rev Genet 26 : 2 39–275.
61. WeiY, YuL, BowenJ, GorovskyMA, AllisCD (1999) Phosphorylation of histone H3 is required for proper chromosome condensation and segregation. Cell 97 : 99–109.
62. BucciarelliE, PellacaniC, NaimV, PalenaA, GattiM, et al. (2009) Drosophila Dgt6 interacts with Ndc80, Msps/XMAP215, and gamma-tubulin to promote kinetochore-driven MT formation. Curr Biol 19 : 1839–1845.
63. Mottier-PavieV, CenciG, VernìF, GattiM, BonaccorsiS (2011) Phenotypic analysis of misato function reveals roles of noncentrosomal microtubules in Drosophila spindle formation. J Cell Sci 124 : 706–717.
64. SperkaT, WangJ, RudolphKL (2012) DNA damage checkpoints in stem cells, ageing and cancer. Nat Rev Mol Cell Biol 13 : 579–590.
65. ShilohY, ZivY (2013) The ATM protein kinase: regulating the cellular response to genotoxic stress, and more. Nat Rev Mol Cell Biol 14 : 197–210.
66. CiapponiL, CenciG, GattiM (2006) The Drosophila Nbs protein functions in multiple pathways for the maintenance of genome stability. Genetics 173 : 1 447–1454.
67. BauerCR, HartlTA, BoscoG (2012) Condensin II promotes the formation of chromosome territories by inducing axial compaction of polyploid interphase chromosomes. PLoS Genet 8 doi: 10.1371/journal.pgen.1002873
68. DeuringR, FantiL, ArmstrongJA, SarteM, PapoulasO, et al. (2000) The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol Cell 5 : 355–365.
69. BadenhorstP, VoasM, RebayI, WuC (2002) Biological functions of the ISWI chromatin remodeling complex NURF. Genes Dev 16 : 3186–3198.
70. SpiererA, SeumC, DelattreM, SpiererP (2005) Loss of the modifiers of variegation Su(var)3–7 or HP1 impacts male X polytene chromosome morphology and dosage compensation. J Cell Sci 118 : 5047–5057.
71. SpiererA, BegeotF, SpiererP, DelattreM (2008) SU(VAR)3-7 links heterochromatin and dosage compensation in Drosophila. PLoS Genet 4 doi: 10.1371/journal.pgen.1000066
72. VermaakD, MalikHS (2009) Multiple roles for heterochromatin protein 1 genes in Drosophila. Annu Rev Genet 43 : 467–492.
73. LängstG, BeckerPB (2001) ISWI induces nucleosome sliding on nicked DNA. Mol Cell 8 : 1085–1092.
74. GelbartME, KurodaMI (2009) Drosophila dosage compensation: a complex voyage to the X chromosome. Development 136 : 1399–1410.
75. LucchesiJC (2009) The structure-function link of compensated chromatin in Drosophila. Curr Opin Genet Dev 19 : 550–556.
76. CoronaDF, ClapierCR, BeckerPB, TamkunJW (2002) Modulation of ISWI function by site specific acetylation. EMBO Rep 3 : 242–247.
77. Varga-WeiszPD, WilmM, BonteE, DumasK, MannM, et al. (1997) Chromatin-remodelling factor CHRAC contains the ATPases ISWI and topoisomerase II. Nature 388 : 598–602.
78. TsaiJ-H, YanR, McKeeBD (2011) Homolog pairing and sister chromatid cohesion in heterochromatin in Drosophila male meiosis I. Chromosoma 120 : 335–351.
79. RoseD, ThomasW, HolmC (1990) Segregation of recombined chromosomes in meiosis requires DNA topoisomerase II. Cell 60 : 1009–1017.
80. RoseD, HolmC (1993) Meiosis-specific arrest revealed in DNA topoisomerase II mutants. Mol Cell Biol 13 : 3445–3455.
81. HartsuikerE, BählerJ, KohliJ (1998) The role of topoisomerase II in meiotic chromosome condensation and segregation in Schizosaccharomyces pombe. Mol Biol Cell 10 : 2739–2750.
82. KallioM, LähdetieJ (1996) Fragmentation of centromeric DNA and prevention of homologous chromosome separation in male meiosis in vivo by the topoisomerase II inhibitor etoposide. Mutagenesis 11 : 435–443.
83. KallioM, LähdetieJ (1997) Effects of the DNA topoisomerase II inhibitor merbarone in male mouse meiotic divisions in vivo: cell cycle arrest and induction of aneuploidy. Environ Mol Mutagen 29 : 16–27.
84. MarchettiF, BishopJB, LoweX, GenerosoWM, HozierJ, et al. (2001) Etoposide induces heritable chromosomal aberrations and aneuploidy during male meiosis in the mouse. Proc Natl Acad Sci USA 98 : 3952–3957.
85. TatenoH, KamiguchiY (2001) Meiotic stage-dependent induction of chromosome aberrations in Chinese hamster primary oocytes exposed to topoisomerase II inhibitor etoposide. Mutat Res 476 : 139–148.
86. CobbJ, ReddyRK, ParkC, HandelMA (1997) Analysis of expression and function of topoisomerase I and II during meiosis in male mice. Mol Reprod Dev 46 : 489–498.
87. AlpheyL, JimenezJ, White-CooperH, DawsonI, NurseP, et al. (1992) twine, a cdc25 homolog that functions in the male and female germline of Drosophila. Cell 69 : 977–988.
88. RebolloE, GonzálezC (2000) Visualizing the spindle checkpoint in Drosophila spermatocytes. EMBO Rep 1 : 65–70.
89. NitissJL (2009) Targeting DNA topoisomerase II in cancer chemotherapy. Nat Rev Cancer 9 : 338–350.
90. HolmC, StearnsT, BotsteinD (1989) DNA topoisomerase II must act at mitosis to prevent nondisjunction and chromosome breakage. Mol Cell Biol 9 : 159–168.
91. ToyodaY, YanagidaM (2006) Coordinated requirements of human Topo II and cohesin for metaphase centromere alignment under Mad2-dependent spindle checkpoint surveillance. Mol Biol Cell 17 : 2287–2302.
92. KäsE, LaemmliUK (1992) In vivo topoisomerase II cleavage of the Drosophila histone and satellite III repeats: DNA sequence and structural characteristics. EMBO J 11 : 705–716.
93. BonaccorsiS, LoheAR (1991) Fine mapping of satellite DNA sequences along the Y chromosome of Drosophila melanogaster: relationships between satellite sequences and fertility factors. Genetics 129 : 177–189.
94. LoheAR, HillikerAJ, RobertsPA (1993) Mapping Simple Repeated DNA Sequences in Heterochromatin of Drosophila melanogaster. Genetics 134 : 1149–1174.
95. PimpinelliS, BerlocoM, FantiL, DimitriP, BonaccorsiS, et al. (1995) Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc Natl Acad Sci USA 92 : 3804–3808.
96. LuoK, YuanJ, ChenJ, LouZ (2009) Topoisomearase II alpha controls the decatenation checkpoint. Nat Cell Biol 11 : 204–210.
97. CugusiS, RamosE, LingH, YokoyamaR, LukKM, et al. (2013) Topoisomerase II plays a role in dosage compensation in Drosophila. Transcription 4 : 1–13.
98. WangCI, AlekseyenkoAA, LeRoyG, EliaAE, GorchakovAA, et al. (2013) Chromatin proteins captured by ChIP-mass spectrometry are linked to dosage compensation in Drosophila. Nat Struct Mol Biol 20 : 202–209.
99. KurzEU, WilsonSE, LeaderKB, SampeyBP, AllanWP, et al. (2002) The histone deacetylase inhibitor sodium butyrate induces DNA topoisomerase II alpha expression and confers hypersensitivity to etoposide in human leukemic cell lines. Mol Cancer Ther 1 : 121–131.
100. KimMS, BlakeM, BaekJH, KohlhagenG, PommierY, et al. (2003) Inhibition of histone deacetylase increases cytotoxicity to anticancer drugs targeting DNA. Cancer Res 63 : 7291–7300.
101. KoundakjianEJ, CowanDM, HardyRW, BeckerAH (2004) The Zuker collection: a resource for the analysis of autosomal gene function in Drosophila melanogaster. Genetics 167 : 203–206.
102. LattaoR, BonaccorsiS, GuanX, WassermanSA, GattiM (2011) Tubby-tagged balancers for the Drosophila X and second chromosomes. Fly (Austin) 5 : 369–370.
103. LaurençonA, PurdyA, SekelskyJ, HawleyRS, SuTT (2003) Phenotypic analysis of separation-of-function alleles of MEI-41, Drosophila ATM/ATR. Genetics 164 : 589–601.
104. SilvaE, TiongS, PedersenM, HomolaE, RoyouA, et al. (2004) ATM is required for telomere maintenance and chromosome stability during Drosophila development. Curr Biol 14 : 1341–1347.
105. SommaMP, FasuloB, CenciG, CundariE, GattiM (2002) Molecular dissection of cytokinesis by RNA interference in Drosophila cultured cells. Mol Biol Cell 13 : 2448–60.
106. BonaccorsiS, GiansantiMG, GattiM (2000) Spindle assembly in Drosophila neuroblasts and ganglion mother cells. Nat Cell Biol 2 : 54–56.
107. CiapponiL, CenciG, DucauJ, FloresC, Johnson-SchlitzD, et al. (2004) The Drosophila Mre11/Rad50 complex is required to prevent both telomeric fusion and chromosome breakage. Curr Biol 14 : 1360–6.
108. RaffaGD, CenciG, SiriacoG, GoldbergML, GattiM (2005) The putative Drosophila transcription factor Woc is required to prevent telomeric fusions. Mol Cell 20 : 821–31.
Štítky
Genetika Reprodukční medicína
Článek Oligoasthenoteratozoospermia and Infertility in Mice Deficient for miR-34b/c and miR-449 LociČlánek The Kinesin AtPSS1 Promotes Synapsis and is Required for Proper Crossover Distribution in MeiosisČlánek Payoffs, Not Tradeoffs, in the Adaptation of a Virus to Ostensibly Conflicting Selective PressuresČlánek Examination of Prokaryotic Multipartite Genome Evolution through Experimental Genome ReductionČlánek BMP-FGF Signaling Axis Mediates Wnt-Induced Epidermal Stratification in Developing Mammalian SkinČlánek Role of STN1 and DNA Polymerase α in Telomere Stability and Genome-Wide Replication in ArabidopsisČlánek RNA-Processing Protein TDP-43 Regulates FOXO-Dependent Protein Quality Control in Stress ResponseČlánek Integrating Functional Data to Prioritize Causal Variants in Statistical Fine-Mapping StudiesČlánek Salt-Induced Stabilization of EIN3/EIL1 Confers Salinity Tolerance by Deterring ROS Accumulation inČlánek Ethylene-Induced Inhibition of Root Growth Requires Abscisic Acid Function in Rice ( L.) SeedlingsČlánek Metabolic Respiration Induces AMPK- and Ire1p-Dependent Activation of the p38-Type HOG MAPK PathwayČlánek Signature Gene Expression Reveals Novel Clues to the Molecular Mechanisms of Dimorphic Transition inČlánek A Mouse Model Uncovers LKB1 as an UVB-Induced DNA Damage Sensor Mediating CDKN1A (p21) DegradationČlánek Dominant Sequences of Human Major Histocompatibility Complex Conserved Extended Haplotypes from to
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 10- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- An Deletion Is Highly Associated with a Juvenile-Onset Inherited Polyneuropathy in Leonberger and Saint Bernard Dogs
- Licensing of Yeast Centrosome Duplication Requires Phosphoregulation of Sfi1
- Oligoasthenoteratozoospermia and Infertility in Mice Deficient for miR-34b/c and miR-449 Loci
- Basement Membrane and Cell Integrity of Self-Tissues in Maintaining Immunological Tolerance
- The Kinesin AtPSS1 Promotes Synapsis and is Required for Proper Crossover Distribution in Meiosis
- Germline Mutations in Are Associated with Familial Gastric Cancer
- POT1a and Components of CST Engage Telomerase and Regulate Its Activity in
- Controlling Meiotic Recombinational Repair – Specifying the Roles of ZMMs, Sgs1 and Mus81/Mms4 in Crossover Formation
- Payoffs, Not Tradeoffs, in the Adaptation of a Virus to Ostensibly Conflicting Selective Pressures
- FHIT Suppresses Epithelial-Mesenchymal Transition (EMT) and Metastasis in Lung Cancer through Modulation of MicroRNAs
- Genome-Wide Mapping of Yeast RNA Polymerase II Termination
- Examination of Prokaryotic Multipartite Genome Evolution through Experimental Genome Reduction
- White Cells Facilitate Opposite- and Same-Sex Mating of Opaque Cells in
- BMP-FGF Signaling Axis Mediates Wnt-Induced Epidermal Stratification in Developing Mammalian Skin
- Genome-Wide Association Study of CSF Levels of 59 Alzheimer's Disease Candidate Proteins: Significant Associations with Proteins Involved in Amyloid Processing and Inflammation
- COE Loss-of-Function Analysis Reveals a Genetic Program Underlying Maintenance and Regeneration of the Nervous System in Planarians
- Fat-Dachsous Signaling Coordinates Cartilage Differentiation and Polarity during Craniofacial Development
- Identification of Genes Important for Cutaneous Function Revealed by a Large Scale Reverse Genetic Screen in the Mouse
- Sensors at Centrosomes Reveal Determinants of Local Separase Activity
- Genes Integrate and Hedgehog Pathways in the Second Heart Field for Cardiac Septation
- Systematic Dissection of Coding Exons at Single Nucleotide Resolution Supports an Additional Role in Cell-Specific Transcriptional Regulation
- Recovery from an Acute Infection in Requires the GATA Transcription Factor ELT-2
- HIPPO Pathway Members Restrict SOX2 to the Inner Cell Mass Where It Promotes ICM Fates in the Mouse Blastocyst
- Role of and in Development of Abdominal Epithelia Breaks Posterior Prevalence Rule
- The Formation of Endoderm-Derived Taste Sensory Organs Requires a -Dependent Expansion of Embryonic Taste Bud Progenitor Cells
- Role of STN1 and DNA Polymerase α in Telomere Stability and Genome-Wide Replication in Arabidopsis
- Keratin 76 Is Required for Tight Junction Function and Maintenance of the Skin Barrier
- Encodes the Catalytic Subunit of N Alpha-Acetyltransferase that Regulates Development, Metabolism and Adult Lifespan
- Disruption of SUMO-Specific Protease 2 Induces Mitochondria Mediated Neurodegeneration
- Caudal Regulates the Spatiotemporal Dynamics of Pair-Rule Waves in
- It's All in Your Mind: Determining Germ Cell Fate by Neuronal IRE-1 in
- A Conserved Role for Homologs in Protecting Dopaminergic Neurons from Oxidative Stress
- The Master Activator of IncA/C Conjugative Plasmids Stimulates Genomic Islands and Multidrug Resistance Dissemination
- An AGEF-1/Arf GTPase/AP-1 Ensemble Antagonizes LET-23 EGFR Basolateral Localization and Signaling during Vulva Induction
- The Proteomic Landscape of the Suprachiasmatic Nucleus Clock Reveals Large-Scale Coordination of Key Biological Processes
- RNA-Processing Protein TDP-43 Regulates FOXO-Dependent Protein Quality Control in Stress Response
- A Complex Genetic Switch Involving Overlapping Divergent Promoters and DNA Looping Regulates Expression of Conjugation Genes of a Gram-positive Plasmid
- ZTF-8 Interacts with the 9-1-1 Complex and Is Required for DNA Damage Response and Double-Strand Break Repair in the Germline
- Integrating Functional Data to Prioritize Causal Variants in Statistical Fine-Mapping Studies
- Tpz1-Ccq1 and Tpz1-Poz1 Interactions within Fission Yeast Shelterin Modulate Ccq1 Thr93 Phosphorylation and Telomerase Recruitment
- Salt-Induced Stabilization of EIN3/EIL1 Confers Salinity Tolerance by Deterring ROS Accumulation in
- Telomeric (s) in spp. Encode Mediator Subunits That Regulate Distinct Virulence Traits
- Ethylene-Induced Inhibition of Root Growth Requires Abscisic Acid Function in Rice ( L.) Seedlings
- Ancient Expansion of the Hox Cluster in Lepidoptera Generated Four Homeobox Genes Implicated in Extra-Embryonic Tissue Formation
- Mechanism of Suppression of Chromosomal Instability by DNA Polymerase POLQ
- A Mutation in the Mouse Gene Leads to Impaired Hedgehog Signaling
- Keeping mtDNA in Shape between Generations
- Targeted Exon Capture and Sequencing in Sporadic Amyotrophic Lateral Sclerosis
- TIF-IA-Dependent Regulation of Ribosome Synthesis in Muscle Is Required to Maintain Systemic Insulin Signaling and Larval Growth
- At Short Telomeres Tel1 Directs Early Replication and Phosphorylates Rif1
- Evidence of a Bacterial Receptor for Lysozyme: Binding of Lysozyme to the Anti-σ Factor RsiV Controls Activation of the ECF σ Factor σ
- Hsp40s Specify Functions of Hsp104 and Hsp90 Protein Chaperone Machines
- Feeding State, Insulin and NPR-1 Modulate Chemoreceptor Gene Expression via Integration of Sensory and Circuit Inputs
- Functional Interaction between Ribosomal Protein L6 and RbgA during Ribosome Assembly
- Multiple Regulatory Systems Coordinate DNA Replication with Cell Growth in
- Fast Evolution from Precast Bricks: Genomics of Young Freshwater Populations of Threespine Stickleback
- Mmp1 Processing of the PDF Neuropeptide Regulates Circadian Structural Plasticity of Pacemaker Neurons
- The Nuclear Immune Receptor Is Required for -Dependent Constitutive Defense Activation in
- Genetic Modifiers of Neurofibromatosis Type 1-Associated Café-au-Lait Macule Count Identified Using Multi-platform Analysis
- Juvenile Hormone-Receptor Complex Acts on and to Promote Polyploidy and Vitellogenesis in the Migratory Locust
- Uncovering Enhancer Functions Using the α-Globin Locus
- The Analysis of Mutant Alleles of Different Strength Reveals Multiple Functions of Topoisomerase 2 in Regulation of Chromosome Structure
- Metabolic Respiration Induces AMPK- and Ire1p-Dependent Activation of the p38-Type HOG MAPK Pathway
- The Specification and Global Reprogramming of Histone Epigenetic Marks during Gamete Formation and Early Embryo Development in
- The DAF-16 FOXO Transcription Factor Regulates to Modulate Stress Resistance in , Linking Insulin/IGF-1 Signaling to Protein N-Terminal Acetylation
- Genetic Influences on Translation in Yeast
- Analysis of Mutants Defective in the Cdk8 Module of Mediator Reveal Links between Metabolism and Biofilm Formation
- Ribosomal Readthrough at a Short UGA Stop Codon Context Triggers Dual Localization of Metabolic Enzymes in Fungi and Animals
- Gene Duplication Restores the Viability of Δ and Δ Mutants
- Selection on a Variant Associated with Improved Viral Clearance Drives Local, Adaptive Pseudogenization of Interferon Lambda 4 ()
- Break-Induced Replication Requires DNA Damage-Induced Phosphorylation of Pif1 and Leads to Telomere Lengthening
- Dynamic Partnership between TFIIH, PGC-1α and SIRT1 Is Impaired in Trichothiodystrophy
- Signature Gene Expression Reveals Novel Clues to the Molecular Mechanisms of Dimorphic Transition in
- Mutations in Moderate or Severe Intellectual Disability
- Multifaceted Genome Control by Set1 Dependent and Independent of H3K4 Methylation and the Set1C/COMPASS Complex
- A Role for Taiman in Insect Metamorphosis
- The Small RNA Rli27 Regulates a Cell Wall Protein inside Eukaryotic Cells by Targeting a Long 5′-UTR Variant
- MMS Exposure Promotes Increased MtDNA Mutagenesis in the Presence of Replication-Defective Disease-Associated DNA Polymerase γ Variants
- Coexistence and Within-Host Evolution of Diversified Lineages of Hypermutable in Long-term Cystic Fibrosis Infections
- Comprehensive Mapping of the Flagellar Regulatory Network
- Topoisomerase II Is Required for the Proper Separation of Heterochromatic Regions during Female Meiosis
- A Splice Mutation in the Gene Causes High Glycogen Content and Low Meat Quality in Pig Skeletal Muscle
- KDM5 Interacts with Foxo to Modulate Cellular Levels of Oxidative Stress
- H2B Mono-ubiquitylation Facilitates Fork Stalling and Recovery during Replication Stress by Coordinating Rad53 Activation and Chromatin Assembly
- Copy Number Variation in the Horse Genome
- Unifying Genetic Canalization, Genetic Constraint, and Genotype-by-Environment Interaction: QTL by Genomic Background by Environment Interaction of Flowering Time in
- Spinster Homolog 2 () Deficiency Causes Early Onset Progressive Hearing Loss
- Genome-Wide Discovery of Drug-Dependent Human Liver Regulatory Elements
- Developmentally-Regulated Excision of the SPβ Prophage Reconstitutes a Gene Required for Spore Envelope Maturation in
- Protein Phosphatase 4 Promotes Chromosome Pairing and Synapsis, and Contributes to Maintaining Crossover Competence with Increasing Age
- The bHLH-PAS Transcription Factor Dysfusion Regulates Tarsal Joint Formation in Response to Notch Activity during Leg Development
- A Mouse Model Uncovers LKB1 as an UVB-Induced DNA Damage Sensor Mediating CDKN1A (p21) Degradation
- Notch3 Interactome Analysis Identified WWP2 as a Negative Regulator of Notch3 Signaling in Ovarian Cancer
- An Integrated Cell Purification and Genomics Strategy Reveals Multiple Regulators of Pancreas Development
- Dominant Sequences of Human Major Histocompatibility Complex Conserved Extended Haplotypes from to
- The Vesicle Protein SAM-4 Regulates the Processivity of Synaptic Vesicle Transport
- A Gain-of-Function Mutation in Impeded Bone Development through Increasing Expression in DA2B Mice
- Nephronophthisis-Associated Regulates Cell Cycle Progression, Apoptosis and Epithelial-to-Mesenchymal Transition
- Beclin 1 Is Required for Neuron Viability and Regulates Endosome Pathways via the UVRAG-VPS34 Complex
- The Not5 Subunit of the Ccr4-Not Complex Connects Transcription and Translation
- Abnormal Dosage of Ultraconserved Elements Is Highly Disfavored in Healthy Cells but Not Cancer Cells
- Genome-Wide Distribution of RNA-DNA Hybrids Identifies RNase H Targets in tRNA Genes, Retrotransposons and Mitochondria
- The Chromosomal Association of the Smc5/6 Complex Depends on Cohesion and Predicts the Level of Sister Chromatid Entanglement
- Cell-Autonomous Progeroid Changes in Conditional Mouse Models for Repair Endonuclease XPG Deficiency
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- The Master Activator of IncA/C Conjugative Plasmids Stimulates Genomic Islands and Multidrug Resistance Dissemination
- A Splice Mutation in the Gene Causes High Glycogen Content and Low Meat Quality in Pig Skeletal Muscle
- Keratin 76 Is Required for Tight Junction Function and Maintenance of the Skin Barrier
- A Role for Taiman in Insect Metamorphosis
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



