-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Meta-Analysis of Genome-Wide Association Studies in African Americans Provides Insights into the Genetic Architecture of Type 2 Diabetes
Despite the higher prevalence of type 2 diabetes (T2D) in African Americans than in Europeans, recent genome-wide association studies (GWAS) were examined primarily in individuals of European ancestry. In this study, we performed meta-analysis of 17 GWAS in 8,284 cases and 15,543 controls to explore the genetic architecture of T2D in African Americans. Following replication in additional 6,061 cases and 5,483 controls in African Americans, and 8,130 cases and 38,987 controls of European ancestry, we identified two novel and three previous reported T2D loci reaching genome-wide significance. We also examined 158 loci previously reported to be associated with T2D or regulating glucose homeostasis. While 56% of these loci were shared between African Americans and the other populations, the strongest associations in African Americans are often found in nearby single nucleotide polymorphisms (SNPs) instead of the original SNPs reported in other populations due to differential genetic architecture across populations. Our results highlight the importance of performing genetic studies in non-European populations to fine map the causal genetic variants.
Published in the journal: . PLoS Genet 10(8): e32767. doi:10.1371/journal.pgen.1004517
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004517Summary
Despite the higher prevalence of type 2 diabetes (T2D) in African Americans than in Europeans, recent genome-wide association studies (GWAS) were examined primarily in individuals of European ancestry. In this study, we performed meta-analysis of 17 GWAS in 8,284 cases and 15,543 controls to explore the genetic architecture of T2D in African Americans. Following replication in additional 6,061 cases and 5,483 controls in African Americans, and 8,130 cases and 38,987 controls of European ancestry, we identified two novel and three previous reported T2D loci reaching genome-wide significance. We also examined 158 loci previously reported to be associated with T2D or regulating glucose homeostasis. While 56% of these loci were shared between African Americans and the other populations, the strongest associations in African Americans are often found in nearby single nucleotide polymorphisms (SNPs) instead of the original SNPs reported in other populations due to differential genetic architecture across populations. Our results highlight the importance of performing genetic studies in non-European populations to fine map the causal genetic variants.
Introduction
The prevalence of type 2 diabetes (T2D) among adults in the USA is currently 11.3%, with substantially higher prevalence in African Americans (18.7%) than in European Americans (10.2%) [1]. To date, genome-wide association studies (GWAS) have identified >70 susceptibility loci for T2D [2]–[8]. While it is known that T2D is heritable in African Americans [9], it is unclear how much heritability is explained by the known genetic associations discovered primarily from European ancestry populations and whether there are risk loci specific to African Americans. Given that individuals of African ancestry tend to harbor more genetic diversity than individuals of other ancestries [10], we hypothesized that large-scale association analyses in African Americans could shed light on the genetic architecture of T2D and the risk attributable to cosmopolitan vs. population-specific variants.
Results
Study overview
We conducted a meta-analysis of 17 African American GWAS on T2D comprising 8,284 cases and 15,543 controls (Tables S1 and S2). Missing genotypes in individual studies were imputed to one of the HapMap reference panels (Phase II release 21–24 CEU+YRI, Phase II release 22 all populations, Phase II+III release 27 CEU+YRI, Phase II+III release 27 CEU+YRI+ASW or Phase II+III release 27 all populations) using MACH, IMPUTE2 or BEAGLE (Table S3). Genomic control corrections [11] were applied to each study (λ = 1.01–1.08) and after meta-analysis (λ = 1.06) due to modest inflated association results (Table S3) [12]. Association results for ∼2.6M SNPs were subsequently examined.
From stage 1 meta-analysis, 49 SNPs moderately associated with T2D (P<1×10−5) and two candidate SNPs near the p value threshold (rs231356 at KCNQ1, P = 2.84×10−5 and rs2244020 at HLA-B, P = 1.02×10−5) totaling 51 SNPs in 21 loci were followed up for replication. rs231356 is 14 kb downstream of the reported T2D index SNP, rs231362, in Europeans [3]. Moderate associations have also been observed across the HLA region in Europeans [3]. The stage 2 replication included in silico and de novo replication in up to 11,544 African American T2D cases and controls, as well as in silico replication in 47,117 individuals of European ancestry from DIAGRAMv2 [3] (Table S4). Meta-analyses were performed to combine results from African Americans (stage 1+2a, n≤35,371, Table S4) and both African Americans and Europeans (stage 1+2a+2b, n≤82,488, Table S4).
T2D loci reaching genome-wide significance
Five independent loci reached genome-wide significance (P<5×10−8). Stage 1 meta-analysis identified the established TCF7L2 locus. Stage 1+2a meta-analysis identified the established KCNQ1 and HMGA2 loci. Stage 1+2a+2b meta-analysis identified a second signal at KCNQ1 and a novel HLA-B locus. Secondary analysis including body mass index (BMI) adjustment in stage 1+2a meta-analysis identified the second novel locus at INS-IGF2 (Table 1 and Figure 1). None of the most strongly associated SNPs at these loci demonstrated significant heterogeneity of effect sizes among studies within each stage, between African Americans in stages 1 and 2a, or between African Americans in stage 1+2a and Europeans in stage 2b after Bonferroni correction of multiple comparisons (Phet>0.001) (Figure S1).
Fig. 1. Association results of stage 1 meta-analysis in African Americans in a model adjusted for age, sex, study sites and study-specific principle components. 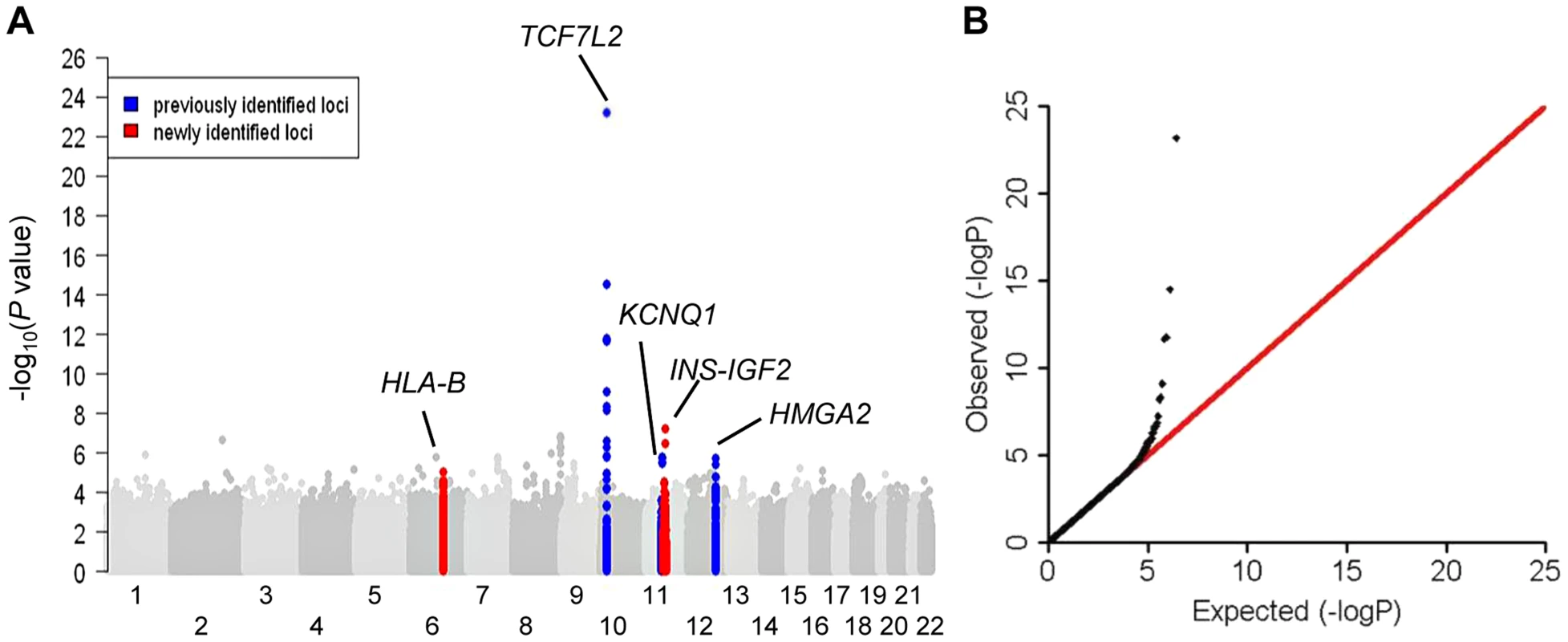
(A) Manhattan plot. Previously identified loci are denoted in red. Novel loci identified in this study are denoted in blue. (B) Quantile-quantile plot. The observed P values (y axis) were compared with the expected P values under the null distribution (x axis). Tab. 1. Novel and previously identified loci associated with T2D at P<5×10−8. 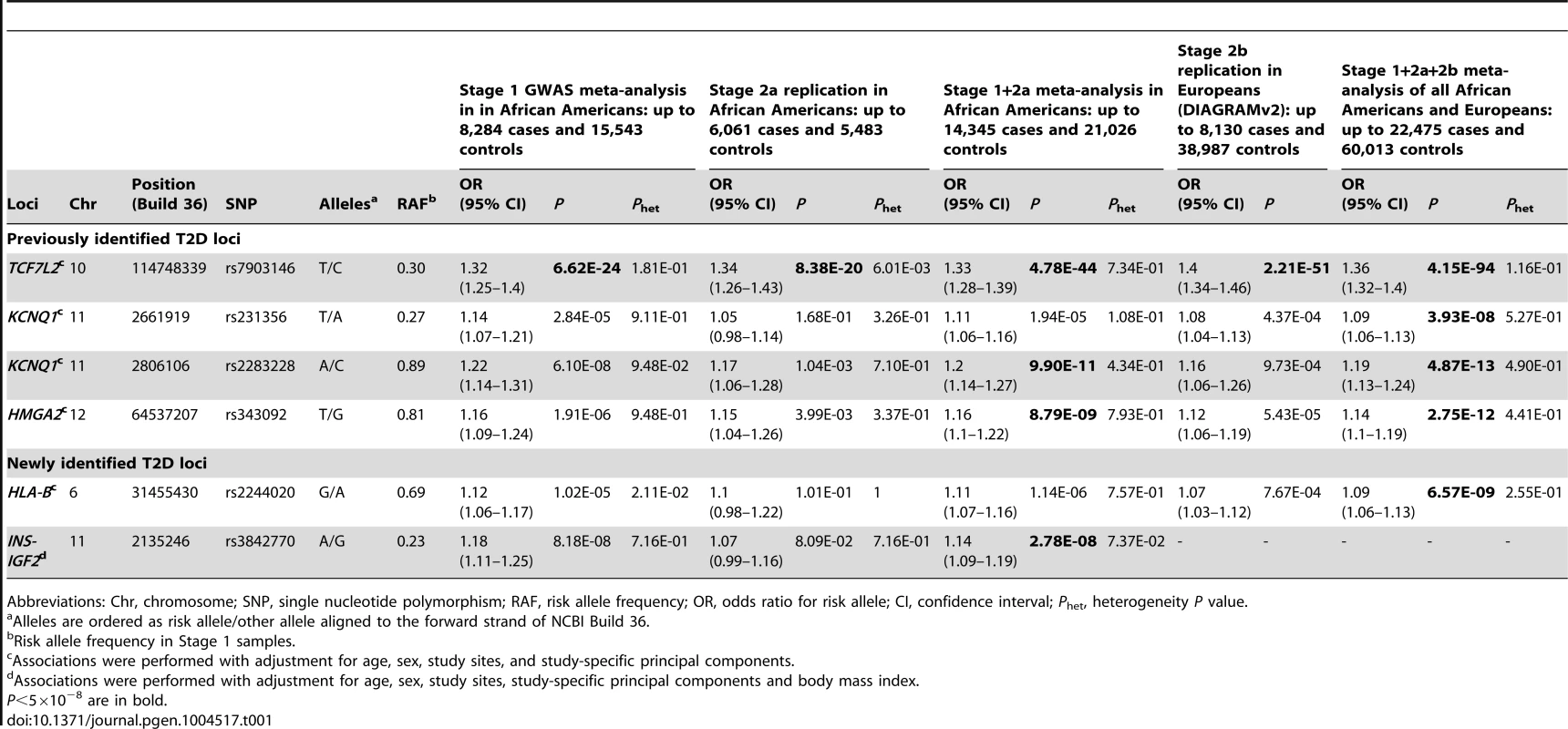
Abbreviations: Chr, chromosome; SNP, single nucleotide polymorphism; RAF, risk allele frequency; OR, odds ratio for risk allele; CI, confidence interval; Phet, heterogeneity P value. At the TCF7L2 locus, the most strongly associated SNP in stage 1+2a African Americans samples was rs7903146 (OR = 1.33, P = 4.78×10−44, Table 1 and Figure 2). rs7903146 is also the index SNP (most significantly associated with T2D in prior studies) in Europeans (OR = 1.40, P = 2.21×10−51) [3], South Asians (OR = 1.25, P = 3.4×10−19) [4] and East Asians (OR = 1.48, P = 2.44×10−15) [13].
Fig. 2. Regional plots of five previously and newly identified T2D loci in African Americans. 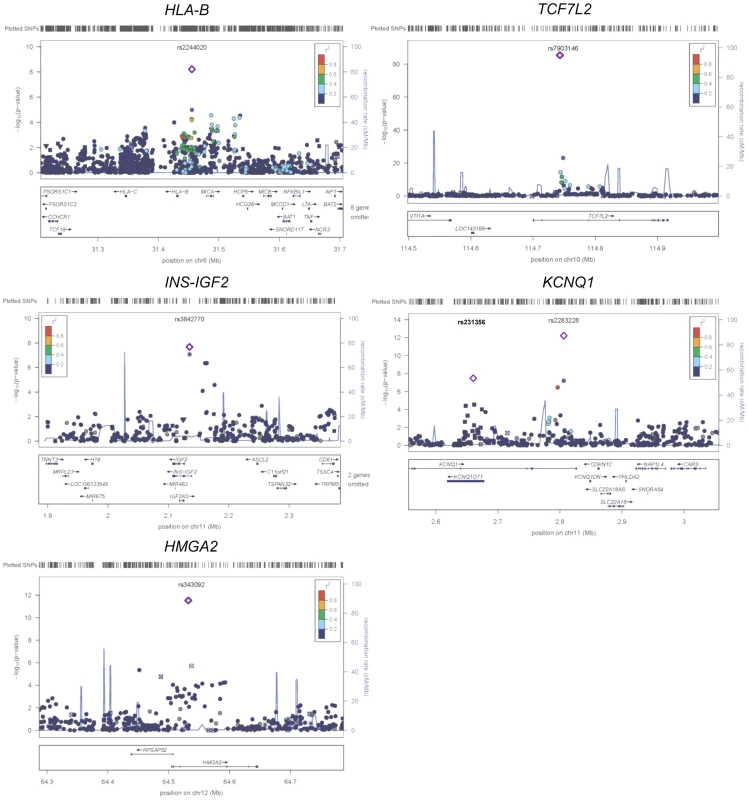
Association P values (on a −log10 scale) of genotyped and imputed SNPs from stage 1 meta-analysis are plotted as a function of genomic position (NCBI Build 36). Plots for HLA-B, TCF7L2, KCNQ1, and HMGA2 used the model without BMI adjustment whereas plots for INS-IGF2 used the model with BMI adjustment. In each panel, the most strongly associated SNP from stage 1 and stage 1+2a+2b meta-analysis is denoted by a purple circle and a purple diamond, respectively. The color of all other SNPs indicates LD with the most strongly associated SNP based on the HapMap 2 YRI data. At KCNQ1, two independent signals are shown. Two association signals were observed at KCNQ1 (Table 1 and Figure 2). The first association signal was represented by rs2283228 located at the 3′ end of KCNQ1 (stage 1+2a OR = 1.20, P = 9.90×10−11; stage 1+2a+2b OR = 1.19, P = 4.87×10−13). Using data from individuals of African ancestry in Southwest USA (ASW) from the 1000 Genomes Project (1KGP) [14], rs2283228 mapped to the same linkage disequilibrium (LD)-based interval as index SNPs from other populations (rs2283228 [15] and rs2237892 [16]–[17] in Japanese, rs2237892 in Hispanics [18], rs163182 [19] and rs2237895 [20] in Han Chinese). The second association signal was represented by rs231356 (r2 = 0 with rs2283228 in both ASW and CEU) (stage 1+2a OR = 1.11, P = 1.94×10−5; stage 1+2a+2b OR = 1.09, P = 3.93×10−8), located 144 kb upstream of the first signal. rs231356 is located at the same LD interval as the index SNPs rs231362 in Europeans [3] and rs231359 in Chinese [20].
At the HMGA2 locus, the most strongly associated SNP was rs343092 (stage 1+2a OR = 1.16, P = 8.79×10−9; stage 1+2a+2b OR = 1.14, P = 2.75×10−12; Table 1 and Figure 2). rs343092 is located 76 kb downstream and at the same LD interval as of the index SNP rs1531343 reported in Europeans [3].
Two novel T2D loci were identified. The effect sizes of rs2244020 located near HLA-B were similar in African Americans and Europeans (OR = 1.11 vs. 1.07, Phet = 0.26; stage 1+2a+2b P = 6.57×10−9) (Table 1 and Figure 2). HLA-B encodes the class I major histocompatibility complex involved in antigen presentation in immune responses.
The most strongly associated SNP near INS-IGF2 was rs3842770 in African Americans (OR = 1.14, P = 2.78×10−8, stage 1+2a BMI adjusted, Table 1 and Figure 2) but the risk A allele was absent in the CEU population. Insulin plays a key role in glucose homeostasis. Mutations at INS lead to neonatal diabetes, type 1 diabetes, and hyperinsulinemia [21]. Insulin-like growth factor 2 (IGF2) is involved in growth and development. IGF2 overexpression in transgenic mice leads to islet hyperplasia [22] and IGF2 deficiency in the Goto–Kakizaki rat leads to beta cell mass anomaly [23].
Associations at previously reported T2D and glucose homeostasis loci
We investigated index SNPs from 158 independent loci associated with T2D and/or glucose homeostasis from prior genome-wide and candidate gene studies in individuals of European, East Asian, South Asian, or African American ancestry (Table S5). Among the 104 T2D-associated index SNPs, 19 were associated with T2D in stage 1 African American samples (P<0.05). Most of the 17 T2D-associated SNPs that showed consistent direction of effects had similar effect sizes between this study and prior reports, despite that rs10440833 at CDKAL1 had substantially stronger effect size in Europeans (OR = 1.25) than in African Americans (OR = 1.06, Phet = 5.86×10−6). Additionally, 3 out of 54 trait-increasing alleles from glucose homeostasis-associated index SNPs were associated with increased T2D risk in African Americans (P<0.05).
We also performed a locus-wide analysis to test for associations of all SNPs within the LD region at r2≥0.3 with the previously reported index SNPs and results were corrected for the effective number of SNPs [24]. Since the causal variant(s) at each locus may be different or reside on different haplotypes across populations with different LD structures, this approach allows the identification of the most strongly associated SNPs in African Americans that may or may not be in LD with the index SNPs reported in other populations. A total of 55 T2D - and 29 glucose-associated loci were associated with T2D in African Americans (Plocus<0.05, corrected for LD in ASW for SNPs within a locus; Table S6). We compared the genetic architecture between the previously reported index SNPs and our fine-mapped SNPs for these 84 loci. The respective average risk allele frequencies were 0.51 and 0.46, and the distributions or pairwise differences of risk allele frequencies were not significantly different (P = 0.255, Wilcoxon rank sum test; and P = 0.295, Wilcoxon signed-rank test, respectively, Figure S2). In contrast, the average odds ratios for the risk alleles were higher for the fine-mapped SNPs as compared to the index SNPs (1.14 vs. 1.05). The distributions and pairwise differences of risk allele odds ratios were significantly different (P = 1.18×10−19 and 5.55×10−14, respectively, Figure S2). Thus, the locus-wide analysis identified variants with larger effect sizes and similar allele frequencies.
We leveraged differences in LD between African Americans and Europeans to fine-map and re-annotate several established loci. The association signal spanning ∼100 kb at INTS8 in African Americans overlapped the ∼200 kb TP53INP1 T2D locus in Europeans [3]. The most strongly associated SNP in MEDIA tended to have larger effect size in African Americans than in Europeans (rs17359493, OR = 1.13 vs. 1.06, P = 1.39×10−7 vs. 3.20×10−2, respectively, Phet = 0.06) (Table S4). However, rs17359493 at intron 10 of INTS8 was only in weak LD with the reported index SNP rs896854 in Europeans (r2 = 0.21 in CEU, 0.10 in ASW). Neither the reported index SNP rs896854 nor its proxies from the CEU data demonstrated significant association to T2D in African Americans (Table S6 and Figure S3a,b), suggesting that rs17359493 may be an independent novel signal. INTS8 encodes a subunit of the integrator complex which is involved in the cleavage of small nuclear RNAs. At KCNQ1, the most strongly associated SNP rs231356 was in weak LD with the index SNP rs231362 reported in Europeans [3] (r2 = 0.24 in CEU and 0.17 in ASW). Given rs231362 was modestly associated with T2D in African American (P = 0.04) and was in weak LD (r2 = 0.21 to 0.46 in CEU) with other associated SNPs in this region (Table S6 and Figure S3c,d), the results suggest a refinement of the localization of causal variant(s) to variants in strong LD with rs231356. At HMGA2, the most strongly associated SNP rs343092 was in moderate LD with the index SNP rs1531343 (r2 = 0.60 in CEU and 0.32 in ASW). Despite rs1531343 and its proxies in high LD were not associated with T2D in African Americans (P>0.05), several SNPs in moderate LD, including rs343092, showed nominal to strong associations (Table S6 and Figure S3e,f). Trans-ethic fine mapping will be particularly useful to dissect the causal variant(s) at this locus.
Effect of obesity on T2D susceptibility loci
We investigated the influence of obesity by comparing the stage 1 meta-analysis results with or without adjustment for BMI at the 51 most significantly associated SNPs from the GWAS for follow up (Tables S4 and S7) and 158 established T2D or glucose homeostasis index SNPs (Table S5). Association results were highly similar with and without BMI adjustment (correlation coefficients were 0.99 for both effect sizes and −logP values). Of particular note, FTO is suggested to influence T2D primarily through modulation of adiposity in Europeans [3], [25], but evidence is contradictory across multiple ethnic groups [26]–[28]. The index SNP rs11642841 was not significantly associated with T2D in African Americans without and with BMI adjustment (P = 0.06 and 0.23, respectively) (Table S5). The frequency of the risk A allele was 0.13 in this study. It had 100% power to detect association at the reported OR of 1.13 at type 1 error rate of 0.05, suggesting that FTO is unlikely a key T2D susceptibility gene in African Americans.
Gene expression and bioinformatics analyses
Among the six genome-wide significant loci (Table 1), we found no coding variants in the most significantly associated SNPs or their proxies. These SNPs demonstrated only weak associations with expression quantitative trait loci (eQTLs) (P>0.001, Table S8). Examination of the ENCODE data [29] revealed that several SNPs at TCF7L2, KCNQ1, and HMGA2 were located at protein binding sites or were predicted to alter motif affinity for transcription factors implicated in energy homeostasis (Table S9). The most strongly associated SNP rs7903146 in TCF7L2 is predicted to alter the binding affinity for a POU3F2 regulatory motif [30]. POU3F2 is a neural transcription factor that enhances the activation of genes regulated by corticotropin-releasing hormone which stimulates adrenocorticotropic hormone (ACTH). ACTH is synthesized from pre-pro-opiomelanocortin (pre-POMC) which regulates energy homeostasis. For the 3′ signal at KCNQ1, several tag SNPs are predicted to alter the binding affinity for regulatory motifs, including SREBP, CTCF and HNF4A. SREBP is a transcription factor involved in sterol biosynthesis. CTCF regulates the expression of IGF2 [31]. HNF4A is a master regulator of hepatocyte and islet transcription. The tag SNP rs2257883 at HMGA2 is predicted to alter the binding affinity of MEF2, which regulates GLUT4 transcription in insulin responsive tissues [32].
Discussion
We have performed the largest genetic association analysis to date for T2D in African Americans. Our data support the hypothesis that risk for T2D is partly attributable to a large number of common variants with small effects [7]. We identified HLA-B and INS-IGF2 as novel T2D loci, the latter specific to African Americans. We found evidence supporting association for 88 previously identified T2D and glucose homeostasis loci. Taken together, these 90 loci yielded a sibling relative risk of 1.19. The phenotypic variance measured on the liability scale is substantially larger in African Americans than in European Americans (17.5% vs. 5.7%) [7] due to larger effect sizes upon fine-mapping as well as higher disease prevalence in African Americans.
The two novel T2D loci, HLA-B and INS-IGF2, have been implicated in type 1 diabetes (T1D) risk in Europeans [33]–[35]. One limitation of our study is the lack of autoantibody measurement. However, our results are unlikely to be confounded by the presence of misclassified patients. Among diabetic youth aged <20 years, T2D characterized by insulin resistance without autoimmunity is more prevalent in African Americans (40.1%) than in European Americans (6.2%), while African Americans less often present with autoimmunity and insulin deficiency resembling T1D compared to European Americans (32.5% vs. 62.9%, respectively) [36]. Autoimmunity is also uncommon in African American diabetic adults [37]. Furthermore, associations for T1D are stronger at HLA class II (HLA-DRB1, -DQA1, and -DQB1) than HLA class I regions in Europeans [33]–[34], [38]–[41] (http://www.t1dbase.org). In African Americans, T1D individuals showed both shared and unique risk and protective HLA class II haplotypes as compared to European T1D individuals [42]–[43]. More importantly, these individuals also showed substantially stronger associations at HLA class II (P<1×10−25) than class I regions (P<1×10−5) [42], which is in contradiction with our finding of stronger associations at HLA class I than class II regions in T2D individuals (HLA-B, Figure S4). The observed HLA-B association may be due to LD with nearby causal gene(s) since there is long range LD in this region. Recently, rs3130501 near POU5F1 and TCF19 was reported for association with T2D in a trans-ancestry meta-analysis [8]. rs3130501 was located 211 kb upstream of rs2244020 and mapped to the same LD interval. However, the two SNPs were not correlated in both CEU (D′ = 0.57, r2 = 0.05) and ASW (D′ = 0.68, r2 = 0.16) from 1KGP nor strongly associated with T2D in the stage 1 meta-analysis (P = 0.04). Other potential non-HLA candidate genes may include TNFA which regulates immune and inflammatory response. It has been hypothesized that activated innate and adaptive immune cells stimulate release of cytokines such as TNFα and IL-1β, which promote both systemic insulin resistance and β-cell damage [44]. On the other hand, evidence has implicated T1D loci HLA-DQ/DR, GLIS3 and INS in the susceptibility of latent autoimmune diabetes in adults (LADA) and/or T2D [7], [34], [45]–[46], while T2D loci such as PPARG and TCF7L2 was associated with T1D [47] and LADA [46], [48], respectively. More comprehensive studies are needed to understand the shared and distinct genetic risks in different forms of diabetes which will facilitate diagnosis and personalized treatment.
Our results have several implications regarding the genetic architecture of T2D. First, fine-mapping suggests that currently known loci explain more of the risk than previously estimated. Second, the loci conferring the largest risk for T2D appear to act through regulatory rather than protein-coding changes. Third, many, but not all, of the previously identified T2D loci are shared across ancestries. The differential LD structure of African-ancestry populations at shared loci provides an opportunity for fine mapping in trans-ethnic meta-analysis. Fourth, the ∼2.6M MEDIA SNPs achieved only 43.3% coverage of the 1KGP ASW common SNPs, suggesting that risk loci that are specific to African-ancestry individuals are difficult to discover with the genotyping arrays being used. Large-scale sequencing studies, such as those focusing on whole genomes, exomes, and targeted resequencing for associated non-coding regions, will be necessary to further delineate the causal variants for T2D risk in African Americans.
Materials and Methods
Samples and clinical characterization
Stage 1 discovery samples included 17 T2D GWAS studies (ARIC, CARDIA, CFS, CHS, FamHS, GeneSTAR, GENOA, HANDLS, Health ABC, HUFS, JHS, MESA, MESA Family, SIGNET-REGARDS, WFSM, FIND, and WHI) with up to 23,827 African American subjects (8,284 cases and 15,543 controls). Stage 2 replication samples included up to 11,544 African American subjects (6,061 cases and 5,483 controls), using in silico replication of GWAS data from eMERGE and IPM Biobank and de novo genotyping in IRAS, IRASFS, SCCS, and WFSM. In general, T2D cases were defined as having at least one of the following: fasting plasma glucose ≥126 mg/dl, 2 hour glucose during oral glucose tolerance test (OGTT) ≥200 mg/dl, random glucose ≥200 mg/dl, oral hypoglycemic agent or insulin treatment, or physician-diagnosed diabetes. All cases were diagnosed at ≥25 years (or age at study ≥25 years if age at diagnosis was not available). For cohort studies, individuals who met the criteria at any of the visits were defined as cases. Controls with normal glucose tolerance (NGT) were defined by satisfying all the following criteria: fasting plasma glucose <100 mg/dl, 2 hour OGTT<140 mg/dl (if available), no treatment of diabetes, and age ≥25 years. For cohort studies, individuals who met the criteria at all visits were defined as controls. All study participants provided written informed consent, except for eMERGE that use an opt out program, and approval was obtained from the institutional review board (IRB) from the respective local institutions. Detailed descriptions of the participating studies are provided in Text S1.
Genotyping, imputation and quality control
For stage 1 and 2 GWAS studies, genotyping was performed with Affymetrix or Illumina genome-wide SNP arrays. Imputation of missing genotypes was performed using MACH [49], IMPUTE2 [50] or BEAGLE [51] using HapMap reference haplotypes. For each study, samples reflecting duplicates, low call rate, gender mismatch, or population outliers were excluded. In general, SNPs were excluded by the following criteria: call rate <0.95, minor allele frequency (MAF)<0.01, minor allele count <10, Hardy-Weinberg P-value <1×10−4, or imputation quality score <0.5 (Table S3). For de novo replication studies, genotyping was performed using the Sequenom MassArray platform (Sequenom; San Diego, CA). Sample and SNP quality controls were performed as with GWAS data.
Statistical analysis
Single SNP association was performed for each study by regressing T2D case/control status on genotypes. To account for uncertainty of genotype calls during imputation, genotype probabilities or dosage were used for association tests in imputed SNPs. The association tests assumed an additive genetic model and adjusted for age, sex, study centers, and principal components. Principal components were included to control for confounding effects of admixture proportion and population structure. Secondary analysis with additional adjustment for BMI was performed for SNPs with P<1×10−5 in stage 1 meta-analysis and index SNPs previously reported to be associated with T2D or glucose homeostasis traits. BMI adjustment allows increasing power to detect T2D loci independent of BMI effect and diminish associations at T2D loci with effects modulated through BMI. Logistic regression was used for samples of unrelated individuals. Generalized estimating equations [52] or SOLAR [53] were used for samples of related individuals. Association results with extreme values (absolute beta coefficient or standard error >10), primarily due to low cell counts resulting from small sample sizes and/or low minor allele frequencies, were excluded (Table S3).
Meta-analysis
In stage 1, association results were combined by a fixed effect model with inverse variance weighted method using the METAL software [12]. Genomic control correction [11] was applied to each study before meta-analysis, and to the overall results after meta-analysis. Results from SNPs genotyped in <10,000 samples and those with allele frequency difference >0.3 among studies were excluded. A total of 2,579,389 SNPs were analyzed in the meta-analysis (Table S3). In stage 2a, association results from African American replication studies were also combined using a fixed effect inverse variance weighted method. To assess the overall effects in African Americans (stage 1+2a) and both African Americans and Europeans (stage 1+2a+2b), association results from studies in the respective stages were combined using a fixed effect inverse variance weighted method. Genome-wide significance is declared at P<5×10−8 from the meta-analysis result of all stages, which has better power than the replication-based strategy [54].
Among the 51 SNPs carried forward for replication, heterogeneity of effect sizes across studies within each stage was assessed using Cochran's Q statistic implemented in METAL. Meta-analysis results from stages 1 and 2a, stage 1+2a and 2b were used to assess heterogeneity of effect sizes between discovery and replication stages in African Americans, and between African Americans and Europeans, respectively. For SNPs with significant heterogeneous effect size after multiple comparison corrections (Phet<0.001), meta-analysis results including studies of all stages assessed by the random effect model implemented in GWAMA [55] were reported. Heterogeneous associations may partly due to differences in ascertainment scheme across studies. For index SNPs reported in prior studies, assessment of heterogeneity using Cochran's Q statistic between prior studies and this study were also reported.
Transferability analysis
Index SNPs associated with T2D or glucose homeostasis traits from prior GWAS and candidate gene studies were examined for association with T2D in African Americans (Table S5). For the index SNP association tests, a per-SNP P value <0.05 was defined as significant. In the locus-wide analysis, the boundaries of a locus were defined by the most distant markers (within ±500 kb) using the 1KGP CEU data with r2≥0.3 with the index SNP. All MEDIA SNPs within these bounds were examined for association analysis. All pairwise LD values within each locus were estimated using the 1KGP CEU and ASW data. To estimate the effective number of SNPs at a locus, we retrieved genotypes from the 1KGP ASW data for markers present in MEDIA, estimated the sample covariance matrix from those genotypes, and spectrally decomposed the covariance matrix [24]. The effective number of SNPs was estimated using the relationship , in which λk is the kth eigenvalue of the K×K covariance matrix for the K SNPs in the locus [24]. The per-locus significance level was defined as 0.05/effective number of SNPs (Table S6). By accounting for all SNPs within the bounds of LD, the per-locus significance level is corrected to account for markers in LD with the index SNP as well as markers not in LD with the index SNP, thereby potentially allowing for discovery of new associations at markers not tagged by the index SNP.
Liability-scale variance explained
For each independent locus, we estimated the sibling relative risk using the most strongly associated SNP within that locus. Let pi and ψi be the risk allele frequency and the corresponding odds ratio at the ith SNP, respectively. Assuming the additive genetic model and independence between SNPs, the contribution to the sibling relative risk λs for a set of N SNPs is given by [56]. Let K be the disease prevalence. The liability-scale variance explained by the set of N SNPs is given by , in which , , and , with representing the standard normal quantile function and z representing the standard normal density at T [57].
Coverage
The coverage of MEDIA SNPs to the human genome was estimated using HaploView [58] via pairwise tagging at the r2 = 0.8 threshold. We used all SNPs with minor allele frequencies ≥1% in both MEDIA and the 1KGP ASW sequence data. Coverage was estimated using non-overlapping bins of 1,000 SNPs.
Power analysis
Study power was calculated using the genetic power calculator [59]. For SNPs with MAF≥0.3, our study had >80% power to detect odds ratios for T2D at OR≥1.06 and ≥1.13 at P<0.05 and P<5×10−8, respectively, in stage 1 samples under an additive model. The observed odds ratios among our stage 1 most significantly associated SNPs with P<1×10−5 ranged from 1.11 to 1.56 (Table S4). Given our African American sample size in stage 1+2a, our study had >80% power to detect OR≥1.1 at P<5×10−8 at MAF≥0.3, thus provided good power to detect genome-wide significance among the most significantly associated SNPs using all African American samples. For T2D SNPs reported from the literature, power was also calculated from the reported effect size using the risk allele frequency from this study for stage 1 samples at P<0.05 and P<5×10−8, respectively (Table S5).
Gene expression analysis
The MuTHER resource (www.muther.ac.uk) includes lymphoblastoid cell lines (LCLs), skin, and adipose tissue derived simultaneously from a subset of well-phenotyped healthy female twins from the TwinsUK adult registry [60]. Whole-genome expression profiling of the samples, each with either two or three technical replicates, was performed using the Illumina Human HT-12 V3 BeadChips (Illumina Inc.) according to the protocol supplied by the manufacturer. Log2-transformed expression signals were normalized separately per tissue as follows: quantile normalization was performed across technical replicates of each individual followed by quantile normalization across all individuals. Genotyping was performed with a combination of Illumina arrays (HumanHap300, HumanHap610Q, 1M-Duo, and 1.2MDuo 1M). Untyped HapMap2 SNPs were imputed using the IMPUTE2 software package. In total, 776 adipose and 777 LCL samples had both expression profiles and imputed genotypes. Association between all SNPs (MAF>5%, IMPUTE info >0.8) within a gene or within 1 Mb of the gene transcription start or end site and normalized expression values were performed with the GenABEL/ProbABEL packages [61]–[62] using the polygenic linear model incorporating a kinship matrix in GenABEL followed by the ProbABEL mmscore score test with imputed genotypes. Age and experimental batch were included as cofactors.
Genotype and gene expression in LCL in HapMap samples were also available [63]. Association of genotypes and gene expression of transcripts within 1 MB of tested SNPs were analyzed separately for CEU and YRI populations. The variance components model implemented in SOLAR was used for association analysis which accounts for correlation among related individuals [53].
In this study, we examined the association of the most significantly associated SNPs from the six genome-wide significant loci and their proxies (r2≥0.8 in ASW) within 1 Mb of the associated SNPs with cis-expression quantitative trait loci (eQTLs) in peripheral blood leukocytes (LCL) and adipose tissue (Table S8).
ENCODE data analysis
We examined putative function of non-coding genome-wide significant SNPs and their proxies within 1 Mb (r2≥0.8 in 1KGP ASW) using HaploReg [30] and RegulomeDB [64]. These databases interrogated multiple chromatin features from the Encyclopedia of DNA Elements (ENCODE) project [29]. High priority was given to variants annotated as protein-binding via ChIP-seq, and motif-changing via position weight matrices, with the respective transcription factors implicated in diabetes pathogenesis and related biological processes.
Supporting Information
Zdroje
1. Centers for Disease Control and Prevention (2011) National diabetes fact sheet: National estimates and general information on diabetes and prediabetes in the United States. Atlanta, GA: U.S. Department of Health and Human Services, Centers for Disease Control and Prevention.
2. McCarthyMI (2010) Genomics, type 2 diabetes, and obesity. N Engl J Med 363 : 2339–2350.
3. VoightBF, ScottLJ, SteinthorsdottirV, MorrisAP, DinaC, et al. (2010) Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat Genet 42 : 579–589.
4. KoonerJS, SaleheenD, SimX, SehmiJ, ZhangW, et al. (2011) Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat Genet 984–989.
5. ChoYS, ChenCH, HuC, LongJ, Hee OngRT, et al. (2011) Meta-analysis of genome-wide association studies identifies eight new loci for type 2 diabetes in East Asians. Nat Genet 44 : 67–72.
6. PalmerND, McDonoughCW, HicksPJ, RohBH, WingMR, et al. (2012) A genome-wide association search for type 2 diabetes genes in African Americans. PLoS ONE 7: e29202.
7. MorrisAP, VoightBF, TeslovichTM, FerreiraT, SegreAV, et al. (2012) Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet 44 : 981–990.
8. DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium, Asian Genetic Epidemiology Network Type 2 Diabetes (AGEN-T2D) Consortium, South Asian Type 2 Diabetes (SAT2D) Consortium, Mexican American Type 2 Diabetes (MAT2D) Consortium, Type 2 Diabetes Genetic Exploration by Next-generation sequencing in multi-Ethnic Samples (T2D-GENES) Consortium (2014) Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet 46 : 234–244.
9. RotimiC, CooperR, CaoG, SundarumC, McGeeD (1994) Familial aggregation of cardiovascular diseases in African-American pedigrees. Genet Epidemiol 11 : 397–407.
10. AbecasisGR, AutonA, BrooksLD, DePristoMA, DurbinRM, et al. (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491 : 56–65.
11. DevlinB, RoederK (1999) Genomic control for association studies. Biometrics 55 : 997–1004.
12. WillerCJ, LiY, AbecasisGR (2010) METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26 : 2190–2191.
13. HaraK, FujitaH, JohnsonTA, YamauchiT, YasudaK, et al. (2014) Genome-wide association study identifies three novel loci for type 2 diabetes. Hum Mol Genet 23 : 239–246.
14. Genomes Project Consortium (2010) A map of human genome variation from population-scale sequencing. Nature 467 : 1061–1073.
15. UnokiH, TakahashiA, KawaguchiT, HaraK, HorikoshiM, et al. (2008) SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat Genet 40 : 1098–1102.
16. YasudaK, MiyakeK, HorikawaY, HaraK, OsawaH, et al. (2008) Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat Genet 40 : 1092–1097.
17. TakeuchiF, SerizawaM, YamamotoK, FujisawaT, NakashimaE, et al. (2009) Confirmation of multiple risk loci and genetic impacts by a genome-wide association study of type 2 diabetes in the Japanese population. Diabetes 58 : 1690–1699.
18. ParraEJ, BelowJE, KrithikaS, ValladaresA, BartaJL, et al. (2011) Genome-wide association study of type 2 diabetes in a sample from Mexico City and a meta-analysis of a Mexican-American sample from Starr County, Texas. Diabetologia 54 : 2038–2046.
19. CuiB, ZhuX, XuM, GuoT, ZhuD, et al. (2011) A genome-wide association study confirms previously reported loci for type 2 diabetes in Han Chinese. PLoS ONE 6: e22353.
20. TsaiFJ, YangCF, ChenCC, ChuangLM, LuCH, et al. (2010) A genome-wide association study identifies susceptibility variants for type 2 diabetes in Han Chinese. PLoS Genet 6: e1000847.
21. StoyJ, SteinerDF, ParkSY, YeH, PhilipsonLH, et al. (2010) Clinical and molecular genetics of neonatal diabetes due to mutations in the insulin gene. Rev Endocr Metab Disord 11 : 205–215.
22. PetrikJ, PellJM, AranyE, McDonaldTJ, DeanWL, et al. (1999) Overexpression of insulin-like growth factor-II in transgenic mice is associated with pancreatic islet cell hyperplasia. Endocrinology 140 : 2353–2363.
23. CalderariS, GangnerauMN, ThibaultM, MeileMJ, KassisN, et al. (2007) Defective IGF2 and IGF1R protein production in embryonic pancreas precedes beta cell mass anomaly in the Goto-Kakizaki rat model of type 2 diabetes. Diabetologia 50 : 1463–1471.
24. RamosE, ChenG, ShrinerD, DoumateyA, GerryNP, et al. (2011) Replication of genome-wide association studies (GWAS) loci for fasting plasma glucose in African-Americans. Diabetologia 54 : 783–788.
25. FraylingTM, TimpsonNJ, WeedonMN, ZegginiE, FreathyRM, et al. (2007) A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316 : 889–894.
26. HertelJK, JohanssonS, SonestedtE, JonssonA, LieRT, et al. (2011) FTO, type 2 diabetes, and weight gain throughout adult life: a meta-analysis of 41,504 subjects from the Scandinavian HUNT, MDC, and MPP studies. Diabetes 60 : 1637–1644.
27. LiH, KilpelainenTO, LiuC, ZhuJ, LiuY, et al. (2012) Association of genetic variation in FTO with risk of obesity and type 2 diabetes with data from 96,551 East and South Asians. Diabetologia 55 : 981–995.
28. BinhTQ, PhuongPT, NhungBT, ThoangDD, LienHT, et al. (2013) Association of the common FTO-rs9939609 polymorphism with type 2 diabetes, independent of obesity-related traits in Vietnamese population. Gene 513 : 31–35.
29. DunhamI, KundajeA, AldredSF, CollinsPJ, DavisCA, et al. (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489 : 57–74.
30. WardLD, KellisM (2012) HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 40: D930–934.
31. BellAC, FelsenfeldG (2000) Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405 : 482–485.
32. OshelKM, KnightJB, CaoKT, ThaiMV, OlsonAL (2000) Identification of a 30-base pair regulatory element and novel DNA binding protein that regulates the human GLUT4 promoter in transgenic mice. J Biol Chem 275 : 23666–23673.
33. Wellcome Trust Case Control Consortium (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447 : 661–678.
34. BarrettJC, ClaytonDG, ConcannonP, AkolkarB, CooperJD, et al. (2009) Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet 41 : 703–707.
35. PlagnolV, HowsonJM, SmythDJ, WalkerN, HaflerJP, et al. (2011) Genome-wide association analysis of autoantibody positivity in type 1 diabetes cases. PLoS Genet 7: e1002216.
36. DabeleaD, PihokerC, TaltonJW, D'AgostinoRBJr, FujimotoW, et al. (2011) Etiological approach to characterization of diabetes type: the SEARCH for Diabetes in Youth Study. Diabetes Care 34 : 1628–1633.
37. Barinas-MitchellE, PietropaoloS, ZhangYJ, HendersonT, TruccoM, et al. (2004) Islet cell autoimmunity in a triethnic adult population of the Third National Health and Nutrition Examination Survey. Diabetes 53 : 1293–1302.
38. HakonarsonH, GrantSF, BradfieldJP, MarchandL, KimCE, et al. (2007) A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 448 : 591–594.
39. ErlichH, ValdesAM, NobleJ, CarlsonJA, VarneyM, et al. (2008) HLA DR-DQ haplotypes and genotypes and type 1 diabetes risk: analysis of the type 1 diabetes genetics consortium families. Diabetes 57 : 1084–1092.
40. HowsonJM, WalkerNM, ClaytonD, ToddJA (2009) Confirmation of HLA class II independent type 1 diabetes associations in the major histocompatibility complex including HLA-B and HLA-A. Diabetes Obes Metab 11 Suppl 1 : 31–45.
41. EikeMC, BeckerT, HumphreysK, OlssonM, LieBA (2009) Conditional analyses on the T1DGC MHC dataset: novel associations with type 1 diabetes around HLA-G and confirmation of HLA-B. Genes Immun 10 : 56–67.
42. HowsonJM, RoyMS, ZeitelsL, StevensH, ToddJA (2013) HLA class II gene associations in African American Type 1 diabetes reveal a protective HLA-DRB1*03 haplotype. Diabet Med 30 : 710–716.
43. NobleJA, JohnsonJ, LaneJA, ValdesAM (2013) HLA class II genotyping of African American type 1 diabetes patients reveals associations unique to African haplotypes. Diabetes 62 : 3292–3299.
44. OdegaardJI, ChawlaA (2012) Connecting type 1 and type 2 diabetes through innate immunity. Cold Spring Harb Perspect Med 2: a007724.
45. RichSS, FrenchLR, SprafkaJM, ClementsJP, GoetzFC (1993) HLA-associated susceptibility to type 2 (non-insulin-dependent) diabetes mellitus: the Wadena City Health Study. Diabetologia 36 : 234–238.
46. CervinC, LyssenkoV, BakhtadzeE, LindholmE, NilssonP, et al. (2008) Genetic similarities between latent autoimmune diabetes in adults, type 1 diabetes, and type 2 diabetes. Diabetes 57 : 1433–1437.
47. RajSM, HowsonJM, WalkerNM, CooperJD, SmythDJ, et al. (2009) No association of multiple type 2 diabetes loci with type 1 diabetes. Diabetologia 52 : 2109–2116.
48. LukacsK, HosszufalusiN, DinyaE, BakacsM, MadacsyL, et al. (2012) The type 2 diabetes-associated variant in TCF7L2 is associated with latent autoimmune diabetes in adult Europeans and the gene effect is modified by obesity: a meta-analysis and an individual study. Diabetologia 55 : 689–693.
49. LiY, WillerCJ, DingJ, ScheetP, AbecasisGR (2010) MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 34 : 816–834.
50. HowieBN, DonnellyP, MarchiniJ (2009) A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5: e1000529.
51. BrowningSR, BrowningBL (2007) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81 : 1084–1097.
52. ChenMH, YangQ (2010) GWAF: an R package for genome-wide association analyses with family data. Bioinformatics 26 : 580–581.
53. AlmasyL, BlangeroJ (1998) Multipoint quantitative-trait linkage analysis in general pedigrees. Am J Hum Genet 62 : 1198–1211.
54. SkolAD, ScottLJ, AbecasisGR, BoehnkeM (2006) Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat Genet 38 : 209–213.
55. MagiR, MorrisAP (2010) GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11 : 288.
56. LinS, ChakravartiA, CutlerDJ (2004) Exhaustive allelic transmission disequilibrium tests as a new approach to genome-wide association studies. Nat Genet 36 : 1181–1188.
57. WrayNR, YangJ, GoddardME, VisscherPM (2010) The genetic interpretation of area under the ROC curve in genomic profiling. PLoS Genet 6: e1000864.
58. BarrettJC, FryB, MallerJ, DalyMJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21 : 263–265.
59. PurcellS, ChernySS, ShamPC (2003) Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19 : 149–150.
60. NicaAC, PartsL, GlassD, NisbetJ, BarrettA, et al. (2011) The architecture of gene regulatory variation across multiple human tissues: the MuTHER study. PLoS Genet 7: e1002003.
61. AulchenkoYS, RipkeS, IsaacsA, van DuijnCM (2007) GenABEL: an R library for genome-wide association analysis. Bioinformatics 23 : 1294–1296.
62. AulchenkoYS, StruchalinMV, van DuijnCM (2010) ProbABEL package for genome-wide association analysis of imputed data. BMC Bioinformatics 11 : 134.
63. StrangerBE, ForrestMS, DunningM, IngleCE, BeazleyC, et al. (2007) Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315 : 848–853.
64. BoyleAP, HongEL, HariharanM, ChengY, SchaubMA, et al. (2012) Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 22 : 1790–1797.
65. Wellcome Trust Case Control Consortium (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447 : 661–678.
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 8- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- DNA Double Strand Break Repair in Mitosis Is Suppressed by Phosphorylation of XRCC4
- Inference of Transposable Element Ancestry
- The Population Genetics of Evolutionary Rescue
- Retinoic Acid Activates Two Pathways Required for Meiosis in Mice
- Pooled Segregant Sequencing Reveals Genetic Determinants of Yeast Pseudohyphal Growth
- Comprehensive Identification of Single Nucleotide Polymorphisms Associated with Beta-lactam Resistance within Pneumococcal Mosaic Genes
- SMA-Causing Missense Mutations in Display a Wide Range of Phenotypes When Modeled in
- Branch Migration Prevents DNA Loss during Double-Strand Break Repair
- Transcriptome Sequencing from Diverse Human Populations Reveals Differentiated Regulatory Architecture
- Genetic Deletion of SEPT7 Reveals a Cell Type-Specific Role of Septins in Microtubule Destabilization for the Completion of Cytokinesis
- Tethering Sister Centromeres to Each Other Suggests the Spindle Checkpoint Detects Stretch within the Kinetochore
- Global Genetic Variations Predict Brain Response to Faces
- Demography and the Age of Rare Variants
- The Response to High CO Levels Requires the Neuropeptide Secretion Component HID-1 to Promote Pumping Inhibition
- Sp6 and Sp8 Transcription Factors Control AER Formation and Dorsal-Ventral Patterning in Limb Development
- The Groucho Co-repressor Is Primarily Recruited to Local Target Sites in Active Chromatin to Attenuate Transcription
- A Transposable Element Insertion Confers Xenobiotic Resistance in Drosophila
- The Genomic Architecture of Population Divergence between Subspecies of the European Rabbit
- Human Social Genomics
- Gene Expansion Shapes Genome Architecture in the Human Pathogen : An Evolutionary Genomics Analysis in the Ancient Terrestrial Mucorales (Mucoromycotina)
- Canonical Non-Homologous End Joining in Mitosis Induces Genome Instability and Is Suppressed by M-phase-Specific Phosphorylation of XRCC4
- Roles of Type 1A Topoisomerases in Genome Maintenance in
- The TRIM-NHL Protein LIN-41 Controls the Onset of Developmental Plasticity in
- Wnt-Mediated Repression via Bipartite DNA Recognition by TCF in the Hematopoietic System
- KDM6 Demethylase Independent Loss of Histone H3 Lysine 27 Trimethylation during Early Embryonic Development
- Integration of UPR and Oxidative Stress Signaling in the Control of Intestinal Stem Cell Proliferation
- miR171-Targeted Scarecrow-Like Proteins Bind to GT -Elements and Mediate Gibberellin-Regulated Chlorophyll Biosynthesis under Light Conditions
- Syndecan-1 Is Required to Maintain Intradermal Fat and Prevent Cold Stress
- LIN-3/EGF Promotes the Programmed Cell Death of Specific Cells in by Transcriptional Activation of the Pro-apoptotic Gene
- A System for Genome-Wide Histone Variant Dynamics In ES Cells Reveals Dynamic MacroH2A2 Replacement at Promoters
- Multiple Regulation of Rad51-Mediated Homologous Recombination by Fission Yeast Fbh1
- A Genome-Wide Association Study of the Maize Hypersensitive Defense Response Identifies Genes That Cluster in Related Pathways
- The RNA Helicases AtMTR4 and HEN2 Target Specific Subsets of Nuclear Transcripts for Degradation by the Nuclear Exosome in
- Asymmetric Division and Differential Gene Expression during a Bacterial Developmental Program Requires DivIVA
- A Model-Based Approach for Identifying Signatures of Ancient Balancing Selection in Genetic Data
- Chromatin Insulator Factors Involved in Long-Range DNA Interactions and Their Role in the Folding of the Drosophila Genome
- Conditional Inactivation of Upstream Binding Factor Reveals Its Epigenetic Functions and the Existence of a Somatic Nucleolar Precursor Body
- Evidence for Divisome Localization Mechanisms Independent of the Min System and SlmA in
- Patterns of Admixture and Population Structure in Native Populations of Northwest North America
- Response Regulator Heterodimer Formation Controls a Key Stage in S Development
- A Genetic Strategy to Measure Circulating Insulin Reveals Genes Regulating Insulin Production and Secretion
- EVA-1 Functions as an UNC-40 Co-receptor to Enhance Attraction to the MADD-4 Guidance Cue in
- Dysfunction of the CNS-Heart Axis in Mouse Models of Huntington's Disease
- An Otx/Nodal Regulatory Signature for Posterior Neural Development in Ascidians
- Phosphorylation of a Central Clock Transcription Factor Is Required for Thermal but Not Photic Entrainment
- Genome-Wide Patterns of Genetic Variation within and among Alternative Selective Regimes
- EF-P Dependent Pauses Integrate Proximal and Distal Signals during Translation
- Ku-Mediated Coupling of DNA Cleavage and Repair during Programmed Genome Rearrangements in the Ciliate
- Functional Specialization Among Members Of Knickkopf Family Of Proteins In Insect Cuticle Organization
- Playing RNase P Evolution: Swapping the RNA Catalyst for a Protein Reveals Functional Uniformity of Highly Divergent Enzyme Forms
- The Translational Regulators GCN-1 and ABCF-3 Act Together to Promote Apoptosis in
- Meta-Analysis of Genome-Wide Association Studies in African Americans Provides Insights into the Genetic Architecture of Type 2 Diabetes
- A -Regulatory Mutation of Causes Silky-Feather in Chickens
- VIB1, a Link between Glucose Signaling and Carbon Catabolite Repression, Is Essential for Plant Cell Wall Degradation by
- A Population Genetic Signal of Polygenic Adaptation
- A Conserved Dopamine-Cholecystokinin Signaling Pathway Shapes Context–Dependent Behavior
- The MAP Kinase p38 Is Part of Circadian Clock
- The Cohesin Subunit Rad21 Is Required for Synaptonemal Complex Maintenance, but Not Sister Chromatid Cohesion, during Drosophila Female Meiosis
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Meta-Analysis of Genome-Wide Association Studies in African Americans Provides Insights into the Genetic Architecture of Type 2 Diabetes
- KDM6 Demethylase Independent Loss of Histone H3 Lysine 27 Trimethylation during Early Embryonic Development
- The RNA Helicases AtMTR4 and HEN2 Target Specific Subsets of Nuclear Transcripts for Degradation by the Nuclear Exosome in
- EF-P Dependent Pauses Integrate Proximal and Distal Signals during Translation
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



