-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariérní portál
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaThe Development of Therapeutic Antibodies That Neutralize Homologous and Heterologous Genotypes of Dengue Virus Type 1
Antibody protection against flaviviruses is associated with the development of neutralizing antibodies against the viral envelope (E) protein. Prior studies with West Nile virus (WNV) identified therapeutic mouse and human monoclonal antibodies (MAbs) that recognized epitopes on domain III (DIII) of the E protein. To identify an analogous panel of neutralizing antibodies against DENV type-1 (DENV-1), we immunized mice with a genotype 2 strain of DENV-1 virus and generated 79 new MAbs, 16 of which strongly inhibited infection by the homologous virus and localized to DIII. Surprisingly, only two MAbs, DENV1-E105 and DENV1-E106, retained strong binding and neutralizing activity against all five DENV-1 genotypes. In an immunocompromised mouse model of infection, DENV1-E105 and DENV1-E106 exhibited therapeutic activity even when administered as a single dose four days after inoculation with a heterologous genotype 4 strain of DENV-1. Using epitope mapping and X-ray crystallographic analyses, we localized the neutralizing determinants for the strongly inhibitory MAbs to distinct regions on DIII. Interestingly, sequence variation in DIII alone failed to explain disparities in neutralizing potential of MAbs among different genotypes. Overall, our experiments define a complex structural epitope on DIII of DENV-1 that can be recognized by protective antibodies with therapeutic potential.
Published in the journal: . PLoS Pathog 6(4): e32767. doi:10.1371/journal.ppat.1000823
Category: Research Article
doi: https://doi.org/10.1371/journal.ppat.1000823Summary
Antibody protection against flaviviruses is associated with the development of neutralizing antibodies against the viral envelope (E) protein. Prior studies with West Nile virus (WNV) identified therapeutic mouse and human monoclonal antibodies (MAbs) that recognized epitopes on domain III (DIII) of the E protein. To identify an analogous panel of neutralizing antibodies against DENV type-1 (DENV-1), we immunized mice with a genotype 2 strain of DENV-1 virus and generated 79 new MAbs, 16 of which strongly inhibited infection by the homologous virus and localized to DIII. Surprisingly, only two MAbs, DENV1-E105 and DENV1-E106, retained strong binding and neutralizing activity against all five DENV-1 genotypes. In an immunocompromised mouse model of infection, DENV1-E105 and DENV1-E106 exhibited therapeutic activity even when administered as a single dose four days after inoculation with a heterologous genotype 4 strain of DENV-1. Using epitope mapping and X-ray crystallographic analyses, we localized the neutralizing determinants for the strongly inhibitory MAbs to distinct regions on DIII. Interestingly, sequence variation in DIII alone failed to explain disparities in neutralizing potential of MAbs among different genotypes. Overall, our experiments define a complex structural epitope on DIII of DENV-1 that can be recognized by protective antibodies with therapeutic potential.
Introduction
Dengue virus (DENV) is a member of the Flaviviridae family and is related to the viruses that cause yellow fever, and the Japanese, St. Louis, and the West Nile encephalitides [1]. DENV infection after mosquito inoculation causes a spectrum of clinical disease ranging from a self-limited febrile illness (DF) to a life threatening hemorrhagic and capillary leak syndrome (Dengue Hemorrhagic Fever (DHF)/Dengue Shock Syndrome (DSS)). Globally, there is significant diversity among DENV strains, including four distinct serotypes (DENV-1, DENV-2, DENV-3, and DENV-4) that differ at the amino acid level in the viral envelope proteins by 25 to 40 percent. There is additional complexity within a given DENV serotype, as genotypes vary further by up to ∼6% and 3% at the nucleotide and amino acid levels, respectively [2],[3]. At present, no approved antiviral treatment or vaccine is available, and therapy is supportive. DENV causes an estimated 25 to 100 million infections and 250,000 cases of DHF/DSS per year worldwide, with 2.5 billion people at risk [4],[5].
DENV is an enveloped virus with a single-stranded, positive-sense RNA genome [6]. The 10.7 kilobase genome is translated as a single polyprotein, which is cleaved into three structural proteins (C, prM/M, E) and seven nonstructural (NS) proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, NS5). The mature DENV virion has a well-organized outer protein shell, a lipid membrane bilayer, and a less-defined inner nucleocapsid core [7],[8]. The ectodomains of DENV E proteins are assembled as dimers with each subunit comprised of three discrete domains [9]–[11]. Domain I (DI) is a central, eight-stranded β-barrel, which contains a single N-linked glycosylation site in most DENV strains. Domain II (DII) is a long, finger-like protrusion from DI and contains a second N-linked glycan that binds to DC-SIGN [12]–[15] and the highly conserved fusion peptide at its distal end. Domain III (DIII), which adopts an immunoglobulin-like fold, has been argued to contain a cell surface receptor recognition site [16]–[19]. Exposure to mildly acidic conditions in the trans-Golgi secretory pathway promotes virus maturation through a structural rearrangement of the flavivirus E proteins and cleavage of prM to M by a furin-like protease [20],[21]. Mature DENV virions are covered by 90 anti-parallel E protein homodimers, which are arranged flat along the surface with quasi-icosahedral symmetry.
Many flavivirus neutralizing antibodies recognize the structural E protein (reviewed in [22]). Serotype-specific MAbs against DENV reportedly have the greatest neutralizing activity [23],[24] although some sub-complex specific MAbs, which recognize some but not all DENV serotypes, also are inhibitory [25]–[27]. Protection in animals by antibodies correlates generally with the degree of neutralizing activity in vitro [24], [28]–[31]. Several type-specific strongly neutralizing antibodies against individual flaviviruses (e.g., West Nile virus (WNV), Japanese encephalitis virus (JEV), DENV-2, DENV-4, and yellow fever virus (YFV)) have been localized to epitopes in DI and DIII [26], [29], [32]–[42]. Based on their potency of neutralization and protection in vivo, anti-WNV humanized or human MAbs are in clinical development as candidate therapeutics (reviewed in [43]). Although type-specific neutralizing MAbs against DENV-1 have been characterized [44],[45], few have been mapped to specific amino acids or structural determinants. To date, only two neutralizing IgM against DENV-1 were mapped by neutralization escape selection to amino acids E279 and E293, which lie outside of DIII [46].
To develop neutralizing MAbs against DENV-1 with possible clinical potential, mice were infected with a genotype 2 strain of a DENV-1 virus, and boosted with recombinant protein of the homologous virus. 79 new MAbs against DENV-1 were generated: 15 of these were DIII-specific IgG and strongly neutralized the homologous DENV-1 strain. Of these, 9 were serotype-specific, 5 were sub-complex-specific, and only 1 MAb cross-reacted with all 4 serotypes of DENV. Several MAbs failed to neutralize at least one DENV-1 strain of a distinct genotype, suggesting that antibody recognition of the neutralizing epitopes on DIII varied among genotypes.
As a first step towards generating antibody therapeutics against DENV-1, we evaluated the protective capacity of these MAbs in an immunocompromised IFN-αβR−/− × IFN-γR−/− (AG129) mouse infection model with a heterologous DENV-1 genotype 4 strain. Among the strongly neutralizing antibodies tested, only four were highly protective. Two MAbs, DENV1-E105 and DENV1-E106, which neutralized all genotypes, were therapeutically active even when administered as a single dose four days after infection. These studies define the complexity of epitopes on DIII of DENV-1 that is recognized by highly protective antibodies with therapeutic potential.
Results
MAb generation
Previous studies with WNV and DENV-2 demonstrated that MAbs against DIII of the E protein protect against infection of cells [26],[27],[29],[32],[35],[41] and, in the case of WNV, in animals [29]. To date, few DENV-1 MAbs have been characterized functionally or mapped structurally. To develop neutralizing MAbs, mice were infected with the 16007 (genotype 2) prototype strain of DENV-1 and in some cases, boosted with recombinant DIII from the homologous DENV-1 strain. Initial immunization studies were performed with wild type BALB/c mice as done previously with WNV [29]. Because low neutralizing titers (<1/100) were achieved, likely because of poor replication of DENV-1, we switched to an immunodeficient (IFN-αβR−/−) strain in the C57BL/6 background. After screening more than 3,000 hybridoma clones as part of five independent fusions, we isolated and cloned 79 new MAbs that recognized cells infected with DENV-1 (Table S1).
All MAbs were initially tested semi-quantitatively for neutralization of the homologous DENV-1 strain by a single endpoint plaque reduction assay in Vero or BHK21-15 cells using neat hybridoma supernatant (∼10 µg/ml). Of the MAbs generated, 18 showed no inhibitory activity (<15% neutralization), 45 had modest inhibitory activity (15–75% neutralization) and 16 were strongly inhibitory (>95% neutralizing). All MAbs were screened for E protein domain recognition using yeast that expressed DENV-1 DI-II or DIII on their surface (Table S1): 14 bound to yeast expressing DI-DII, 53 recognized DIII, and the remaining 12 did not bind either DI-DII or DIII, indicating they recognized epitopes that are not present on the truncated yeast-expressed forms of DENV-1 E protein. All MAbs were tested for cross-reactivity using cells infected with different serotypes of DENV or WNV. Five of the fourteen DI-DII-specific MAbs and 26 of the 53 DIII-specific MAbs cross-reacted with other serotypes of DENV. Only 3 of the 78 MAbs bound all DENV serotypes and WNV. One of the neutralizing MAbs, DENV1-E50, was of the IgM isotype and not evaluated further here.
Characterization of strongly neutralizing MAbs against DENV-1
Studies with neutralizing mouse MAbs against DENV-2 defined a type-specific epitope on the lateral ridge of DIII (DIII-LR) and an adjacent more conserved β-strand termed the sub-complex-specific DIII A-strand epitope [25]–[27],[41]. As our single endpoint titer neutralization experiments showed that all 15 inhibitory IgG MAbs localized to DIII of DENV-1, we analyzed their reactivity patterns with other strains. We found that 9 of the 15 neutralizing IgG MAbs were serotype-specific and showed no cross-reactivity to DENV-2, DENV-3, DENV-4 or WNV-infected cells. Five MAbs were sub-complex specific and bound some but not all DENV serotypes. Only one MAb, DENV1-E102, cross-reacted with all DENV serotypes but not with WNV-infected cells (Table 1).
Tab. 1. Binding of MAbs to cells infected with other DENV serotypes. 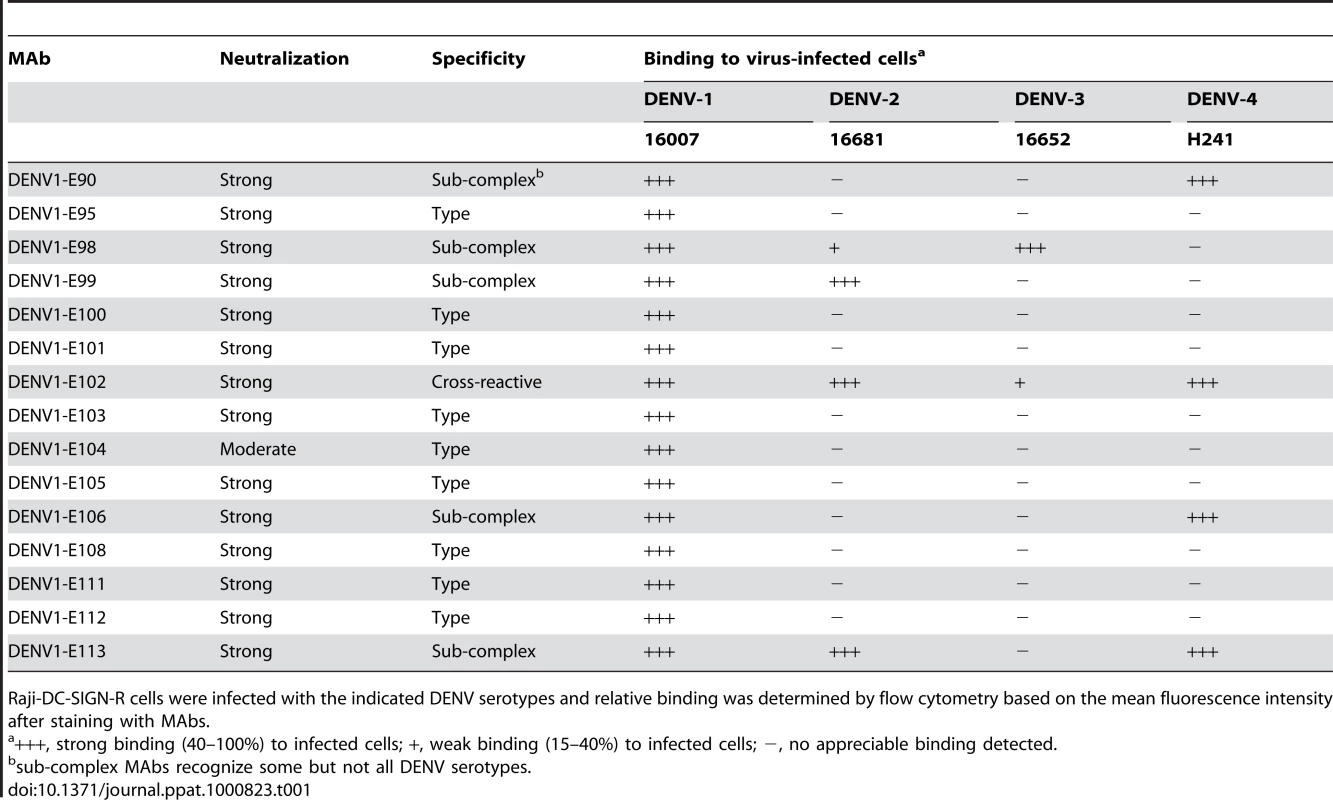
Raji-DC-SIGN-R cells were infected with the indicated DENV serotypes and relative binding was determined by flow cytometry based on the mean fluorescence intensity after staining with MAbs. Genotype-specific reactivity of anti-DENV-1 MAbs
As variation exists in the genetic composition of viruses within a given DENV serotype, an intra-serotype genotype classification system was developed [2],[3], with a given genotype of a serotype having no more than 6% nucleotide sequence divergence in the E gene [47]. To address whether the neutralizing MAbs were affected by the variation within the serotype, we tested reactivity and neutralizing potential against individual strains that encompass the five distinct DENV-1 genotypes: Asian genotype 1 (strain TVP-2130), Thai genotype 2 (strain 16007), Malaysian genotype 3 (TVP-5175), South Pacific genotype 4 (strain West-Pac 74), and American/African genotype 5 (strain 3146 SL).
(a) Binding to different DENV-1 genotypes
To determine whether antibodies recognized different genotypes of DENV-1, we infected Raji-DC-SIGN-R or C6/36 cells (depending on permissiveness) with strains of the five different genotypes of DENV-1. Infected cells were incubated with hybridoma supernatants and analyzed by flow cytometry. All neutralizing MAbs tested efficiently recognized genotypes 1, 2, 4, and 5. However, a few of the MAbs (DENV1-E100, DENV1-E104, and DENV1-E111) showed reduced binding to cells infected with the genotype 3 strain (Table 2). In contrast, other DIII-specific non-neutralizing MAbs (e.g., DENV1-E91) showed reduced binding to several of the heterologous DENV-1 genotypes (Fig 1).
Fig. 1. Binding of MAbs to cells infected with different genotypes of DENV-1. 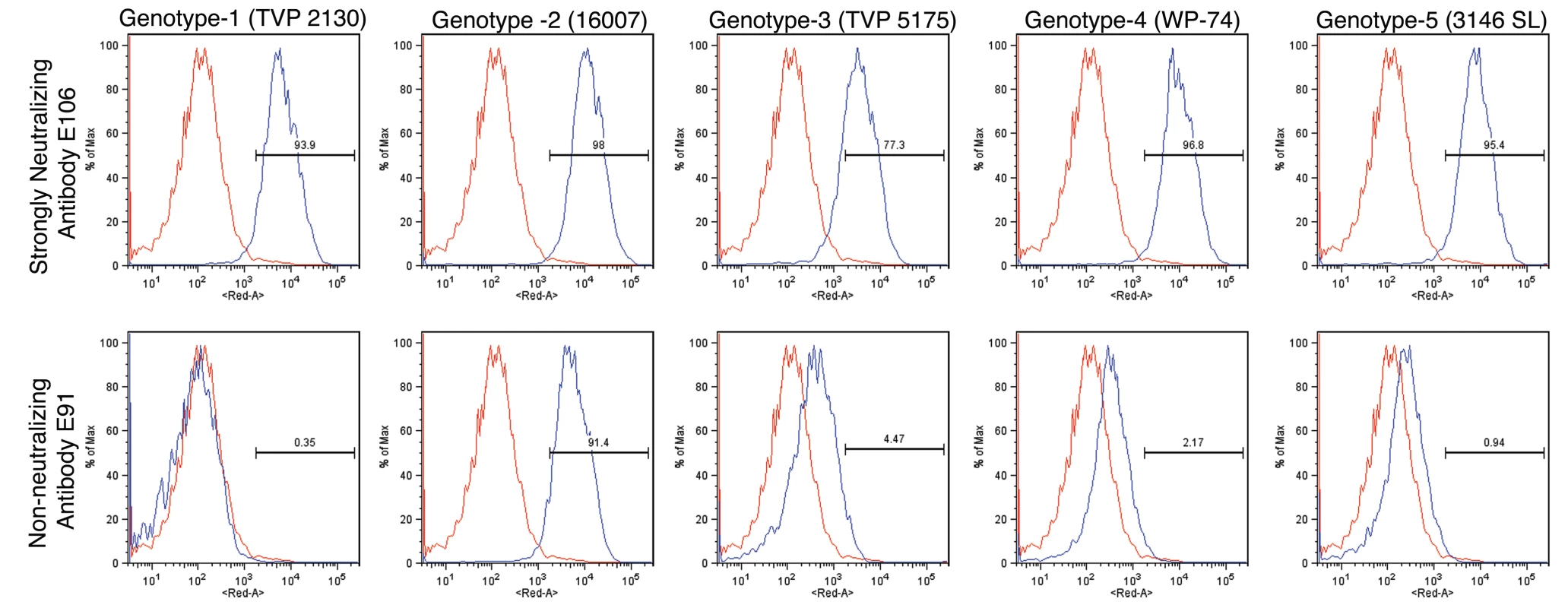
C6/36 insect cells were infected with strains of DENV-1 virus corresponding to all five genotypes. After fixation and permeabilization, MAbs were incubated with infected cells and binding was assessed by flow cytometry. The data shown are representative histograms from two independent experiments with a neutralizing (DENV1-E106, upper panels) and non-neutralizing (DENV1-E91, lower panels) MAb. Tab. 2. MAb binding to cells infected with different genotypes of DENV-1 virus. 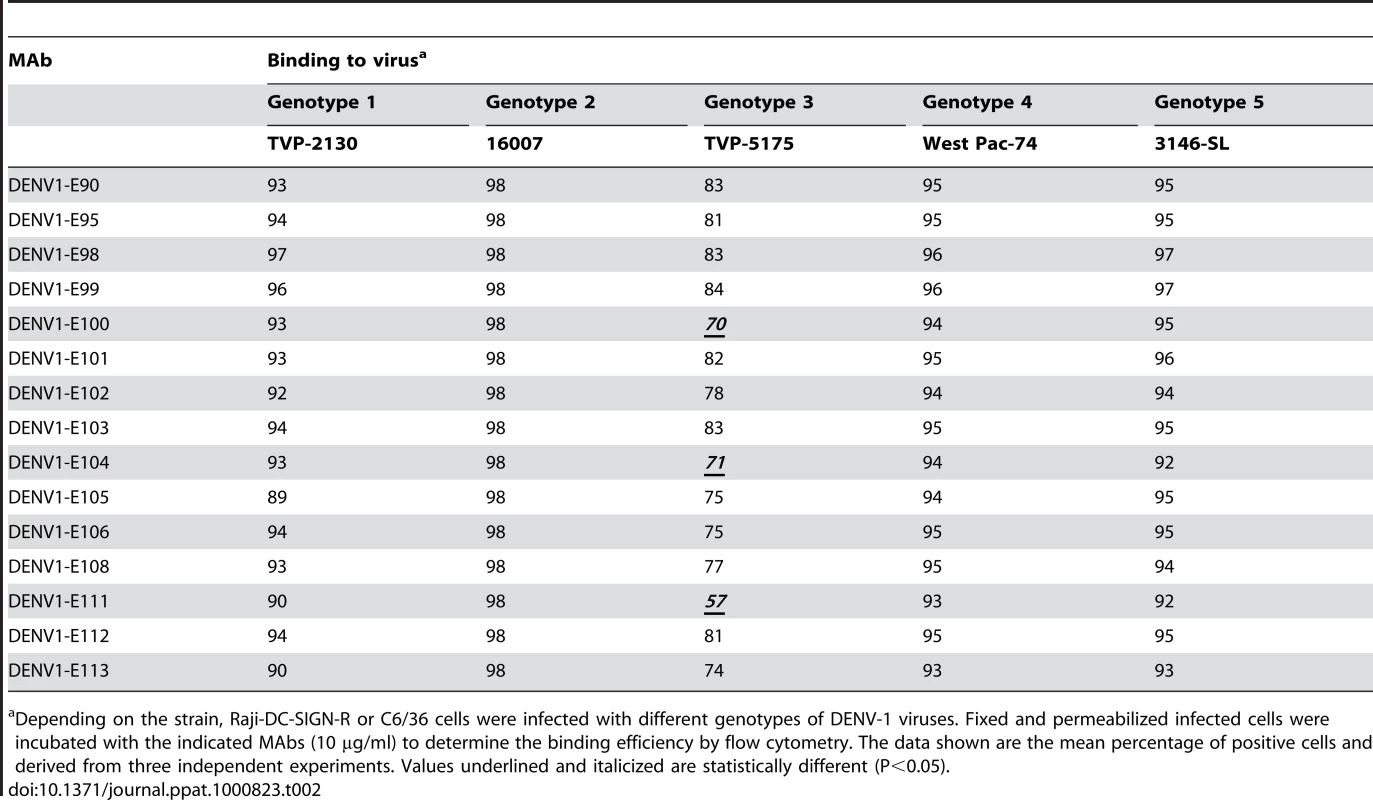
Depending on the strain, Raji-DC-SIGN-R or C6/36 cells were infected with different genotypes of DENV-1 viruses. Fixed and permeabilized infected cells were incubated with the indicated MAbs (10 µg/ml) to determine the binding efficiency by flow cytometry. The data shown are the mean percentage of positive cells and derived from three independent experiments. Values underlined and italicized are statistically different (P<0.05). (b) Neutralizing potential of MAbs against different DENV-1 genotypes
Although most MAbs retained binding to cells infected with many of the genotypes, we questioned whether this translated into efficient cross-genotypic neutralization. A recent study of neutralization escape mutants with human MAbs against WNV showed that retention of binding did not necessarily sustain neutralizing activity [48]. To evaluate this, we assessed semi-quantitatively the inhibitory activity of hybridoma supernatants of 15 neutralizing MAbs against all five genotypes by a standard plaque reduction assay with 102 PFU of virus in Vero or BHK21-15 cells, depending on the strain (Table 3). Only 8 of 15 hybridoma supernatants neutralized strains from all five genotypes by greater than 90%, with three MAbs (DENV1-E103, DENV1-E105, and DENV1-E106) showing 100% neutralization of all genotypes. Several MAbs showed reduced neutralizing potential for individual genotypes even at the relatively high (∼10 µg/ml) concentration of MAb used in the assay. For example, 9, 12, 12, and 14 MAbs neutralized infection by DENV genotypes 1, 3, 4, and 5, respectively. This was especially surprising for the genotype 1 strain as no appreciable reduction in binding to infected cells was observed (Table 2). As an example, DENV1-E104 showed relatively normal binding to the genotype 1 strain yet showed a 92% loss in neutralization activity. Some MAbs (DENV1-E100 and DENV1-E111), which had decreased binding to genotype 3 infected cells, also showed reduced inhibitory activity. Others (e.g., DENV1-E104) sustained normal inhibitory activity despite attenuated binding to fixed, permeabilized cells. Although more studies are necessary, this could reflect an epitope that is more sensitive to fixation in the context of recognition of specific genotypes.
Tab. 3. MAb neutralization of different DENV-1 genotypes. 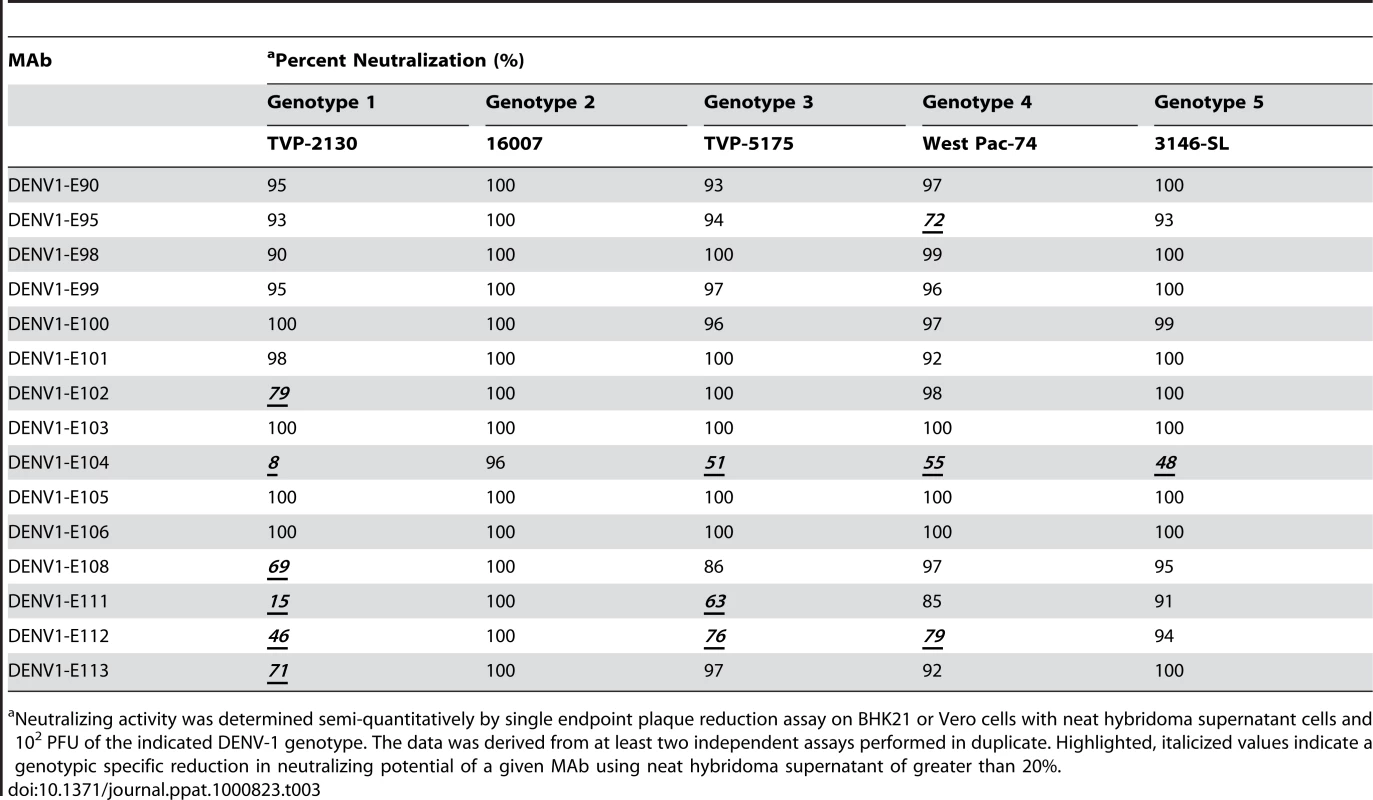
Neutralizing activity was determined semi-quantitatively by single endpoint plaque reduction assay on BHK21 or Vero cells with neat hybridoma supernatant cells and 102 PFU of the indicated DENV-1 genotype. The data was derived from at least two independent assays performed in duplicate. Highlighted, italicized values indicate a genotypic specific reduction in neutralizing potential of a given MAb using neat hybridoma supernatant of greater than 20%. To more rigorously characterize the neutralizing potency, we purified MAbs and assessed their inhibitory activity against the homologous genotype 2 (16007) strain in cell culture by performing a dose-response curve and determining the concentration of MAb (PRNT50, expressed here as ng/ml of antibody) that blocked plaque formation by 50% (Table 4). For the homologous genotype 2 16007 strain, 8 of 15 MAbs potently neutralized infection with PRNT50 values below 5 ng/ml. This value is significant as it is lower than that observed with our most inhibitory anti-WNV MAb (E16), which functions successfully as a post-exposure therapeutic agent [29],[49],[50]. Six other MAbs also showed strong inhibitory activity with PRNT50 values against 16007 of between 10 and 70 ng/ml. Only one MAb, E104 showed modest activity with a PRNT50 value of ∼600 ng/ml.
Tab. 4. PRNT50 values of MAbs against a homologous and heterologous DENV-1 strain. 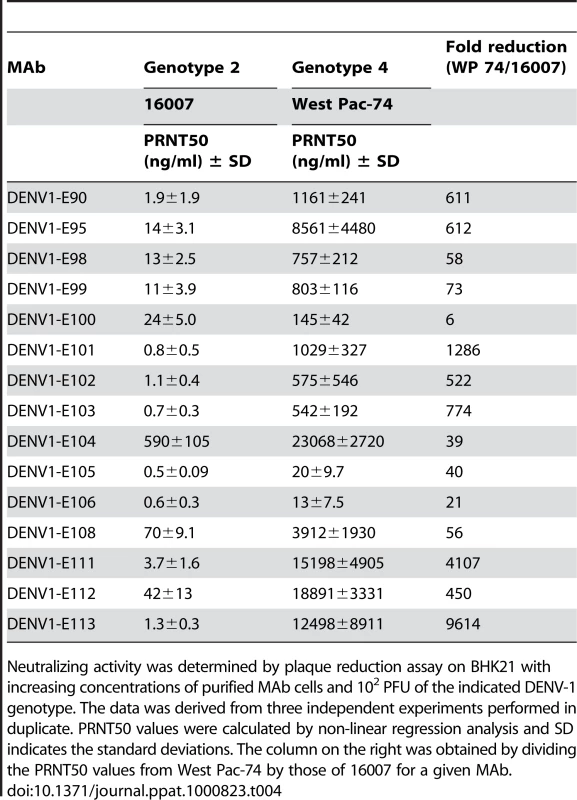
Neutralizing activity was determined by plaque reduction assay on BHK21 with increasing concentrations of purified MAb cells and 102 PFU of the indicated DENV-1 genotype. The data was derived from three independent experiments performed in duplicate. PRNT50 values were calculated by non-linear regression analysis and SD indicates the standard deviations. The column on the right was obtained by dividing the PRNT50 values from West Pac-74 by those of 16007 for a given MAb. Given our findings of genotype variation with respect to single endpoint neutralization using hybridoma supernatants, we repeated the dose-response curve analysis with a heterologous genotype 4 (West Pac-74) (Table 4). Surprisingly, the neutralizing activity was markedly decreased for all MAbs against the heterologous DENV-1 genotype (e.g., 6 to 9600-fold for DENV1-E100 and DENV1-E113, respectively). Only 2 MAbs, DENV1-E105 and DENV1-E106, retained strong neutralizing activity (≤20 ng/ml) against the West Pac-74 strain. Seven of 15 MAbs neutralized West-Pac 74 poorly with PRNT50 values greater than 1 µg/ml of antibody. Thus, the neutralizing efficiency of several DIII-specific anti-DENV-1 MAbs was significantly altered against strains of a different DENV-1 genotype.
Crystal structure of DENV-1 DIII and analysis of genotypic variation
To begin to understand the disparity in inhibitory potential of the DIII-specific neutralizing MAbs among different genotypes, we determined the X-ray crystal structure of recombinant DIII of the 16007 (genotype 2) strain at 2.25-Å resolution (Fig 2 and Table S2). Among the DENV-1 strains that represent the five different genotypes used in this study, variation in DIII was limited to nine residues, six of which were conservative substitutions (Fig 2 and Fig 3B). Four of the conservative substitutions (V/I380, L/V351, T/S339, and D/N341) are predicted to be partially or completely solvent inaccessible in the isolated DENV-1 DIII crystal structure and cryoelectron microscopy reconstruction of the mature virion [51], and thus, should not directly affect antibody engagement. In contrast, the conservative substitution K/R361 should be accessible due to its location in the DE loop. S/T397, the remaining conservative substitution is located after the G-strand in a region that links DIII with the first α-helix of the stem anchor motif. Of the three non-conservative substitutions, A/T369 is completely buried and although A/I/V345 is largely solvent accessible in the DIII crystal structure, it is likely buried in the virion according to cryoelectron microscopy reconstructions. The remaining non-conservative substitution (M/V297) occurs in the N-terminal linker region that connects to DI, which is solvent accessible in the DIII crystal structure, but less so in the mature virion structure. In summary, amino acid variation in DIII among the five different DENV-1 genotypes is modest, with the majority of changes being solvent inaccessible. Indeed, none of the observed genotypic variation occurs in residues previously implicated as critical to the creation of neutralizing antibody epitopes in DENV or related flaviviruses.
Fig. 2. Crystal structure of DENV-1 DIII (strain 16007) and analysis of genotypic variation. 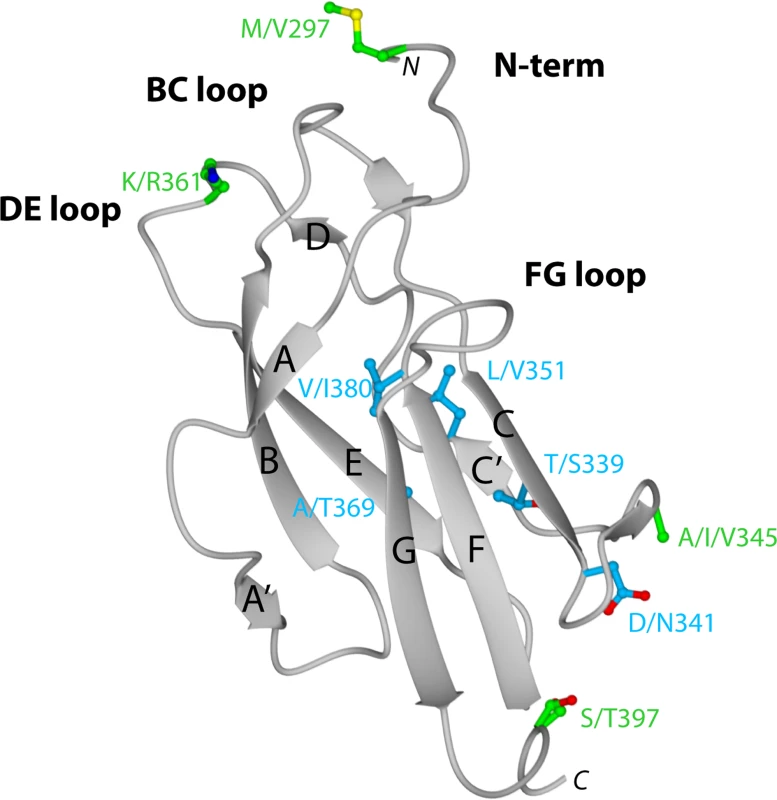
Ribbon diagram of Ig-like fold of DENV-1 DIII with β-strands and loops labeled accordingly after solution of the X-ray crystallographic structure at 2.25 Å resolution. Amino acid sequences in DIII that vary among the five different DENV-1 genotypes are indicated, with solvent inaccessible and accessible residues depicted in blue and green, respectively. Fig. 3. Epitope localization of anti-DENV-1 MAbs. 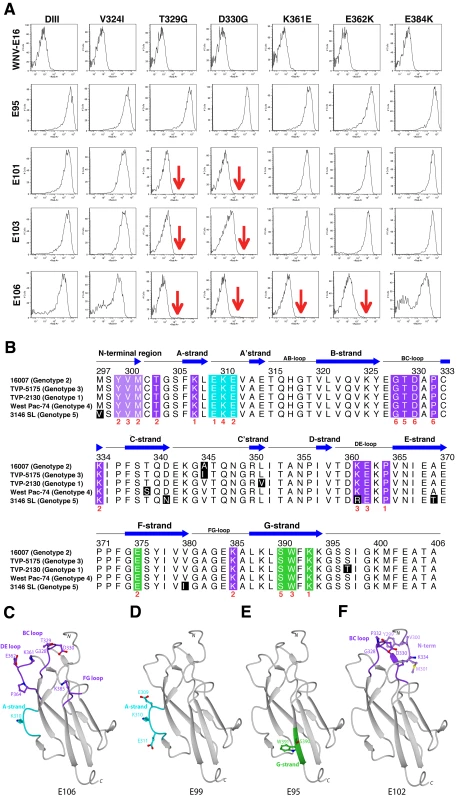
A. Flow cytometry histograms of loss-of-function DIII variants (V324I, T329G, D330G, K361E, E362K, and E384K) with individual neutralizing MAbs. Representative histograms are shown for the MAbs WNV E16 (negative control), DENV1-E95, DENV1-101, DENV1-103, and DENV1-106 with the wild type DIII and each of the variants. Data shown are representative of three independent experiments. B. Sequence alignment of different DENV-1 genotypes and mapping of neutralizing MAbs. The sequence and secondary structure of DIII from DENV-1 (strain 16007, genotype 2) E protein is aligned with other DENV-1 genotypes (genotype 1, strain TVP-2130; genotype 3, strain TVP 5175; genotype 4, strain Western Pacific-74; genotype 5, strain 3146 Sri Lanka). The secondary structure of DENV-1 E DIII residues 294 to 395 from the strains that had not been crystallized was predicted by DSSP [112] using the 16007 strain coordinates. Black blocks highlight residues of genotypic variation. The results of the yeast surface-display epitope mapping (see Table 5) are denoted underneath in red to indicate the number of neutralizing MAbs in our panel that lose binding when a specific amino acid is changed. Colored boxes correspond to specific neutralizing antibody and structural recognition determinants: N-terminal region, light purple; lateral ridge (BC, DE, and FG loops), dark purple; A-strand, cyan; and F- and G-strand, green. C–F. Localization of neutralizing epitopes on DENV-1 DIII as determined by yeast surface display. Structure of DENV-1 DIII (strain 16007) with amino acid residues marked in blue that significantly affect binding of neutralizing MAbs (C) DENV1-E106, (D) DENV1-E99, (E) DENV1-E95, and (F) DENV1-E102. Epitope mapping of DIII neutralizing MAbs
On the basis of studies that mapped DIII-specific MAbs against WNV and DENV-2 [25]–[27],[29],[41], we initially used reverse genetics to engineer corresponding mutations (S305P, K307E, E309K, K310E, V324I, T329G, D330G, K361E, E362K, E384K, K385E, and K393E) on residues of the BC, DE, and FG loops and the A β-strand (A-strand) of DIII of the genotype 2 (16007) strain, and displayed these variants on the surface of yeast. All DIII-specific MAbs were screened for loss-of-binding to the mutants to identify potentially critical recognition residues. Using this strategy, we localized binding of seven MAbs (DENV1-E100, DENV1-E101, DENV1-E102, DENV1-E103, DENV1-E104, DENV1-E105, and DENV1-E106) to individual or combinations of amino acids on the BC, DE, and FG loops of the lateral ridge of DIII (Table 5, Fig 3A, B, and C). Analogously, five MAbs (DENV1-E99, DENV1-E100, DENV1-E104, DENV1-E106, and DENV1-E113) exhibited reduced binding after mutation of A-strand (K307, E309, K310, and/or E311) residues (Fig 3D). Three of the neutralizing MAbs (DENV1-E95, DENV1-E111, and DENV1-E112), however, showed no appreciable reduction in binding with analogous mutations in the lateral ridge or A-strand residues that were identified from DENV-2 yeast screens.
Tab. 5. Summary of MAb binding to DENV-1 DIII mutants expressed on the surface of yeast. 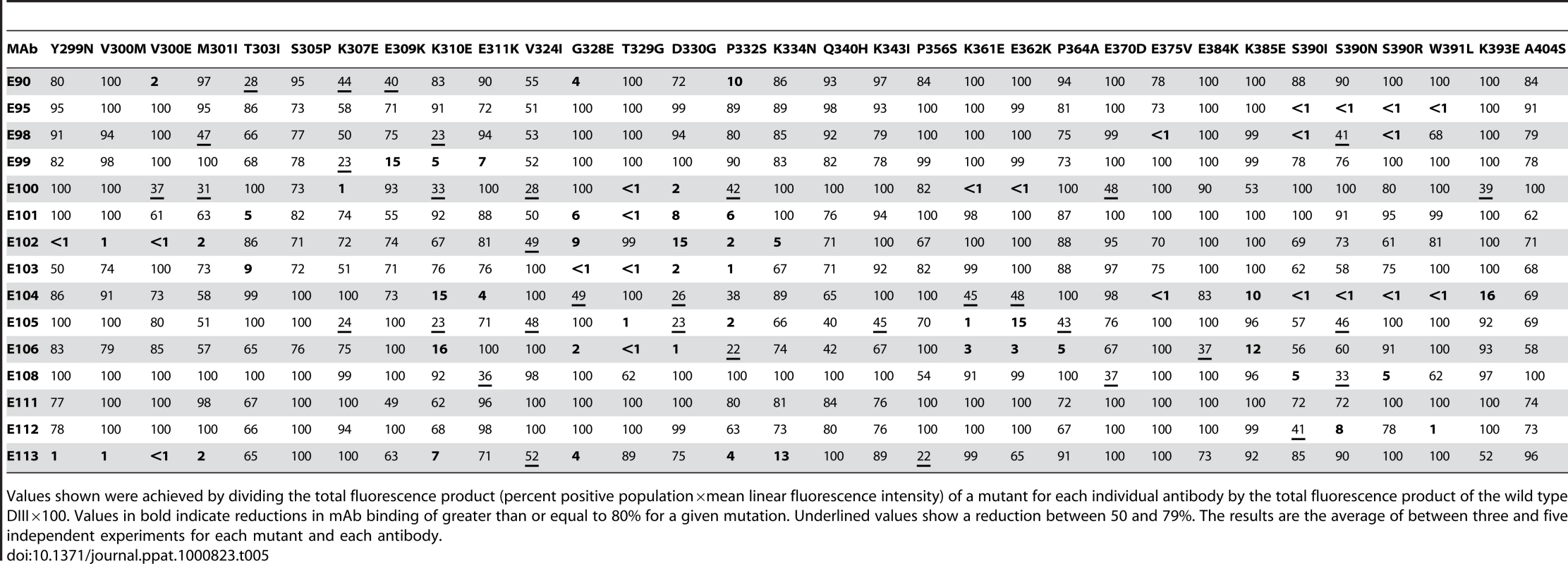
Values shown were achieved by dividing the total fluorescence product (percent positive population×mean linear fluorescence intensity) of a mutant for each individual antibody by the total fluorescence product of the wild type DIII×100. Values in bold indicate reductions in mAb binding of greater than or equal to 80% for a given mutation. Underlined values show a reduction between 50 and 79%. The results are the average of between three and five independent experiments for each mutant and each antibody. Forward genetic screens subsequently were performed to define additional residues that affected MAb binding. Error-prone PCR introduced random point mutations within DIII of DENV-1 E protein. Yeast were incubated sequentially with an Alexa Fluor 647-conjugated individual MAb and an Alexa Fluor 488-conjugated oligoclonal pool of MAbs to eliminate mutants that abolished surface expression of DIII (see Materials and Methods). Yeast that selectively lost expression of individual MAb epitopes were identified, subjected to plasmid recovery, sequenced, and tested for reactivity by flow cytometry against the remainder of the MAb panel (Table 5). In total, 21 DIII residues were identified that when changed resulted in a decrease in yeast staining by one or more MAbs, with mutations in BC-loop residues most frequently observed (Fig 3B). Only one of these residues (K/R361) is variable amongst the five genotypes, an unexpected observation based on the distinct genotypic specificities of the different MAbs.
The two most protective MAbs in vivo (see below), DENV1-E105 (type-specific) and DENV1-E106 (sub-complex specific), lost binding when residues on the BC (G328, T329 and D330), DE (K361E and E362K), and FG (K385) loops of the DIII-LR region were altered. Similarly, other protective MAbs (DENV1-E100 and DENV1-E103) also lost binding when residues in the BC or DE loop were altered. Type-specific MAbs (DENV1-E95, DENV1-E101, DENV1-E104, DENV1-E108, DENV1-E111, and DENV1-E112) that inhibited genotype 2 (16007) yet poorly neutralized the genotype 4 (West Pac-74) strain mapped to additional sites in DIII. DENV1-E95, DENV1-E104, DENV1-E108, and DENV1-E112 exhibited reduced binding with mutations in the G-strand (S390, W391, and K393) (Fig 3E) whereas DENV1-E101 was affected by changes in the N-terminal linker (T303) and BC-loop (G328, T329, and D330). Surprisingly, none of the mutations altered DENV-E111 binding, and forward genetic screens with this MAb also failed to identify a loss-of-binding variant (data not shown). DENV1-E104, which showed the weakest neutralizing activity, localized to residues in the A (K310 and E311), F (E375), and G (S390 and W391) β-strands.
Because of analogous studies with DENV-2, we anticipated that sub-complex-specific and cross-reactive MAbs would recognize the A-strand epitope [25]–[27]. Indeed, mutation of S305, K307, E309, K310, and E311 reduced binding of several sub-complex MAbs (DENV1-E98, DENV1-E99, DENV1-E106, and DENV1-E113) in our panel. For the cross-reactive neutralizing MAb DENV1-E102, its epitope localized to residues in the N-terminal linker region (Y299, V300, and M301) and the BC loop (D330 and P332) (Fig 3F). Although Y299, M301, and P332 are highly conserved among strains of different DENV serotypes, V300 is specific for DENV-1 and D330 is present in DENV-1 and DENV-3; this may explain why DENV1-E102 fails to neutralize DENV-2 and DENV-3 efficiently (S. Sukupolvi-Petty, J. Brien, and M. Diamond, unpublished results).
Despite the extensive epitope mapping data (summarized in Fig 3B and 4), no structural explanation was readily apparent for the decrease in neutralization of the genotype 4 West Pac-74 strain by several antibodies. MAbs that showed markedly depressed neutralization of West Pac-74 localized to residues that were instead conserved among DENV-1 genotypes. To begin to address this, we evaluated whether individual MAbs differentially recognized DIII from 16007 or West Pac-74 strains when expressed on yeast; the DIII of these two strain differ by only two amino acids at positions 339 and 345 in the C strand and C-C′ loop (Fig 3B). Notably, all neutralizing MAbs equivalently recognized yeast displaying DIII of 16007 or West Pac-74 over a range of concentrations (Fig 5A, and data not shown). Thus, sequences differences in DIII of West Pac-74 and 16007 did not explain the differential patterns of neutralization. Because of this, we hypothesized that sequences variation in other domains of the West Pac-74 E protein could directly alter MAb binding or perhaps influence the display of DIII on the infectious virion. To test this, we performed a virion capture ELISA with 16007 and West Pac-74 viruses and selected MAbs having different neutralization profiles against the two strains (Fig 5B). DENV1-E98, which exhibited a ∼50-fold reduction in neutralization showed little difference in binding over several concentrations of antibody. In contrast, DENV1-E111, which had ∼4,000-fold less neutralizing activity, exhibited a depressed ability to bind West Pac-74 compared to 16007. Surprisingly, DENV1-E90 and DENV1-E113, which had ∼600 to 9,600-fold differences in PRNT50 values with West Pac-74 compared to 16007 showed virtually no variation in binding in the capture ELISA. Thus, differential neutralization of genotypes 2 and 4 of DENV-1 could be explained by disparate binding to the virion for only a subset of MAbs. For MAbs like DENV1-E90 and DENV1-E113, the profile was similar to that observed with escape variants of a human anti-WNV MAb; in that case, mutations that abolished neutralizing activity did not appear to affect antibody binding to virus [48].
Fig. 4. Summary of neutralizing antibody and structural recognition determinants on DENV-1 DIII. 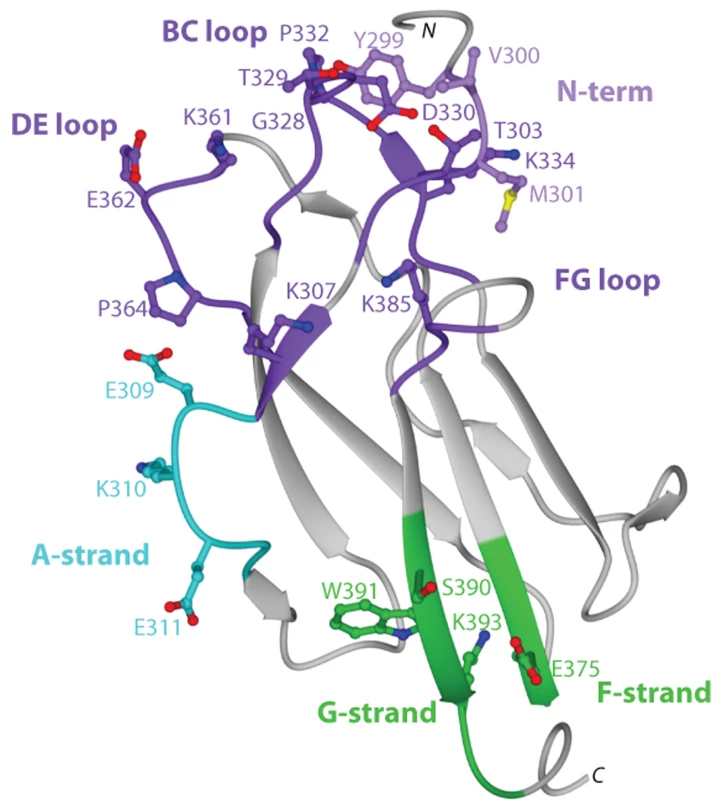
Data from yeast surface display (Table 5) and sequence alignment (Fig 3B) were mapped onto the crystal structure of DENV-1 DIII to visualize the composite epitopes of different neutralizing MAbs. Recognition sites include N-terminal region, light purple; lateral ridge (BC, DE, and FG loops), dark purple; A-strand, cyan; and F- and G-strand, green. Fig. 5. Binding of MAbs to DIII or virions of 16007 and West Pac-74. 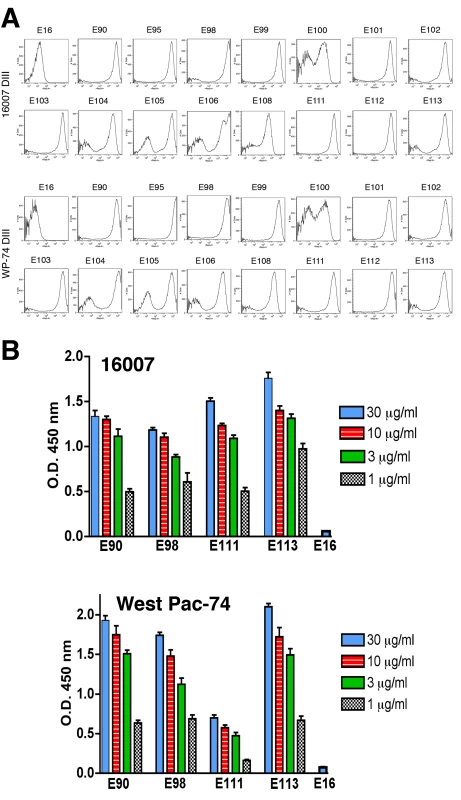
A. MAb binding to DIII of 16007 or West Pac-74 on yeast. Flow cytometry histograms of DIII of 16007 or West Pac-74 (different at residues 339 and 345) with neutralizing MAbs. Data shown are representative of two independent experiments. B. A capture ELISA was used to detect binding of MAbs to 16007 and West Pac-74 virions. Microtiter plates were coated with the indicated MAbs, incubated with 2×105 PFU of 16007 or West Pac-74 at room temperature, and with biotinylated anti-E MAbs. The data is an average of several separate experiments performed in duplicate, and error bars indicate standard deviation. In vivo protection studies with neutralizing MAbs
Pre-exposure passive transfer of neutralizing MAbs against WNV, DENV-2, and DENV-4 protects against lethal infection in wild type and immunodeficient mice [24], [29], [52]–[56]. To confirm that neutralizing anti-DENV-1 MAbs protect in vivo and begin to explore the possibility for antibody therapy, we evaluated the efficacy of our panel of inhibitory MAbs against West Pac-74, the heterologous genotype 4 DENV-1 strain. The heterologous strain was selected for two reasons: (i) the West Pac-74 strain was the only DENV-1 isolate in our collection that caused 100% lethality in an intraperitoneal challenge model in AG129 mice; and (ii) a possible antibody therapeutic must demonstrate efficacy against DENV-1 strains of different genotypes.
A single dose (500 µg) of neutralizing MAbs against DENV-1 was administered as prophylaxis one day prior to infection with 106 PFU of West Pac-74. Two MAbs (DENV1-E105 and DENV1-E106) protected 100% of mice compared to PBS or irrelevant MAb controls (P<0.0001), which had a 0% survival rate (Table 6). Three other MAbs (DENV1-E99, DENV1-E100, and DENV1-E103) exhibited strong yet incomplete protection with 60 to 80% survival rates. However, nine neutralizing MAbs (DENV1-E90, DENV1-E95, DENV1-E98, DENV1-E101, DENV1-E102, DENV1-E108, DENV1-E111, DENV1-E112 and DENV1-E113) protected inefficiently, with an 11 to 38% survival rate. Although the survival rate was relatively low for these MAbs, the mean time to death (MTD) was protracted (23±6.1 to 40±8.6 days) compared to untreated, infected mice (20±3.3 days). One neutralizing MAb, DENV1-E104 showed no protection against lethality although a longer MTD was observed. In general, the protective efficacy in vivo correlated with neutralizing activity against the West Pac-74 genotype 4 strain. As an example, DENV1-E105 and DENV1-E106 had low PRNT50 values (13 to 20 ng/ml) for West Pac-74 and protected against infection whereas DENV1-E104 and DENV1-E112, which protected poorly, had PRNT50 values (18 to 23 µg/ml) that were almost 1,000-fold different.
Tab. 6. In vivo protection of AG129 mice from West Pac-74 (genotype 4) after prophylaxis with neutralizing anti-DENV-1 MAbs. 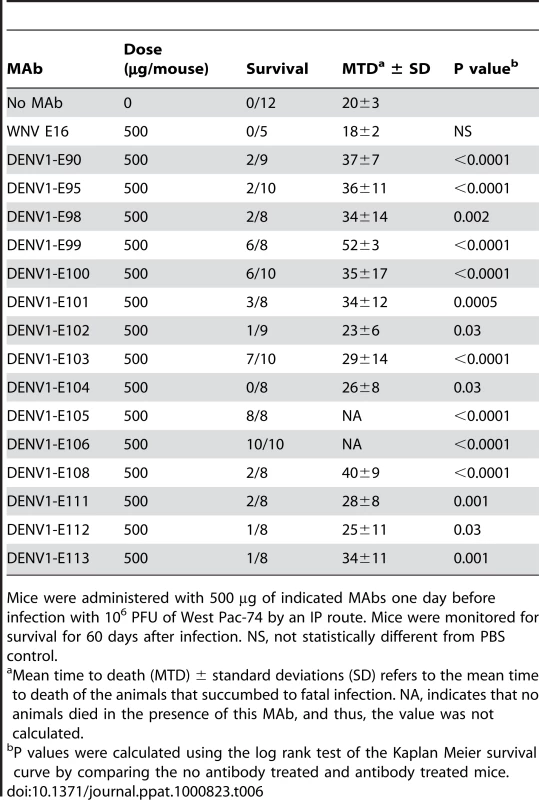
Mice were administered with 500 µg of indicated MAbs one day before infection with 106 PFU of West Pac-74 by an IP route. Mice were monitored for survival for 60 days after infection. NS, not statistically different from PBS control. To begin to define the therapeutic potential of the highly protective MAbs (DENV1-E99, DENV1-E103, DENV1-E105, and DENV1-E106), we administered 5 and 25-fold lower doses (100 µg and 20 µg) one day prior to IP infection of AG129 mice with 106 PFU of West Pac-74. The lower doses of DENV1-E99 and DENV1-E103 provided sub-optimal protection against lethal infection (Fig 6A and B), with only a delay in the MTD observed. In comparison, ∼50% of mice survived West Pac-74 infection after receiving a single dose of 20 or 100 µg of DENV1-E105 antibody (Fig 6C, P≤0.003). More strikingly, a single 20 or 100 µg injection of DENV1-E106 completely protected DENV-1 infected mice (Fig. 6D, P<0.0001). These data suggest that DENV1-E106 was the most protective MAb in our panel of neutralizing antibodies against infection by the heterologous strain, DENV-1 West Pac-74.
Fig. 6. Dose response of protective efficacy MAbs in mice infected with DENV-1. 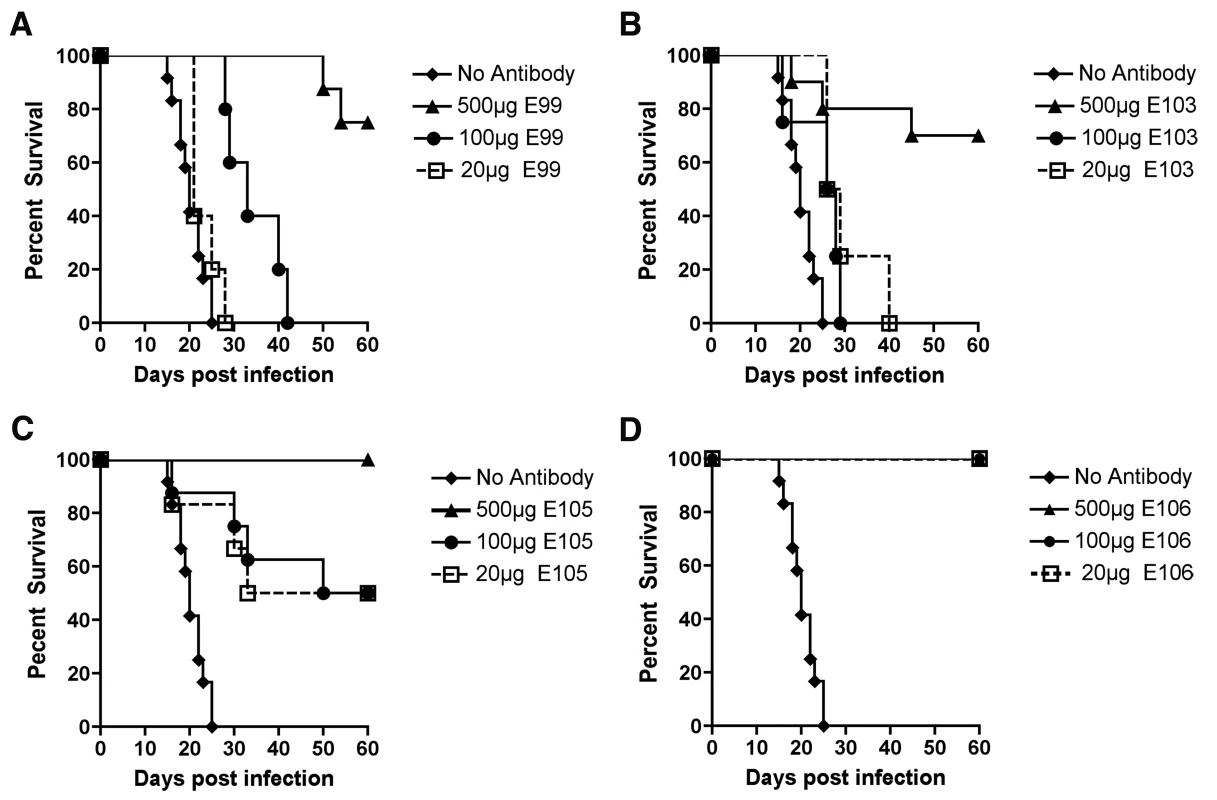
AG129 mice were passively transferred saline or 20, 100, or 500 µg of (A) DENV1-E99, (B) DENV1-E103, (C) DENV1-E105, and (D) DENV1-E106 MAbs one day before infection with 106 PFU of West Pac-74 (genotype 4) by an IP route. Mice were monitored for survival for 60 days after infection. The survival curves were constructed from data of two independent experiments. The number of animals for each antibody dose ranged from 5 to 10 per group. Therapeutic studies with neutralizing MAbs
Antibody-based therapeutics against DENV will need to be administered after infection. To assess the therapeutic efficacy of strongly neutralizing anti-DENV-1 MAbs, mice were infected with 106 PFU of West Pac-74 and two days after infection, a single 500 µg MAb dose (DENV1-E99, DENV1-E103, DENV1-E105, or DENV1-E106) was transferred passively by an IP route. Notably, 500 µg of DENV1-E99 or DENV1-E103 provided no protection with animals dying at a rate and time similar to untreated mice (P>0.2). In contrast, DENV1-E105 and DENV1-E106 protected 75 and 82% (P≤0.0001) of mice, respectively, when administered two days after infection. When a single dose of DENV1-E105 or DENV1-E106 MAb was given four days after infection, 20 and 40% (P≤0.03) of mice, respectively were protected from lethal infection (Fig 7A and B).
Fig. 7. Therapeutic efficacy of strongly neutralizing antibodies in mice after DENV-1 infection. 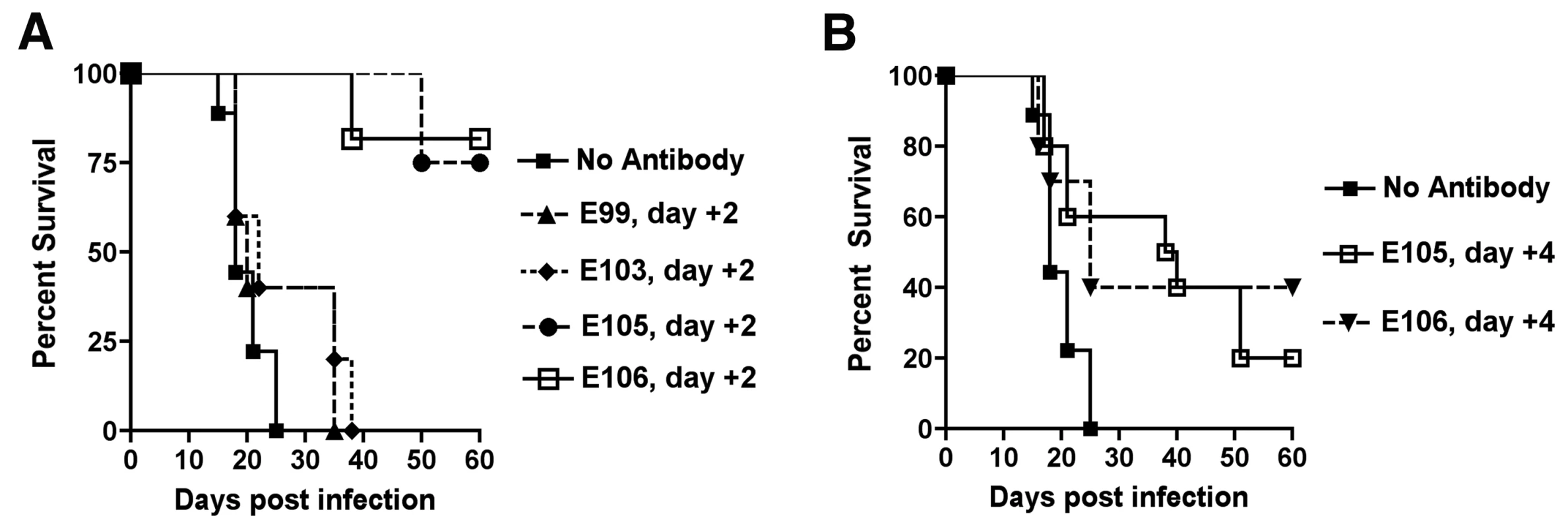
Mice were administered saline or a single 500 µg dose of DENV1-E105 or DENV1-E106 MAbs at day (A) 2 or (B) 4 after infection with 106 PFU of West Pac-74 (genotype 4) by an IP route. Mice were monitored for survival for 60 days after infection. The number of animals for each antibody ranged from 10 to 11 per group. Given the potency of DENV1-E105 and DENV1-E106 in protection against the genotype 4 West Pac-74 strain, we performed PRNT50 analysis with strains corresponding to the remaining genotypes to assess potential clinical utility of these MAbs. Both DENV1-E105 and DENV1-E106 strongly neutralized infection of all DENV-1 genotypes (Fig 8A and B), with PRNT50 values ranging from 0.5 to 59.2 ng/ml. Thus, DENV1-E105 and DENV1-E106 protected AG129 immunocompromised mice as a pre and post-exposure treatment from infection by a heterologous DENV-1 genotype and efficiently inhibited all DENV-1 genotypes in cell culture.
Fig. 8. PRNT50 analysis of different DENV-1 genotypes by DENV1-E105 and DENV1-E106. 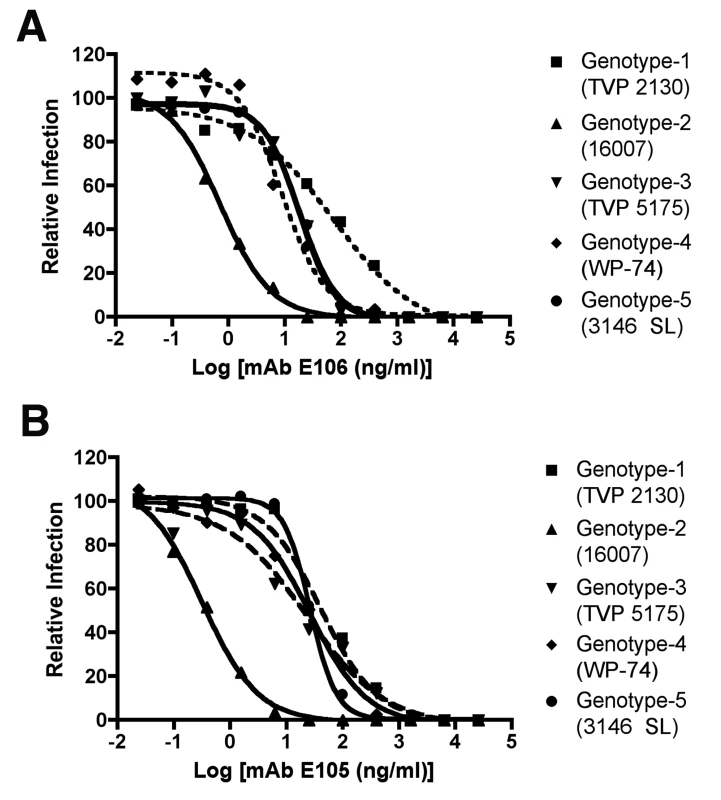
Increasing concentrations of purified (A) DENV1-E106 and (B) DENV1-E105 were mixed with 102 PFU of DENV-1 strains corresponding to all five genotypes and neutralization was assessed by a standard PRNT assay in BHK or Vero cells (depending on the strain). PRNT50 values were calculated after regression analysis using statistical software. The graph is representatives of at least two independent experiments performed in duplicate. Discussion
One primary goal of this study was to generate a collection of strongly neutralizing MAbs that would recognize virtually all DENV-1 strains and protect against infection as post-exposure therapy. In an attempt to achieve this, we generated a panel of 79 new MAbs against DENV-1 E protein. Although neutralizing antibodies can be generated against all three domains of the flavivirus E protein [29],[32],[42],[46],[52],[55],[57],[58], we mapped the epitopes of the MAbs with the greatest inhibitory activity primarily to epitopes on DIII, analogous to that observed with MAbs against DENV-2 [25]–[27],[41]. One surprising finding that has not been reported previously was the disparate neutralizing activity and protective potential of individual MAbs against different DENV-1 genotypes. In comparative studies with strains from genotype 2 and genotype 4, virtually all MAbs showed reduced (from 6 to 9,600-fold) PRNT50 values against the heterologous genotype. Because of this, few (5 of 15) MAbs protected as pre-exposure prophylaxis against infection by the heterologous DENV-1 genotype in AG129 mice, and only two (DENV1-E105 and DENV1-E106) showed utility as post-exposure therapy.
Although many DENV strains can cause a similar debilitating febrile illness in humans, there is significant genetic variation (25 to 40% amino acid difference for serotypes and up to 3% variation among genotypes within a serotype) and phenotypic difference in virulence among individual strains of specific serotypes and genotypes [59]–[63]. Previous studies identified the lateral ridge and A-strand epitopes in DIII as targets of strongly neutralizing type-specific and sub-complex-specific neutralizing MAbs against DENV-2 and DENV-3 [26], [27], [41], [64]–[67] and the fusion loop in DII as a recognition site for many cross-reactive neutralizing DENV MAbs [58],[68],[69]. In comparison, the serotype-specific 5H2 neutralizing MAb against DENV-4 localized to the loop between the G and H β-strand in DI [42]. Prior to our study, little was known as to how neutralizing antibodies recognized DENV-1. Although neutralizing anti-DENV-1 MAbs have been generated [44],[45], few were localized to specific amino acids or structural domains. Two neutralizing IgM against DENV-1 were mapped by neutralization escape selection to amino acids E279 and E293, at the hinges between DI and DII or DI and DIII, respectively [46]. While our experiments establish that highly neutralizing antibodies recognize determinants in DIII of DENV-1, we do not rule out the existence of other epitopes in distinct regions. Rather, the panel of MAbs that we obtained may be skewed by the immunization protocol (boosting with recombinant DIII), primary screen, and genetic background of the animals, which influences the B cell repertoire. Indeed, for WNV we have recently identified strongly neutralizing and fusion-blocking human MAbs that poorly recognize recombinant forms of the E protein and map to the DI-DII hinge and dimer contact regions [48].
This is the first study that describes a post-exposure therapeutic effect of a MAb against any DENV strain in any animal model. In particular, DENV1-E105 and DENV1-E106, which neutralized all five DENV-1 genotypes showed efficacy when administered two or four days after infection of highly immunocompromised AG129 mice. In contrast, three other neutralizing MAbs (DENV1-E99, DENV1-E100, and DENV1-103) failed to protect when added a few days after infection even though they functioned as pre-exposure prophylaxis. Although prior studies demonstrated that anti-E MAbs protect against DENV infection in rodents and non-human primates [42],[55],[56],[70],[71], none have reported efficacy when added after infection. It is noteworthy that the two MAbs with the most neutralizing PRNT50 values were the ones with greatest therapeutic efficacy, suggesting, at least for strongly neutralizing MAbs of a given class, a correlation between in vitro and in vivo activity. It is unclear why therapeutic anti-DENV MAbs have not been previously described. Several post-exposure therapeutic MAbs or polyclonal antibody preparations against WNV, JEV, YFV, and tick-borne encephalitis virus have been reported [29], [30], [53], [57], [72]–[77], with some entering human clinical trials [43]. One speculation is that because viremia associated with DENV infection is several log greater than WNV [78]–[80], a condition of viral antigen excess occurs more rapidly in circulation leading to fewer antibody molecules binding to a given DENV virion. As neutralization is determined in part by the stoichiometry of antibody binding [81],[82], only MAbs that can block infection at low fractional occupancy may be capable of neutralizing in vivo.
MAb-based therapeutics against DENV could be complicated by antibody-dependent enhancement of infection (ADE). Sub-neutralizing concentrations of antibody augment infection of Fc-γ receptor-expressing cells possibly by enhancing the efficiency of virus attachment or entry [83],[84]. Thus, the administration of virus-specific MAbs could adversely impact the outcome of DENV infection. In non-human primates, passive transfer of sub-neutralizing concentrations of monoclonal or polyclonal antibody increased viremia although no change in disease status was observed [85],[86]. Based on the possibility of ADE, MAb therapeutics against DENV in humans would appear to have considerable regulatory hurdles. However, recent studies with amino acid substitutions or deletions in the Fc region of recombinant anti-flavivirus antibodies prevented ADE in vitro and in vivo [86]–[88]. As such, studies are underway to engineer mutations into the Fc region of a humanized version of DENV1-E106 that abolishes the possibility of enhancement by abrogating binding to Fc-γ receptors.
Amino acid contact residues of neutralizing MAbs that react with DENV-2 or DENV-3 have been defined by analyzing neutralization escape mutants [40], chimeric DENV variants [89], site-specific DENV-2 mutants [68],[90], peptide mapping [65],[91],[92], yeast surface display of variant E proteins [25],[26],[66], mutant recombinant proteins [27],[41],[67], and by co-crystallography of recombinant DENV proteins and Fab fragments of neutralizing MAbs [25]. Here, we used genetic strategies to generate mutant DIII of DENV-1 on yeast to map antibody recognition residues. The validity of the yeast surface display mapping for identifying critical contact residues has been confirmed by crystallographic studies that resolved the structural interface between WNV and DENV2 DIII Fab fragments [25],[93] as well as DII-fusion loop specific Fab recognition [94]. For neutralizing type-specific DENV-2 MAbs that mapped to specific amino acid residues in DIII, the majority localized to residues in the BC and FG loops within the lateral ridge [26],[41]. Our yeast mapping experiments confirm that the lateral ridge and A-strand on DIII are also recognized by strongly neutralizing anti-DENV-1 antibodies.
The detailed mapping of neutralizing anti-DENV-1 MAbs identified two additional regions on DIII that comprised protective epitope(s): (a) DENV1-E102 and DENV1-E113, which react with multiple DENV serotypes, showed an almost complete loss-of-binding with mutation of residues (Y299, V300, and M301) in the N-terminal linker region. This result is analogous to that described for neutralizing DENV-2 MAbs where the mutation M301G abolished reactivity of five complex-specific and seven serotype-specific MAbs [27],[41]. The binding of the N-terminal linker by MAbs against WNV has been speculated to contribute to neutralization by inhibiting the significant rotation of DIII during the dimer to trimer transition [93]. (b) DENV1-E95, DENV1-E98, DENV1-E104, DENV1-E108, and DENV1-E112 exhibited loss of binding when residues (S390, W391, and K393) in the G-strand were changed. As the G-strand of the DIII is solvent accessible on the surface of the DENV virion [7], it is structurally feasible that recognition of this region by MAbs will inhibit infection. Indeed, in experiments that assessed the reactivity of a panel of complex-specific neutralizing MAbs against DENV-2 and DENV-3 DIII, mutation of G-strand residues also reduced binding [27],[67]. Additionally, X-ray crystallography studies identified residues K388 and N390 in the G-strand of DENV-2 as contacts for the neutralizing sub-complex-specific MAb 1A1D-2 [25]. These studies suggest that DIII is a complex antigen that is recognized by neutralizing MAbs that localize to one of four overlapping regions: the N-terminal linker, A-strand, lateral ridge loops, and G-strand. What remains uncertain is whether binding to specific determinants on DIII influences the mechanism of antibody neutralization of DENV.
Although our most strongly neutralizing MAbs localized to DIII of DENV-1, these antibodies were derived from immunodeficient mice, which may not necessarily reflect the neutralizing antibody response in humans. The antibody repertoire against DENV-1 or other serotypes in humans remains unknown, as B cell profiling at the epitope level has not been performed. Nonetheless, recent studies suggest that the humoral response against flaviviruses in humans may be directed away from DIII neutralizing epitopes [53],[54],[95]. DIII-specific antibodies comprised only a small fraction of the total antibody in DENV immune sera [96],[97]; whether these antibodies contribute to DENV neutralization in vivo remains controversial. Because highly neutralizing antibodies against DIII of DENV-1, DENV-2, and DENV-3 map to the lateral ridge and A-strand epitopes [26],[27],[41],[67], DIII-based vaccines [98]–[101] that skew the humoral response have the potential to elicit highly protective responses.
One intriguing finding was the disparate neutralizing activity of individual MAbs against different DENV-1 genotypes. Infection with one DENV serotype is believed to confer long-term immunity against strains of the homologous but not heterologous DENV serotypes. Based on this, we assumed it would be straightforward to generate DIII-specific therapeutic MAbs that neutralized all five genotypes within the DENV-1 serotype. Indeed, there are limited amino acid changes in DIII among DENV-1 genotypes with a maximum difference of seven amino acids, with genotype 5 the most diverse although the changes are largely conserved and reside in solvent inaccessible regions. Nonetheless, several MAbs in the panel exhibited markedly depressed neutralizing activity against the heterologous West Pac-74 strain. This was perplexing given that the 16007 and West Pac-74 strains differ in DIII by two amino acids, which were not coincident with our epitope mapping data: a T→S change at residue 339 in the C-strand and an A→V change at residue 345 in the C-C′ loop. Indeed, in comparative binding assays with 16007 and West Pac-74 DIII on yeast, no obvious difference in binding at saturating concentrations of MAb was observed. When studies were repeated with selected MAbs in a virus capture ELISA, one MAb (DENV1-E111) with disparate neutralizing potential showed reduced binding to West Pac-74, but others (DENV1-E90 and DENV1-E113) did not. This was corroborated in surface plasmon resonance studies, which showed an ∼10-fold decrease in the KD of binding of DENV1-E111 for West Pac-74 DIII (K. Austin, M. Diamond, D. Fremont, unpublished results). Although further structural studies are warranted, we propose four hypotheses to explain these results: (a) similar to findings with a recently characterized human anti-WNV MAb [48], mutations that abrogate neutralizing activity do not always reduce measurable antibody binding to the virion; (b) because of sequence variation outside of DIII, the E protein of different DENV-1 genotypes may pack differently on the virion. Some components of an epitope for a given MAb may be differentially exposed on virions of distinct genotype; (c) individual loss-of-function mutations identified by yeast display do not necessarily represent the complete footprint on DIII of bound antibody, as mutation of other residues may have allosteric effects; or (d) neutralizing anti-DENV-1 MAbs in our panel may have additional amino acid contacts outside of DIII that vary among the DENV-1 genotypes. Notably, we did not observe detectable binding of any the DIII-specific neutralizing MAbs to a DI-DII protein displayed on yeast (data not shown). Structural studies with Fab-E protein or Fab-virion complexes will be required to address these possibilities.
Regardless of the mechanism, our results suggest that antibodies against one DENV genotype may have decreased inhibitory potency against a heterologous genotype within the same serotype. This has potential implications for assessing the breadth of the protective efficacy of tetravalent DENV vaccines, which are in advanced clinical trials. In addition to evaluating protection against other serotypes, it may be critical to assess whether antibody responses against the vaccine strain of a given serotype neutralize infection of heterologous genotypes equivalently.
Materials and Methods
Cells and viruses
Vero, BHK21-15 and Raji-DC-SIGN-R cells were cultured in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) (Omega Scientific) and antibiotics (penicillin G and streptomycin) at 37°C in a 5% CO2 incubator. Strains from all five genotypes of DENV-1 included: TVP-2130 (genotype 1), 16007 (genotype 2), TVP-5175 (genotype 3), Western Pacific-74 (genotype 4) and 3146 Sri Lanka (genotype 5). TVP-2130 and TVP-5175 were obtained from Dr. Robert Tesh and the World Reference Center for Emerging Viruses and Arboviruses (University of Texas Medical Branch, Galveston, TX). Western Pacific-74 (West Pac-74) and the 3146 Sri Lanka strain were obtained from Drs. Stephen Whitehead (NIAID, NIH, Bethesda, MD) and Rebeca Rico-Hesse (Southwest Foundation for Biomedical Research, San Antonio, Texas), respectively. Strains corresponding to other DENV serotypes were also used: 16681 (DENV-2), 16652 (DENV-3), and H241 (DENV-4). All viruses were amplified in C6/36 Aedes albopictus cells according to previously described protocols [26].
Generation, purification, and labeling of anti-DENV-1 MAbs
MAbs were generated essentially as described [29] after performing several independent splenocyte-myeloma fusions. To generate anti-DENV-1 MAbs, IFN-αβR−/− C57BL/6 mice were infected with 105 PFU of DENV-1 strain 16007 (genotype 2) via an intraperitoneal route and re-challenged two weeks later with the same strain. Mice with serum having the highest neutralizing titer (>1/1500) were immunized with purified DIII (50 µg, strain 16007) in PBS as a final intravenous boost. Three days later splenocytes were fused to P3X63Ag8.53 myeloma cells [102]. MAbs were subcloned by limiting dilution, isotyped (Pierce Rapid ELISA Kit), and purified using protein A affinity chromatography (Invitrogen). For library sorting experiments, MAbs were labeled with Alexa Fluor 647 or Alexa Fluor 488, using a MAb labeling kit (Molecular Probes) according to the manufacturer's instructions.
In vitro neutralization assay
Plaque reduction neutralization titer (PRNT) assays were performed with the indicated DENV-1 strains with MAbs or serum on BHK21-15 cells as described previously [26],[80]. PRNT50 values were determined using non-linear regression analysis (Graph Pad Prism4).
Cloning and expression of DIII of DENV-1 E protein
A cDNA encoding DENV-1 DIII (strain 16007, residues 293 to 400) of the 16007 strain was amplified from viral RNA by reverse transcriptase and high-fidelity Platinum Taq polymerase chain reaction (PCR) according to the manufacturer's instructions (Invitrogen). The PCR product was cloned into the pET21a bacterial expression plasmid (EMD Biosciences) using flanking NdeI and XhoI restriction sites, sequenced, and then expressed in BL21 Codon Plus (Stratagene) E. coli. Inclusion bodies containing insoluble aggregates were denatured in the presence of 6 M guanidine hydrochloride and 20 mM β-mercaptoethanol and refolded in the presence of 400 mM L-arginine, 100 mM Tris-base (pH 8.0), 2 mM EDTA, 0.2 mM phenylmethylsulfonyl fluoride, and 5 and 0.5 mM reduced and oxidized glutathione, respectively. Refolded protein was separated from aggregates on a Superdex 75 or 200, 16/60 size exclusion column using fast-protein liquid chromatography (GE Healthcare).
X-ray crystallographic structure determination of DV1 DIII
DENV-1 DIII was crystallized by vapor diffusion in hanging drops at 20°C at a concentration of 14 mg/ml. The drop contained equal parts protein and well solution composed of 0.1 M imidazole-HCl at pH 7.5, 0.2 M lithium sulfate, and 30% (w/v) polyethylene glycol 3000. Crystals were cryoprotected after a soak in well solution containing 15% glycerol before rapid cooling in a nitrogen gas stream. Data was collected at ALS beamline 4.2.2 (Lawrence Berkeley Laboratories, Berkeley, CA) at 293° K and at a wavelength of 1.29 Å using a CCD detector. The data were processed and scaled in HKL-3000 [103]. Crystallographic phasing was obtained by molecular replacement using the program Phaser [104] and the corresponding DIII fragment from DV2 (PDB accession 1OAN) as a search model. The resulting atomic model was iteratively built in O [105] and refined in REFMAC [106]. The final model with one DENV1 DIII monomer per asymmetric unit has been refined to 2.25Å resolution and contains 103 amino acids (residues 296 to 399 of DIII) and 53 water molecules. The DENV1 DIII domain is similar in structure to that reported for the post-fusion DENV-1 E ectodomain trimer (3G7T) [11] with a root-mean-square difference of 1.6 Å between Cα atom positions for equivalent core residues 296 through 394 [107]. The coordinates and diffraction data have been deposited in the RCSB with accession number 3IRC.
Domain mapping by yeast surface display
The DNA fragments encoding amino acid residues 1 to 293 (DI-DII) and 294 to 409 (DIII) of DENV-1 E protein were amplified from the DENV-1 strain 16007 or West Pac-74 by RT-PCR with BamHI and XhoI sites added at the 5′ and 3′ end, respectively. The PCR products were cloned as downstream fusions to Aga2 and Xpress epitope tag genes in the yeast surface display vector pYD1 (Invitrogen), under the control of an upstream GAL1 promoter. These constructs were transformed into Saccharomyces cerevisiae strain EBY100 [108],[109] using S.c. EasyComp Transformation Kit (Invitrogen) to generate yeast that expressed DENV-1 DI-DII or DIII. Individual yeast colonies were grown to logarithmic phase at 30°C in tryptophan-free yeast media containing 2% glucose. Fusion protein expression was induced on the surface by growing yeast for additional 48 hrs in tryptophan-free media containing 2% galactose at 20°C. Yeast cells were washed with PBS supplemented with BSA (1 mg/ml) and incubated with 50 µl of neat culture supernatant or purified MAbs at a concentration of 25 µg/ml. After a 30 minute incubation on ice, yeast were washed in PBS with BSA and then incubated with a goat anti-mouse IgG secondary antibody conjugated to Alexa Fluor 647 (Molecular Probes). After fixation with 1% paraformaldehyde in PBS, yeast cells were analyzed on a FACSArray flow cytometer (Becton-Dickinson) using FloJo software.
Yeast library construction and screening
To generate a library of DENV-1 DIII variants on yeast, mutation was accomplished by error-prone PCR, using a GeneMorph II random mutagenesis kit (Stratagene). The library was ligated into the pYD1 vector and transformed into XL2-Blue ultracompetent cells (Stratagene). The colonies were pooled and transformed into yeast cells as described above. In some instances, additional site-specific mutations were engineered into DENV-1 DIII by a reverse genetic approach using the Quick Change II Mutagenesis Kit (Stratagene).
For individual MAbs, the DENV-1 DIII mutant library was screened according to a previously described protocol [29]. To identify yeast that had selectively lost binding to a given MAb, the library was stained with an individual Alexa Fluor 647 conjugated MAb for 30 min on ice. To control for surface expression of appropriately folded DIII of DENV-1, yeast cells were subsequently stained for 30 min on ice with an Alexa Fluor 488-conjugated pool of DIII-specific MAbs (DENV1-E90, DENV1-E95, DENV1-E99, DENV1 - DENV1-E111, DENV1-E112, and DENV1-E113), and then processed by flow cytometry, The population that was single MAb negative but pool oligoclonal MAb positive was selected and sorted. After four to five rounds of sorting, yeast were plated and individual colonies were tested for binding to individual MAbs by flow cytometry. For clones that had lost binding to the desired MAb of interest, the plasmid was recovered using a Zymoprep yeast miniprep kit (Zymo Research), transformed into XL1-Blue competent E. Coli (Stratagene), purified using a QIAprep spin miniprep kit (Qiagen), and sequenced.
Immunostaining of DENV-infected cells
For the study of binding of DENV-1 MAbs to different DENV strains, Raji-DC-SIGN-R or C6/36 cells were infected with different DENV strains at a multiplicity of infection (MOI) of 1. Depending on the DENV-1 strain, Raji-DC-SIGN-R cells were harvested at 48 or 96 hours and C6/36 cells were harvested at day 7 after infection. Cells were washed, fixed in PBS with 4% paraformaldehyde, and permeabilized in Hanks' balanced salt solution (HBSS) supplemented with 10 mM HEPES (pH 7.3), 0.1% (w/v) saponin (Sigma), and 0.02% NaN3. The cells were then incubated with MAbs for 30 minutes on ice, washed and incubated subsequently with an Alexa Fluor 647-conjugated goat anti-mouse IgG (Molecular Probes). After 30 minutes, cells were washed and fixed in PBS with 1% paraformaldehyde and analyzed by flow cytometry. To test whether DENV-1 MAbs cross-reacted with more distantly related flaviviruses, infection and staining experiments were performed with WNV (strain 3000.0259) and Raji-DC-SIGN-R cells as described previously [52].
DENV-1 virus capture ELISA
The capture ELISA for DENV-1 was based on a published assay for WNV with modifications [48]. Briefly, anti-DENV-1 MAbs were coated at indicated concentrations on MaxiSorp (Nunc) polystyrene 96-well microtiter plates in a sodium carbonate (pH 9.3) buffer. Plates were washed three times in ELISA wash buffer (PBS with 0.02% Tween 20) and blocked for 1 hour at 37°C with ELISA block buffer (PBS, 2% bovine serum albumin, and 0.02% Tween 20). DENV-1 virions (2.5 to 5.0×105 PFU of strain 16007 or West Pac-74) diluted in DMEM with 10% heat-inactivated FBS were captured on plates coated with anti-DENV-1 MAbs for 2 hours at room temperature. Plates were rinsed five times in wash buffer and then incubated with biotinylated 4G2 (1 µg/ml, respectively diluted in ELISA block buffer), which recognizes the fusion loop peptide in DII, for 1 hour at room temperature. Plates were washed again five times and then sequentially incubated with 2 µg/ml of horseradish peroxidase-conjugated streptavidin (Vector Laboratories) and tetramethylbenzidine substrate (Dako). The reaction was stopped with the addition of 2 N H2SO4 to the medium, and emission (450 nm) was read using an iMark microplate reader (Bio-Rad).
Mouse experiments
All mouse studies were approved and performed according to the guidelines of the Washington University School of Medicine Animal Safety Committee. IFN-αβR−/− × γR−/− mice on the 129 Sv background (AG129 mice) were a gift from Dr. Skip Virgin (Washington University School of Medicine) and bred in a pathogen-free barrier facility. In prophylaxis experiments, AG129 mice were administered a single dose of individual MAbs one day before infection. Subsequently, mice were challenged with West Pac-74 (genotype 4, 106 PFU) by an IP route and morbidity and mortality were monitored for 60 days. In post-exposure therapeutic experiments, a single 500 µg dose of MAb dose was administered by IP injection two or four days after infection with 106 PFU of West Pac-74.
Mapping of mutations onto the DENV-1 DIII crystal structure
Figures were prepared using the atomic coordinates of DENV-1 DIII (RCSB accession number 3IRC) using the programs CCP4MG [110]. The alignment of DENV-1 DIII from different genotypes was created with the program ALSCRIPT [111].
Statistical analysis
All data were analyzed using Prism software (GraphPad software). For survival analysis, Kaplan-Meier survival curves were analyzed by log-rank test. For neutralization assays an unpaired student's T-test was used to determine significance.
Supporting Information
Zdroje
1. BurkeDS
MonathTP
2001 Flaviviruses.
KnipeDM
HowleyPM
Fields Virology. Fourth Edition ed Philadelphia Lippincott Williams &Wilkins 1043 1125
2. Rico-HesseR
1990 Molecular evolution and distribution of dengue viruses type 1 and 2 in nature. Virology 174 479 493
3. HolmesEC
TwiddySS
2003 The origin, emergence and evolutionary genetics of dengue virus. Infect Genet Evol 3 19 28
4. HalsteadSB
1988 Pathogenesis of dengue: challenges to molecular biology. Science 239 476 481
5. MonathTP
1994 Dengue: the risk to developed and developing countries. Proc Natl Acad Sci U S A 91 2395 2400
6. ChambersTJ
HahnCS
GallerR
RiceCM
1990 Flavivirus genome organization, expression, and replication. Annu Rev Microbiol 44 649 688
7. KuhnRJ
ZhangW
RossmannMG
PletnevSV
CorverJ
2002 Structure of dengue virus: implications for flavivirus organization, maturation, and fusion. Cell 108 717 725
8. ZhangY
CorverJ
ChipmanPR
ZhangW
PletnevSV
2003 Structures of immature flavivirus particles. Embo J 22 2604 2613
9. ModisY
OgataS
ClementsD
HarrisonSC
2003 A ligand-binding pocket in the dengue virus envelope glycoprotein. Proc Natl Acad Sci U S A 100 6986 6991
10. ModisY
OgataS
ClementsD
HarrisonSC
2005 Variable surface epitopes in the crystal structure of dengue virus type 3 envelope glycoprotein. J Virol 79 1223 1231
11. NayakV
DessauM
KuceraK
AnthonyK
LedizetM
2009 Crystal structure of dengue virus type 1 envelope protein in the postfusion conformation and its implications for membrane fusion. J Virol 83 4338 4344
12. MondotteJA
LozachPY
AmaraA
GamarnikAV
2007 Essential Role of Dengue Virus Envelope Protein N Glycosylation at Asparagine-67 during Viral Propagation. J Virol 81 7136 7148
13. PokidyshevaE
ZhangY
BattistiAJ
Bator-KellyCM
ChipmanPR
2006 Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN. Cell 124 485 493
14. TassaneetrithepB
BurgessT
Granelli-PipernoA
TrumpfhellerC
FinkeJ
2003 DC-SIGN (CD209) mediates dengue virus infection of human dendritic cells. J Exp Med 197 823 829
15. Navarro-SanchezE
AltmeyerR
AmaraA
SchwartzO
FieschiF
2003 Dendritic-cell-specific ICAM3-grabbing non-integrin is essential for the productive infection of human dendritic cells by mosquito-cell-derived dengue viruses. EMBO Rep 4 723 728
16. ReyFA
HeinzFX
MandlC
KunzC
HarrisonSC
1995 The envelope glycoprotein from tick-borne encephalitis virus at 2 Angstrom resolution. Nature 375 291 298
17. BhardwajS
HolbrookM
ShopeRE
BarrettAD
WatowichSJ
2001 Biophysical characterization and vector-specific antagonist activity of domain III of the tick-borne flavivirus envelope protein. J Virol 75 4002 4007
18. VolkDE
BeasleyDW
KallickDA
HolbrookMR
BarrettAD
2004 Solution structure and antibody binding studies of the envelope protein domain III from the New York strain of West Nile virus. J Biol Chem 279 38755 38761
19. YuS
WuuA
BasuR
HolbrookMR
BarrettAD
2004 Solution structure and structural dynamics of envelope protein domain III of mosquito - and tick-borne flaviviruses. Biochemistry 43 9168 9176
20. LiL
LokSM
YuIM
ZhangY
KuhnRJ
2008 The flavivirus precursor membrane-envelope protein complex: structure and maturation. Science 319 1830 1834
21. YuIM
ZhangW
HoldawayHA
LiL
KostyuchenkoVA
2008 Structure of the immature dengue virus at low pH primes proteolytic maturation. Science 319 1834 1837
22. PiersonTC
FremontDH
KuhnRJ
DiamondMS
2008 Structural insights into the mechanisms of antibody-mediated neutralization of flavivirus infection: implications for vaccine development. Cell Host Microbe 4 229 238
23. RoehrigJT
MathewsJH
TrentDW
1983 Identification of epitopes on the E glycoprotein of Saint Louis encephalitis virus using monoclonal antibodies. Virology 128 118 126
24. RoehrigJT
StaudingerLA
HuntAR
MathewsJH
BlairCD
2001 Antibody prophylaxis and therapy for flaviviral encephalitis infections. Ann NY Acad Sci 286 297
25. LokSM
KostyuchenkoV
NybakkenGE
HoldawayHA
BattistiAJ
2008 Binding of a neutralizing antibody to dengue virus alters the arrangement of surface glycoproteins. Nat Struct Mol Biol 15 312 317
26. Sukupolvi-PettyS
PurthaWE
AustinSK
OliphantT
NybakkenG
2007 Type - and Sub-Complex-Specific Neutralizing Antibodies Against Domain III of Dengue Virus Type-2 Envelope Protein Recognize Adjacent Epitopes. J Virol 81 12816 12826
27. GromowskiGD
BarrettND
BarrettAD
2008 Characterization of dengue virus complex-specific neutralizing epitopes on envelope protein domain III of dengue 2 virus. J Virol 82 8828 8837
28. DiamondMS
SitatiE
FriendL
ShresthaB
HiggsS
2003 Induced IgM protects against lethal West Nile Virus infection. J Exp Med 198 1 11
29. OliphantT
EngleM
NybakkenG
DoaneC
JohnsonS
2005 Development of a humanized monoclonal antibody with therapeutic potential against West Nile virus. Nature Medicine 11 522 530
30. BrandrissMW
SchlesingerJJ
WalshEE
BriselliM
1986 Lethal 17D yellow fever encephalitis in mice. I. Passive protection by monoclonal antibodies to the envelope proteins of 17D yellow fever and dengue 2 viruses. J Gen Virol 67 229 234
31. GouldEA
BuckleyA
BarrettAD
CammackN
1986 Neutralizing (54K) and non-neutralizing (54K and 48K) monoclonal antibodies against structural and non-structural yellow fever virus proteins confer immunity in mice. J Gen Virol 67 ( Pt 3) 591 595
32. BeasleyDW
BarrettAD
2002 Identification of neutralizing epitopes within structural domain III of the West Nile virus envelope protein. J Virol 76 13097 13100
33. CrillWD
RoehrigJT
2001 Monoclonal antibodies that bind to domain III of dengue virus E glycoprotein are the most efficient blockers of virus adsorption to Vero cells. J Virol 75 7769 7773
34. RoehrigJT
BolinRA
KellyRG
1998 Monoclonal antibody mapping of the envelope glycoprotein of the dengue 2 virus, Jamaica. Virology 246 317 328
35. SanchezMD
PiersonTC
McAllisterD
HannaSL
PufferBA
2005 Characterization of neutralizing antibodies to West Nile virus. Virology 336 70 82
36. CeciliaD
GouldEA
1991 Nucleotide changes responsible for loss of neuroinvasiveness in Japanese encephalitis virus neutralization-resistant mutants. Virology 181 70 77
37. SeifSA
MoritaK
MatsuoS
HasebeF
IgarashiA
1995 Finer mapping of neutralizing epitope(s) on the C-terminal of Japanese encephalitis virus E-protein expressed in recombinant Escherichia coli system. Vaccine 13 1515 1521
38. WuSC
LianWC
HsuLC
LiauMY
1997 Japanese encephalitis virus antigenic variants with characteristic differences in neutralization resistance and mouse virulence. Virus Res 51 173 181
39. SchlesingerJJ
ChapmanS
NestorowiczA
RiceCM
GinocchioTE
1996 Replication of yellow fever virus in the mouse central nervous system: comparison of neuroadapted and non-neuroadapted virus and partial sequence analysis of the neuroadapted strain. J Gen Virol 77 ( Pt 6) 1277 1285
40. LinB
ParrishCR
MurrayJM
WrightPJ
1994 Localization of a neutralizing epitope on the envelope protein of dengue virus type 2. Virology 202 885 890
41. GromowskiGD
BarrettAD
2007 Characterization of an antigenic site that contains a dominant, type-specific neutralization determinant on the envelope protein domain III (ED3) of dengue 2 virus. Virology 366 349 360
42. LaiCJ
GoncalvezAP
MenR
WernlyC
DonauO
2007 Epitope determinants of a chimpanzee dengue virus type 4 (DENV-4)-neutralizing antibody and protection against DENV-4 challenge in mice and rhesus monkeys by passively transferred humanized antibody. J Virol 81 12766 12774
43. DiamondMS
2009 Progress on the development of therapeutics against West Nile virus. Antiviral Res 83 214 227
44. SimantiniE
BanerjeeK
1995 Epitope mapping of dengue 1 virus E glycoprotein using monoclonal antibodies. Arch Virol 140 1257 1273
45. ChenYC
HuangHN
LinCT
ChenYF
KingCC
2007 Generation and characterization of monoclonal antibodies against dengue virus type 1 for epitope mapping and serological detection by epitope-based peptide antigens. Clin Vaccine Immunol 14 404 411
46. BeasleyDW
AaskovJG
2001 Epitopes on the dengue 1 virus envelope protein recognized by neutralizing IgM monoclonal antibodies. Virology 279 447 458
47. GoncalvezAP
EscalanteAA
PujolFH
LudertJE
TovarD
2002 Diversity and evolution of the envelope gene of dengue virus type 1. Virology 303 110 119
48. VogtMR
MoeskerB
GoudsmitJ
JongeneelenM
AustinSK
2009 Human Monoclonal Antibodies Induced by Natural Infection Against West Nile Virus Neutralize at a Post-Attachment Step. J Virol 83 6494 6507
49. MorreyJD
SiddharthanV
OlsenAL
RoperGY
WangH
2006 Humanized monoclonal antibody against West Nile virus E protein administered after neuronal infection protects against lethal encephalitis in hamsters. J Infect Dis 194 1300 1308
50. MorreyJD
SiddharthanV
OlsenAL
WangH
JulanderJG
2007 Defining limits of humanized neutralizing monoclonal antibody treatment for West Nile virus neurological infection in a hamster model. Antimicrob Agents Chemother 51 2396 2402
51. ZhangW
ChipmanPR
CorverJ
JohnsonPR
ZhangY
2003 Visualization of membrane protein domains by cryo-electron microscopy of dengue virus. Nat Struct Biol 10 907 912
52. OliphantT
NybakkenGE
EngleM
XuQ
NelsonCA
2006 Antibody recognition and neutralization determinants on domains I and II of West Nile Virus envelope protein. J Virol 80 12149 12159
53. GouldLH
SuiJ
FoellmerH
OliphantT
WangT
2005 Protective and therapeutic capacity of human single chain Fv-Fc fusion proteins against West Nile virus. J Virol 79 14606 14613
54. ThrosbyM
GeuijenC
GoudsmitJ
BakkerAQ
KorimbocusJ
2006 Isolation and characterization of human monoclonal antibodies from individuals infected with West Nile Virus. J Virol 80 6982 6992
55. SultanaH
FoellmerHG
NeelakantaG
OliphantT
EngleM
2009 Fusion loop peptide of the West Nile virus envelope protein is essential for pathogenesis and is recognized by a therapeutic cross-reactive human monoclonal antibody. J Immunol 183 650 660
56. JohnsonAJ
RoehrigJT
1999 New mouse model for dengue virus vaccine testing. J Virol 73 783 786
57. GoncalvezAP
ChienCH
TubthongK
GorshkovaI
RollC
2008 Humanized monoclonal antibodies derived from chimpanzee Fabs protect against Japanese encephalitis virus in vitro and in vivo. J Virol 82 7009 7021
58. GoncalvezAP
PurcellRH
LaiCJ
2004 Epitope determinants of a chimpanzee Fab antibody that efficiently cross-neutralizes dengue type 1 and type 2 viruses map to inside and in close proximity to fusion loop of the dengue type 2 virus envelope glycoprotein. J Virol 78 12919 12928
59. LeitmeyerKC
VaughnDW
WattsDM
SalasR
VillalobosI
1999 Dengue virus structural differences that correlate with pathogenesis. J Virol 73 4738 4747
60. Rico-HesseR
2007 Dengue virus evolution and virulence models. Clin Infect Dis 44 1462 1466
61. Rico-HesseR
HarrisonLM
SalasRA
TovarD
NisalakA
1997 Origins of dengue type 2 viruses associated with increased pathogenicity in the Americas. Virology 230 244 251
62. ArmstrongPM
Rico-HesseR
2003 Efficiency of dengue serotype 2 virus strains to infect and disseminate in Aedes aegypti. Am J Trop Med Hyg 68 539 544
63. Rico-HesseR
2003 Microevolution and virulence of dengue viruses. Adv Virus Res 59 315 341
64. LisovaO
HardyF
PetitV
BedouelleH
2007 Mapping to completeness and transplantation of a group-specific, discontinuous, neutralizing epitope in the envelope protein of dengue virus. J Gen Virol 88 2387 2397
65. ThullierP
DemangelC
BedouelleH
MegretF
JouanA
2001 Mapping of a dengue virus neutralizing epitope critical for the infectivity of all serotypes: insight into the neutralization mechanism. J Gen Virol 82 1885 1892
66. RajamanonmaniR
NkenfouC
ClancyP
YauYH
ShochatSG
2009 On a mouse monoclonal antibody that neutralizes all four dengue virus serotypes. J Gen Virol 90 799 809
67. MatsuiK
GromowskiGD
LiL
SchuhAJ
LeeJC
2009 Characterization of dengue complex-reactive epitopes on dengue 3 virus envelope protein domain III. Virology 384 16 20
68. CrillWD
ChangGJ
2004 Localization and characterization of flavivirus envelope glycoprotein cross-reactive epitopes. J Virol 78 13975 13986
69. CrillWD
TrainorNB
ChangGJ
2007 A detailed mutagenesis study of flavivirus cross-reactive epitopes using West Nile virus-like particles. J Gen Virol 88 1169 1174
70. ChenZ
LiuLM
GaoN
XuXF
ZhangJL
2009 Passive protection assay of monoclonal antibodies against dengue virus in suckling mice. Curr Microbiol 58 326 331
71. KaufmanBM
SummersPL
DuboisDR
EckelsKH
1987 Monoclonal antibodies against dengue 2 virus E-glycoprotein protect mice against lethal dengue infection. Am J Trop Med Hyg 36 427 434
72. EngleM
DiamondMS
2003 Antibody prophylaxis and therapy against West Nile Virus infection in wild type and immunodeficient mice. J Virol 77 12941 12949
73. Ben-NathanD
Gershoni-YahalomO
SaminaI
KhinichY
NurI
2009 Using high titer West Nile intravenous immunoglobulin from selected Israeli donors for treatment of West Nile virus infection. BMC Infect Dis 9 18
74. Ben-NathanD
LustigS
TamG
RobinzonS
SegalS
2003 Prophylactic and therapeutic efficacy of human intravenous immunoglobulin in treating west nile virus infection in mice. J Infect Dis 188 5 12
75. ZhangMJ
WangMJ
JiangSZ
MaWY
1989 Passive protection of mice, goats, and monkeys against Japanese encephalitis with monoclonal antibodies. J Med Virol 29 133 138
76. Kimura-KurodaJ
YasuiK
1988 Protection of mice against Japanese encephalitis virus by passive administration with monoclonal antibodies. J Immunol 141 3606 3610
77. KreilTR
EiblMM
1997 Pre - and postexposure protection by passive immunoglobulin but no enhancement of infection with a flavivirus in a mouse model. J Virol 71 2921 2927
78. VaughnDW
GreenS
KalayanaroojS
InnisBL
NimmannityaS
2000 Dengue Viremia Titer, Antibody Response Pattern, and Virus Serotype Correlate with Disease Severity. J Infect Dis 181 2 9
79. XiaoSY
GuzmanH
ZhangH
Travassos da RosaAP
TeshRB
2001 West Nile virus infection in the golden hamster (Mesocricetus auratus): a model for West Nile encephalitis. Emerg Infect Dis 7 714 721
80. DiamondMS
ShresthaB
MarriA
MahanD
EngleM
2003 B cells and antibody play critical roles in the immediate defense of disseminated infection by West Nile encephalitis virus. J Virol 77 2578 2586
81. PiersonTC
XuQ
NelsonS
OliphantT
NybakkenGE
2007 The stoichiometry of antibody-mediated neutralization and enhancement of West Nile virus infection. Cell Host and Microbe 1 135 145
82. KlassePJ
2007 Modeling how many envelope glycoprotein trimers per virion participate in human immunodeficiency virus infectivity and its neutralization by antibody. Virology 369 245 262
83. GollinsS
PorterfieldJ
1984 Flavivirus infection enhancement in macrophages: radioactive and biological studies on the effect of antibody and viral fate. J Gen Virol 65 1261 1272
84. HalsteadSB
1994 Antibody-dependent enhancement of infection: a mechanism for indirect virus entry into cells. Cellular Receptors for Animal Viruses Cold Spring Harbor Cold Spring Harbor Laboratory Press 493 516
85. HalsteadSB
1979 In vivo enhancement of dengue virus infection in rhesus monkeys by passively transferred antibody. J Infect Dis 140 527 533
86. GoncalvezAP
EngleRE
St ClaireM
PurcellRH
LaiCJ
2007 Monoclonal antibody-mediated enhancement of dengue virus infection in vitro and in vivo and strategies for prevention. Proc Natl Acad Sci U S A 104 9422 9427
87. MehlhopE
Ansarah-SobrinhoC
JohnsonS
EngleM
FremontDH
2007 Complement protein C1q inhibits antibody-dependent enhancement of flavivirus infection in an IgG subclass-specific manner. Cell Host and Microbe 2 417 426
88. BalsitisSJ
WilliamsKL
LachicaR
FloresD
KyleJL
2010 Lethal antibody enhancement of dengue disease in mice is prevented by Fc modification. PLoS Pathog 6 e1000790 doi:10.1371/journal.ppat.1000790
89. HiramatsuK
TadanoM
MenR
LaiCJ
1996 Mutational analysis of a neutralization epitope on the dengue type 2 virus (DEN2) envelope protein: monoclonal antibody resistant DEN2/DEN4 chimeras exhibit reduced mouse neurovirulence. Virology 224 437 445
90. SerafinIL
AaskovJG
2001 Identification of epitopes on the envelope (E) protein of dengue 2 and dengue 3 viruses using monoclonal antibodies. Arch Virol 146 2469 2479
91. MegretF
HugnotJP
FalconarA
GentryMK
MorensDM
1992 Use of recombinant fusion proteins and monoclonal antibodies to define linear and discontinuous antigenic sites on the dengue virus envelope glycoprotein. Virology 187 480 491
92. TrirawatanapongT
ChandranB
PutnakR
PadmanabhanR
1992 Mapping of a region of dengue virus type-2 glycoprotein required for binding by a neutralizing monoclonal antibody. Gene 116 139 150
93. NybakkenG
OliphantT
JohnsonS
BurkeS
DiamondMS
2005 Structural basis for neutralization of a therapeutic antibody against West Nile virus. Nature 437 764 769
94. CherrierMV
KaufmannB
NybakkenGE
LokSM
WarrenJT
2009 Structural basis for the preferential binding of immature flaviviruses by a fusion-loop specific antibody. EMBO 28 3269 3276
95. OliphantT
NybakkenGE
AustinSK
XuQ
BramsonJ
2007 The Induction of Epitope-Specific Neutralizing Antibodies against West Nile virus. J Virol 81 11828 11839
96. WahalaWM
KrausAA
HaymoreLB
Accavitti-LoperMA
de SilvaAM
2009 Dengue virus neutralization by human immune sera: role of envelope protein domain III-reactive antibody. Virology 392 103 113
97. CrillWD
HughesHR
DeloreyMJ
ChangGJ
2009 Humoral immune responses of dengue fever patients using epitope-specific serotype-2 virus-like particle antigens. PLoS ONE 4 e4991 doi:10.1371/journal.pone.0004991
98. HermidaL
BernardoL
MartinJ
AlvarezM
PradoI
2006 A recombinant fusion protein containing the domain III of the dengue-2 envelope protein is immunogenic and protective in nonhuman primates. Vaccine 24 3165 3171
99. ChuJH
ChiangCC
NgML
2007 Immunization of Flavivirus West Nile Recombinant Envelope Domain III Protein Induced Specific Immune Response and Protection against West Nile Virus Infection. J Immunol 178 2699 2705
100. ChenS
YuM
JiangT
DengY
QinC
2007 Induction of tetravalent protective immunity against four dengue serotypes by the tandem domain III of the envelope protein. DNA Cell Biol 26 361 367
101. ValdesI
BernardoL
GilL
PavonA
LazoL
2009 A novel fusion protein domain III-capsid from dengue-2, in a highly aggregated form, induces a functional immune response and protection in mice. Virology 394 249 258
102. HarlowE
LaneD
1988 Antibodies, A laboratory manual Cold Spring Harbor Cold Spring Harbor Laboratory 714
103. MinorW
CymborowskiM
OtwinowskiZ
ChruszczM
2006 HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr D Biol Crystallogr 62 859 866
104. McCoyAJ
Grosse-KunstleveRW
AdamsPD
WinnMD
StoroniLC
2007 Phaser crystallographic software. J Appl Crystallogr 40 658 674
105. JonesEY
WalkerNP
StuartDI
1991 Methodology employed for the structure determination of tumour necrosis factor, a case of high non-crystallographic symmetry. Acta Crystallogr A 47 ( Pt 6) 753 770
106. WinnMD
MurshudovGN
PapizMZ
2003 Macromolecular TLS refinement in REFMAC at moderate resolutions. Methods Enzymol 374 300 321
107. ShindyalovIN
BournePE
1998 Protein structure alignment by incremental combinatorial extension (CE) of the optimal path. Protein Eng 11 739 747
108. BoderET
WittrupKD
1997 Yeast surface display for screening combinatorial polypeptide libraries. Nat Biotechnol 15 553 557
109. BoderET
WittrupKD
1998 Optimal screening of surface-displayed polypeptide libraries. Biotechnol Prog 14 55 62
110. PottertonL
McNicholasS
KrissinelE
GruberJ
CowtanK
2004 Developments in the CCP4 molecular-graphics project. Acta Crystallogr D Biol Crystallogr 60 2288 2294
111. BartonGJ
1993 ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng 6 37 40
112. KabschW
SanderC
1983 Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22 2577 2637
Štítky
Hygiena a epidemiologie Infekční lékařství Laboratoř
Článek Suppression of mRNAs Encoding Tegument Tetraspanins from Results in Impaired Tegument TurnoverČlánek Novel Riboswitch Ligand Analogs as Selective Inhibitors of Guanine-Related Metabolic PathwaysČlánek The Physical Relationship between Infectivity and Prion Protein Aggregates Is Strain-DependentČlánek Rhomboid 4 (ROM4) Affects the Processing of Surface Adhesins and Facilitates Host Cell Invasion by
Článek vyšel v časopisePLOS Pathogens
Nejčtenější tento týden
2010 Číslo 4- Perorální antivirotika jako vysoce efektivní nástroj prevence hospitalizací kvůli COVID-19 − otázky a odpovědi pro praxi
- Parazitičtí červi v terapii Crohnovy choroby a dalších zánětlivých autoimunitních onemocnění
- Vakcíny proti klíšťové encefalitidě
- Kdy je nejlepší očkovat
- Možné vedlejší účinky očkování
-
Všechny články tohoto čísla
- Innate Recognition of Fungal Cell Walls
- Suppression of mRNAs Encoding Tegument Tetraspanins from Results in Impaired Tegument Turnover
- Junín Virus Infection of Human Hematopoietic Progenitors Impairs Proplatelet Formation and Platelet Release via a Bystander Effect Involving Type I IFN Signaling
- The Endosymbiotic Bacterium Induces Resistance to Dengue Virus in
- Natural Regulatory T Cells in Malaria: Host or Parasite Allies?
- Keratinocytes Determine Th1 Immunity during Early Experimental Leishmaniasis
- Spatial and Temporal Association of Outbreaks of H5N1 Influenza Virus Infection in Wild Birds with the 0°C Isotherm
- Novel Riboswitch Ligand Analogs as Selective Inhibitors of Guanine-Related Metabolic Pathways
- RNA Polymerase Activity and Specific RNA Structure Are Required for Efficient HCV Replication in Cultured Cells
- The Physical Relationship between Infectivity and Prion Protein Aggregates Is Strain-Dependent
- Inadequate Clearance of Translocated Bacterial Products in HIV-Infected Humanized Mice
- Topology and Organization of the Type III Secretion Needle Complex Components
- Temperature Modulates Plant Defense Responses through NB-LRR Proteins
- Peptide Inhibitors of Dengue-Virus Entry Target a Late-Stage Fusion Intermediate
- Identification of Host-Dependent Survival Factors for Intracellular through an siRNA Screen
- Exposure to HIV-1 Directly Impairs Mucosal Epithelial Barrier Integrity Allowing Microbial Translocation
- Increased Asymmetric Dimethylarginine in Severe Falciparum Malaria: Association with Impaired Nitric Oxide Bioavailability and Fatal Outcome
- Reconstitution of SARS-Coronavirus mRNA Cap Methylation
- Induces Brain Microvascular Endothelial Cell Detachment from the Matrix and Cleavage of Occludin: A Role for MMP-8
- Two Coregulated Efflux Transporters Modulate Intracellular Heme and Protoporphyrin IX Availability in
- The Type I NADH Dehydrogenase of Counters Phagosomal NOX2 Activity to Inhibit TNF-α-Mediated Host Cell Apoptosis
- Rhomboid 4 (ROM4) Affects the Processing of Surface Adhesins and Facilitates Host Cell Invasion by
- Increased Monocyte Turnover from Bone Marrow Correlates with Severity of SIV Encephalitis and CD163 Levels in Plasma
- The RING-CH Ligase K5 Antagonizes Restriction of KSHV and HIV-1 Particle Release by Mediating Ubiquitin-Dependent Endosomal Degradation of Tetherin
- Molecular Mechanisms of Ethanol-Induced Pathogenesis Revealed by RNA-Sequencing
- Highly Frequent Mutations in Negative Regulators of Multiple Virulence Genes in Group A Streptococcal Toxic Shock Syndrome Isolates
- Emergence and Pathogenicity of Highly Virulent Genotypes in the Northwest United States
- Structural and Functional Analysis of Viral siRNAs
- Prion Shedding from Olfactory Neurons into Nasal Secretions
- a GATA Transcription Factor That Directs Disparate Fates in Including Morphogenesis and Siderophore Biosynthesis
- Three Members of the 6-cys Protein Family of Play a Role in Gamete Fertility
- Complement as an Endogenous Adjuvant for Dendritic Cell-Mediated Induction of Retrovirus-Specific CTLs
- A Genomic Survey of Positive Selection in Provides Insights into the Evolution of Accidental Virulence
- Overcomes Stress of Azole Drugs by Formation of Disomy in Specific Multiple Chromosomes
- Blood Fluke Exploitation of Non-Cognate CD4 T Cell Help to Facilitate Parasite Development
- Antagonism of Tetherin Restriction of HIV-1 Release by Vpu Involves Binding and Sequestration of the Restriction Factor in a Perinuclear Compartment
- The Development of Therapeutic Antibodies That Neutralize Homologous and Heterologous Genotypes of Dengue Virus Type 1
- Deficiencies in Jasmonate-Mediated Plant Defense Reveal Quantitative Variation in Pathogenesis
- Interactions with Bacteria in the Context of Human Health and Disease
- Viral Capsid Is a Pathogen-Associated Molecular Pattern in Adenovirus Keratitis
- Electron Tomography Reveals the Steps in Filovirus Budding
- Selective Condensation Drives Partitioning and Sequential Secretion of Cyst Wall Proteins in Differentiating
- The Effect of Vaccination on the Evolution and Population Dynamics of Avian Paramyxovirus-1
- A Timescale for Evolution, Population Expansion, and Spatial Spread of an Emerging Clone of Methicillin-Resistant
- VacA Toxin/Subunit p34: Targeting of an Anion Channel to the Inner Mitochondrial Membrane
- Production of Extracellular Traps against and in Infected Lung Tissue Is Dependent on Invading Neutrophils and Influenced by Hydrophobin RodA
- A Differential Role for Macropinocytosis in Mediating Entry of the Two Forms of Vaccinia Virus into Dendritic Cells
- Impaired Innate Immunity in Mice but Preserved CD8 T Cell Responses against in -, -, - or -Deficient Mice
- SARS-CoV Pathogenesis Is Regulated by a STAT1 Dependent but a Type I, II and III Interferon Receptor Independent Mechanism
- Proteolysis of Human Thrombin Generates Novel Host Defense Peptides
- Multilayered Mechanism of CD4 Downregulation by HIV-1 Vpu Involving Distinct ER Retention and ERAD Targeting Steps
- PLOS Pathogens
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- The Effect of Vaccination on the Evolution and Population Dynamics of Avian Paramyxovirus-1
- Reconstitution of SARS-Coronavirus mRNA Cap Methylation
- Increased Monocyte Turnover from Bone Marrow Correlates with Severity of SIV Encephalitis and CD163 Levels in Plasma
- Deficiencies in Jasmonate-Mediated Plant Defense Reveal Quantitative Variation in Pathogenesis
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



