-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Fine-mapping of the human leukocyte antigen locus as a risk factor for Alzheimer disease: A case–control study
In a case–control study, Jennifer Yokoyama and colleagues present fine-mapping of the human leukocyte antigen genetic region to identify haplotypes associated with Alzheimer disease.
Published in the journal: . PLoS Med 14(3): e32767. doi:10.1371/journal.pmed.1002272
Category: Research Article
doi: https://doi.org/10.1371/journal.pmed.1002272Summary
In a case–control study, Jennifer Yokoyama and colleagues present fine-mapping of the human leukocyte antigen genetic region to identify haplotypes associated with Alzheimer disease.
Introduction
Alzheimer disease (AD) is a progressive neurodegenerative disorder and has a global burden of approximately 46 million people worldwide, with prevalence projected to double over the next 20 y [1]. The hallmark features of the disease include the accumulation of amyloid plaques, tau neurofibrillary tangles, and neuronal destruction, leading to brain atrophy and loss of cognitive function. The etiology of these processes stems from synergistic interactions of environmental and genetic factors, many of which remain obscure and therefore complicate research efforts aimed at identifying efficacious therapies.
The three largest genetic contributors identified thus far are rare variants in amyloid precursor protein (APP) and presenilin 1 and 2 (PSEN1, PSEN2) [2]. These variants are uncommon, cause an early onset form of the disease, and typically segregate in an autosomal dominant fashion. Studies of late onset AD (typically defined as onset age >65 y) have demonstrated the risk of sequence variants such as the common ε4 allele of apolipoprotein E (APOE), rare variation in TREM2 [3–7], and MAPT [8], as well as numerous common variants contributing modest AD risk [9], including single nucleotide polymorphisms (SNPs) in the following loci: CR1, BIN1, INPP5D, MEF2C, CD2AP, ZCWPW1, NME8, EPHA1, CLU, PICALM, MS4A4, CELF1, FERMT2, ABCA7, CD33, CASS4, PTK2B, SORL1, SLC24A4-RIN3, DSG2, and HLA-DRB5/HLA-DRB1 [9,10]. However, there remain additional unexplained genetic contributions to nonfamilial forms of AD, suggesting polygenic contributors as well as the potential for epistatic and epigenetic interactions [11].
There is increasing support for the role of neuroinflammation in the etiology of AD as well as evidence that inflammatory processes are an early event in the brains of patients with AD [12]. Several studies have provided biochemical and histological evidence of classic immune components, including active microglia [13–15], complement factors [16,17], inflammatory cytokines [18], and C-reactive protein [19] within the parenchyma of AD brains. This is further supported by work in mouse models providing strong evidence for the role of complement-dependent destruction of synapses by phagocytic microglia prior to plaque deposition; similar mechanisms may even contribute to age-related cognitive decline [20,21]. Given these findings, there is a great deal of interest in identifying genetic determinants of inflammation related to AD susceptibility.
Located on chromosome 6p21, the major histocompatibility complex (MHC) is a dense region of approximately 150 genes that encode the human leukocyte antigen (HLA) immunoregulatory proteins [22]. Because of their proximity to each other, many of the MHC genes exist in linkage disequilibrium (LD) and are inherited as haplotypes with varying frequencies in global populations. MHC genes encode cell surface receptors and are classified based on their ability to present endogenous or exogenous antigens to T cells. MHC class I proteins exist on the surface of all nucleated cells and present fragments of antigens generated intracellularly to CD8+ T cells to induce a cytokine-mediated immune response. MHC class II molecules are only expressed by professional antigen-presenting cells, including B cells, macrophages, and microglia, and present exogenous material taken into the cell via endocytic vesicles to CD4+ T cells. Together, the diverse repertoire of the human immune system partly stems from the extremely polymorphic nature of the MHC class I and II regions.
Many associations are established between neurodegenerative and autoimmune diseases, specific class I and II alleles, and combinations of alleles (haplotypes) in the HLA region. Previous genome-wide association studies (GWASs), pleiotropic analyses, and meta-analyses by our group and others have investigated MHC susceptibility loci in a wide range of diseases, including AD [9,23,24]. However, because of the complex genetic organization of the HLA region and differences in the haplotype substructure of different ethnic populations, as well as differences in sequencing and allelic imputation methods, studies have yet to definitively elucidate which genes and specific alleles contribute to the observed association signals.
As mentioned, HLA-DRB5/HLA-DRB1 has been implicated in numerous GWASs as a significant contributor to AD risk [9]. This prior work has established a significant association of the HLA locus to AD risk in over 75,000 individuals, yet the specific allele or alleles contributing to this association remain elusive. We thus used a robust HLA imputation method and case–control approach to fine-map the contributions of HLA polymorphisms and haplotypes to AD in over 11,500 patients and controls from independent cohorts from the University of California, San Francisco (UCSF) Memory and Aging Center (MAC) and the Alzheimer’s Disease Genetics Consortium (ADGC) (S1 File). We also examined longitudinal neuropsychological measures of cognitive function and cross-sectional biomarker data from cerebrospinal fluid (CSF) from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) to assess the clinical relevance of identified risk haplotypes.
Materials and methods
Participants were consented (as described below) for research in accordance with the Institutional Review Board at the University of California, San Francisco, and institutional review boards at each site for multicenter study data approved all aspects of this study as they fall under the purview of the respective research groups (ADNI and ADGC).
Participants
UCSF MAC cohort
The participants included in this study were 309 white individuals over the age of 50 y, including 191 controls and 118 individuals with AD seen at the UCSF MAC between 1999–2012 who were genotyped as part of their participation in longitudinal research on neurodegenerative disease and healthy cognitive aging. DNA from the UCSF MAC cohort was collected from 2000–2012, and genotyping was performed in 2012. Because individuals are followed up longitudinally, we verified clinical diagnosis at the beginning of this study (May 2015). A multidisciplinary team of neurologists, neuropsychologists, and nurses performed a detailed evaluation on individuals with AD and established a diagnosis according to consensus criteria for AD [25]. Individuals included as controls underwent a similar assessment and were diagnosed as having normal cognition for their age. Participants who carried a known genetic risk variant in APP, PSEN1, or PSEN2 were excluded from this study. Participants or surrogates completed written informed consent for all genetic research related to neurodegenerative disease and healthy cognitive aging during their initial visit in accordance with the Institutional Review Board at the University of California, San Francisco.
ADGC
The ADGC is an NIH-funded collection of GWAS data created for the goal of identifying genetic contributions to late-onset AD. Participants included in this study were from 30 merged datasets combined by Boehme, Mukherjee, Crane, and Kauwe and included 28,730 individuals carrying either an AD or cognitively normal control clinical diagnosis [26] A list of the datasets and basic information is included in S1 Table; full details on the datasets and the merging process are available at http://kauwelab.byu.edu/Portals/22/adgc_combined_1000G_12032014.pdf [26]. Analyses were limited to white individuals for maximum statistical power to reduce potential for confounding due to the known population-based contribution to diversity in the HLA region. Participants were recruited and seen between 1984–2012. Written informed consent for genetic studies falling under the purview of the ADGC was obtained from all study participants, and institutional review boards at each site approved all aspects of this study. Specific consent for this study was obtained from the ADGC based on an application describing the proposed work.
ADNI
We also utilized data from 346 individuals recruited for participation in the ADNI study with data from SNP genotyping and longitudinal cognitive scores. All individuals included in this study had a minimum of two clinic assessments. At baseline, 120 individuals were cognitively normal (CN) older adults, 113 individuals were diagnosed with mild cognitive impairment (MCI), and 113 with AD. Of these, 163 individuals also had CSF measurements of plasma biomarkers available (S2 Table). The ADNI cohort is well characterized and has been used in previously published studies [27–29]. The clinical severity of symptoms in the MCI and AD groupings was measured using the Clinical Dementia Rating sum of boxes (CDR-SB) [30]. A clinician diagnosed each participant using a structured protocol that utilized clinical judgment and neuropsychological tests that are provided in S1 Methods. The mean follow-up time was 3.15 ± 2.04 y for control participants (n = 91), 2.39 ± 1.71 y for participants with MCI (n = 148), and 1.37 ± 0.75 y for patients with AD (n = 69). Written informed consent was obtained from all study participants for research studies falling under the purview of ADNI, and the University of California, San Francisco Institutional Review Board approved all aspects of this study.
Genotype acquisition
UCSF MAC cohort
Patient and control genotypes were obtained via genotyping on the Illumina Omni1-Quad array (Illumina, San Diego, California) using manufacturer’s instructions. APOE genotype was determined with a TaqMan Allelic Discrimination Assay for the two SNPs, rs429358 and rs7412, on an ABI 7900HT Fast Real-Time PCR system (Applied Biosystems, Foster City, California) using the manufacturer's instructions.
ADGC
Details of genotyping in the 30 datasets that comprise the combined ADGC dataset are available online [26] and partially described in previously published papers [10].
ADNI
Haplotypes were determined using genotypes from the Human610-Quad BeadChip (Illumina, San Diego, California) as previously described [31]. APOE genotypes were determined by Cogenics (now Beckman Coulter; Pasadena, California).
CSF biomarker measurements
ADNI
Baseline CSF biomarkers levels were measured using the Human DiscoveryMAP panel developed by Rules Based Medicine (Myriad RBM; Austin, Texas). The Human DiscoveryMAP panel is commercially available and measures a collection of metabolic, lipid, inflammatory, and other AD-relevant indicators. We limited our analyses to 28 immune proteins in the panel that were associated with inflammatory or immune processes (S1 List). The samples were processed and analyzed by Myriad RBM and checked for quality by the ADNI Biomarker core. CSF amyloid β 1–42 was measured using the AlzBio3 Luminex xMAP immunoassay (Innogenetics, Ghent, Belgium) according to previously described methods [32]. This method utilizes a monoclonal antibody specific for amyloid β 1–42 that is chemically bonded to color-coded beads along with analyte-specific detector antibodies. Additional details are available in S1 Methods.
Clinical assessments
ADNI
In this study, we analyzed two neuropsychological measures of cognitive function and one measure of clinical severity in ADNI participants. The Rey Auditory Verbal Learning Test (RAVLT) [33] is a test of verbal memory. It begins with the administrator reading a list of 15 unrelated words to the participant, who is then asked to verbally repeat as many of the words as they can. This happens for a total of five learning trials, and the administrator records the number of words correctly recalled after each trial. The test administrator then reads a set of 15 new words to the participant (interference word list), and, immediately following this, the participant is asked to recall as many of the first list of words as possible (immediate recall score). After a 30-min delay during which unrelated tests are administered, the participant is asked to recall as many words as possible from the initial list (delayed recall score). The RAVLT “forgetting score” is calculated as the difference between immediate recall versus delayed recall scores [33]. The forgetting score remains relatively stable over time in individuals with consistent memory function; the forgetting score tends to get smaller as the number of recalled items decreases. The 11-item Alzheimer’s Disease Assessment Scale (ADAS) cognitive subscale assesses learning and memory, orientation, and several aspects of language including production, comprehension, and constructional and ideational praxis [34,35]. Higher scores indicate more impairment. Finally, the Clinical Dementia Rating (CDR) scale is a measure of three cognitive domains (memory, problem solving, and orientation) and three functional domains (self-care, community engagement, and hobbies). Information is collected directly from the study participant, as well as from a study informant. The scores for the six domains are combined into the CDR sum of boxes (CDR-SB) score [36].
Statistical analysis
Cohort demographic summary statistics
Summary statistics for participants’ age, sex, age of onset, and APOE ε4 carrier status were calculated using R.
Imputation of HLA alleles
HLA genotypes were derived from chromosome 6 SNP data using an imputation program, HLA Genotype Imputation with Attribute Bagging (HIBAG) v1.3, which calculates predictions of genotype by averaging HLA-type posterior probabilities over an ensemble of classifiers built on bootstrap samples [37]. It relies on a training set of known HLA and SNP genotypes. We imputed the following HLA genes: A, B, DRB1, DQA1, and DQB1. For the UCSF MAC cohort and the ADGC merged dataset, a training set for four-digit resolution using ethnic-specific models for Europeans based on Omni1_Quad_v1_0_H was used. For the ADNI cohort clinical biomarker analyses, we used four-digit resolution ethnic-specific models for Europeans derived from Illumina Human610-Quad v1.0.
Quality control of HLA imputation
Based on the distribution of posterior probabilities for each of the five imputed alleles (see S1 Fig), we chose a call threshold (CT) of 0.75. As previous studies have shown that a CT of 0.5 leads to HIBAG prediction accuracies of 94.8%–99.2% for individuals of European ancestry [38], we expect our more stringent CT will correspond to similar or higher HIBAG prediction accuracies based on assumed accuracy of imputed ADGC SNPs. After excluding samples with any imputation probability below this cutoff at any locus, our final ADGC cohort size was 11,381.
Calculating locus and haplotype odds ratios (ORs)
OR estimates for patients with AD and cognitively normal controls were calculated using a statistical package designed to specifically probe associations with the HLA (BIGDAWG), including tests of Hardy-Weinberg equilibrium and case–control association analyses for haplotypes as previously described [39]. Analyses were performed for each cohort (UCSF + ADGC) separately and in combination. As this was a fine-mapping study based on a previous genome-wide significant, and replicated, finding at HLA-DRB5; and considering that this study represents a first analysis of the highly polymorphic HLA region in the context of AD, a complex disease, we did not require a multiple testing correction. To strike a balance between reducing Type I error while also allowing for full exploration of the loci underlying this association with the MHC region, we implemented a stepwise assessment of HLA gene contributions to AD: using allelic information, we established a priori significance at p < 0.05 at the haplotype level based on the prior GWAS-significant results. We then examined the contingency table from which the haplotype result was derived to identify the specific allele(s) contributing to the association signal. We accepted allele-level significance at p < 0.05 given the haplotype-level significance [40,41]. Based on a sample size of 11,690 in our combined UCSF+ADGC cohort, with 326 degrees of freedom and an alpha of 0.05, we had 64.1% power to detect an OR of 1.21 based on the haplotype frequencies of AD versus cognitively normal controls for the top associated five-allele haplotype.
Biomarker and cognitive data
Discrete and continuous demographic variables were compared across the ADNI cohort using chi-squared and ANOVA analyses, respectively. Linear mixed effects models were used to assess the relationship between the risk haplotype of interest and changes in the longitudinal cognitive measurements, ADAS and RAVLT, while controlling for baseline and time interactions of age, sex, education, baseline CDR-SB score (to account for baseline differences in clinical severity/diagnosis), and APOE ε4 carrier status. Use of linear mixed effects models allowed us to account for variable data missingness across participants by estimating subject-specific slopes. This enabled us to estimate cognitive changes for each individual despite varying numbers of visits. Missing data were omitted from the analyses, and all participants were required to have at least two time points to be included in the analysis. All interactions and main effects were modeled as fixed effects with random slopes and intercepts across individuals. The main effects of all variables were included in all longitudinal analyses but have been omitted from the definitions below to improve their clarity.
The linear mixed effects model for ADAS11 scores was defined as follows:
The linear mixed effects model for the RAVLT forgetting score was defined as follows:
Cross-sectional CSF biomarker analyses
Linear models were used to test for an association between baseline CSF biomarker levels and the haplotype of interest. We controlled for age, sex, education, baseline CDR-SB score (to account for baseline differences in clinical severity/diagnosis), and APOE ε4 dosage.
Results
Five-allele haplotype analysis implicated DR15 in AD risk
The discovery UCSF cohort consisted of 309 individuals with clinically diagnosed AD and cognitively normal older adult controls (Table 1). Because of the small sample size, all imputed alleles were included in the haplotype analysis (HLA A, B, DRB1, DQA1, and DQB1). We performed association analysis on the four haplotypes with sufficient frequency in this small cohort. Of these four, one showed a significant association with AD risk: HLA A*02:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 (OR = 3.69; 95% confidence interval [CI] 1.16–13.69; p = 0.01) (Table 2).
Tab. 1. Cohort demographics.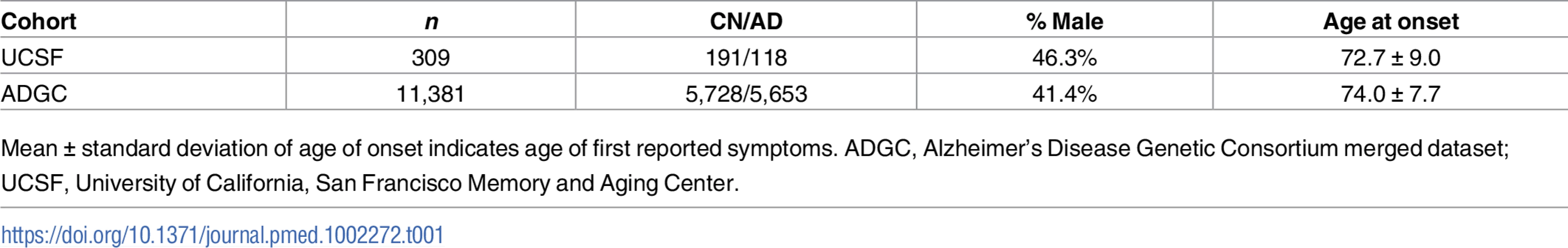 
Mean ± standard deviation of age of onset indicates age of first reported symptoms. ADGC, Alzheimer’s Disease Genetic Consortium merged dataset; UCSF, University of California, San Francisco Memory and Aging Center. Tab. 2. Five-allele haplotype risk associations in two clinical cohorts and combined dataset. 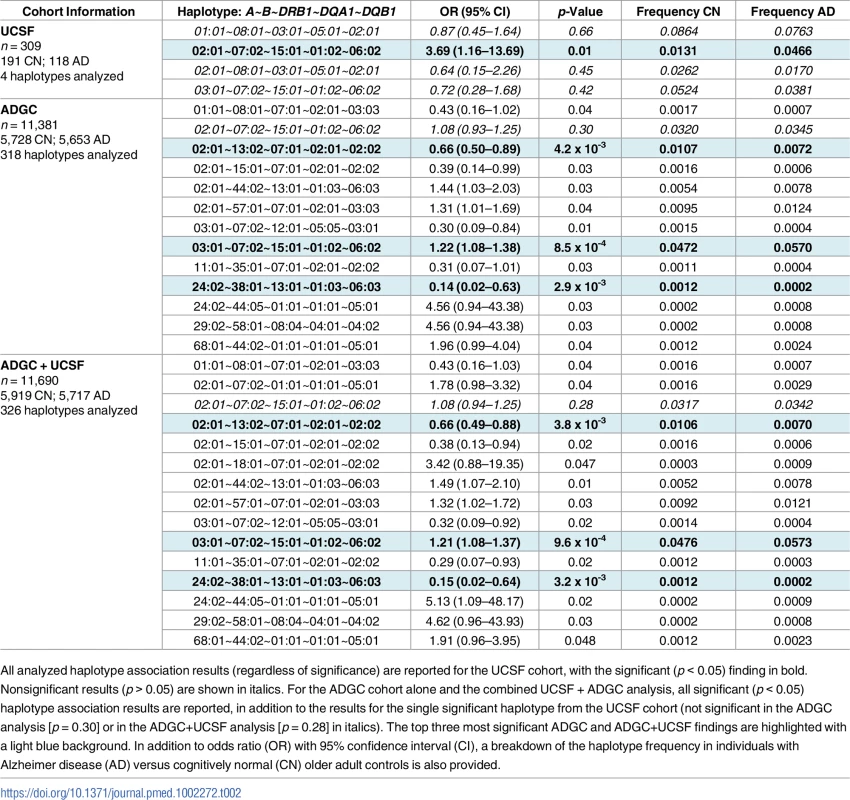
All analyzed haplotype association results (regardless of significance) are reported for the UCSF cohort, with the significant (p < 0.05) finding in bold. Nonsignificant results (p > 0.05) are shown in italics. For the ADGC cohort alone and the combined UCSF + ADGC analysis, all significant (p < 0.05) haplotype association results are reported, in addition to the results for the single significant haplotype from the UCSF cohort (not significant in the ADGC analysis [p = 0.30] or in the ADGC+UCSF analysis [p = 0.28] in italics). The top three most significant ADGC and ADGC+UCSF findings are highlighted with a light blue background. In addition to odds ratio (OR) with 95% confidence interval (CI), a breakdown of the haplotype frequency in individuals with Alzheimer disease (AD) versus cognitively normal (CN) older adult controls is also provided. After quality control, 11,381 individuals were available for analysis in the validation ADGC cohort (Table 1). Of the 318 haplotypes available for analysis, 12 five-allele haplotypes were significantly associated with AD risk (p < 0.05, Table 2). The strongest association was HLA A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 (OR = 1.22 [1.08–1.38], p = 8.5 x 10−4). This haplotype differed from the UCSF finding by one allele, at HLA-A. The third most significant haplotype association in the ADGC cohort was A*02:01~B*13:02~DRB1*07:01~DQA1*02:01~DQB1*02:02, which showed a protective effect, (OR = 0.66 [0.50–0.89], p = 4.2 x 10−3). This haplotype shared the HLA-A allele associated with AD risk in the UCSF discovery analysis. The full A*02:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype associated with AD in the UCSF cohort was not significant in the ADGC cohort (p = 0.30).
In combined analysis of both the UCSF and ADGC cohorts, 326 haplotypes were available for analysis (additional haplotypes beyond the 4 + 318 haplotypes analyzed in the separate UCSF and ADGC cohorts resulted when sufficient numbers of AD and CN controls for rare haplotypes became available in the combined UCSF + ADGC dataset). HLA A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 (OR = 1.21 [1.08–1.37), p = 9.6 x 10−4) and A*02:01~B*13:02~DRB1*07:01~DQA1*02:01~DQB1*02:02, [OR = 0.66 [0.49–0.88], p = 3.8 x 10−3] remained as two of the three most significant associations with AD (Table 2). Locus-level analyses of the combined cohort showed independent AD associations of B*07:02, DRB1*15:01, DQA1*01:02, and DQB1*06:02 (Table 3).
Tab. 3. Individual alleles with significant risk associations in combined cohort. 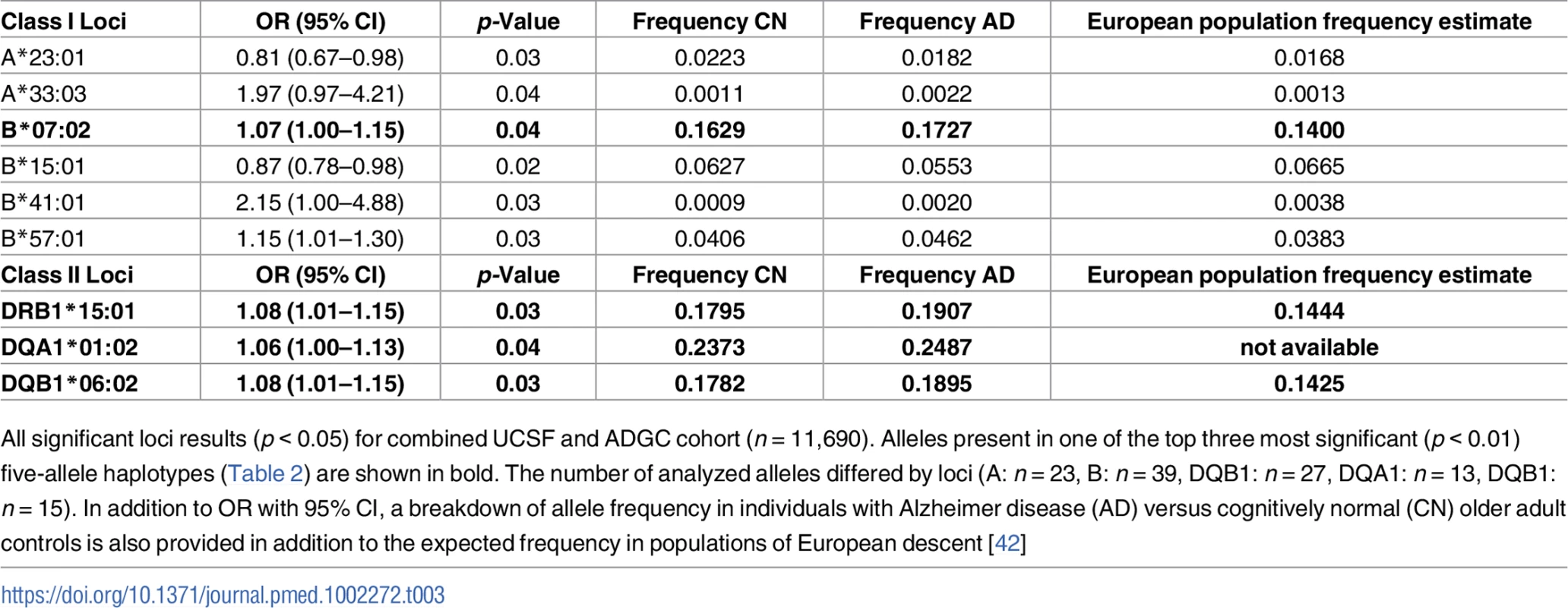
All significant loci results (p < 0.05) for combined UCSF and ADGC cohort (n = 11,690). Alleles present in one of the top three most significant (p < 0.01) five-allele haplotypes (Table 2) are shown in bold. The number of analyzed alleles differed by loci (A: n = 23, B: n = 39, DQB1: n = 27, DQA1: n = 13, DQB1: n = 15). In addition to OR with 95% CI, a breakdown of allele frequency in individuals with Alzheimer disease (AD) versus cognitively normal (CN) older adult controls is also provided in addition to the expected frequency in populations of European descent [42] Five-allele haplotype contributed to AD risk independently of APOE ɛ4 and may be driven by ɛ4-negative individuals
We next assessed whether the strong genetic AD risk factor APOE ɛ4 can account for the most significant five-allele haplotype association we identified in the combined UCSF+ADGC cohort. We recoded individuals as carriers or noncarriers of the A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype and assessed in a logistic regression framework the independent contributions of the risk haplotype, APOE ɛ4 carrier status, and whether there was an interaction between the two. As expected, APOE ɛ4 was strongly associated with AD risk (p < 2 x 10−16). The five-allele risk haplotype remained a significant contributor to AD risk (p = 0.036), but there was no statistically significant interaction between haplotype and APOE ɛ4 (p = 0.19). However, dividing the cohort by APOE ɛ4 carrier status showed that the frequency of the A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype was higher only in individuals with AD who are negative for ɛ4 (Table 4). Analysis of variance performed separately in ɛ4 carriers and noncarriers resulted in a significant association of the five-allele haplotype with AD only in ɛ4-negative individuals (p = 0.036 in ɛ4 noncarriers; p = 0.90 in ɛ4 carriers).
Tab. 4. AD/CN distribution by APOE ɛ4 and A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype carrier status. 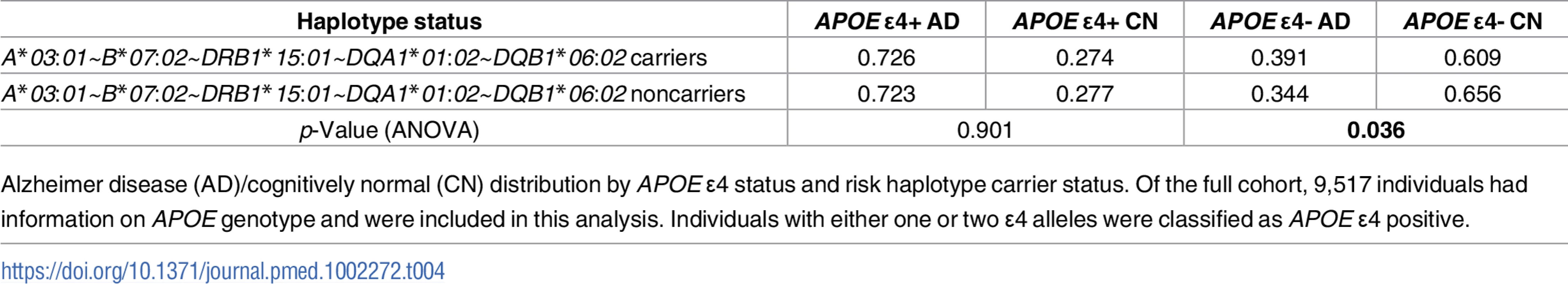
Alzheimer disease (AD)/cognitively normal (CN) distribution by APOE ɛ4 status and risk haplotype carrier status. Of the full cohort, 9,517 individuals had information on APOE genotype and were included in this analysis. Individuals with either one or two ɛ4 alleles were classified as APOE ɛ4 positive. Separate class I and class II haplotype analyses corroborated A*03:01~B*07:02 and DR15 in AD risk
Given the different roles of HLA receptors in recognizing endogenous (class I) or exogenous (class II) ligands, we also assessed class I (HLA A~B) and class II (HLA DRB1~DQA1~DQB1) haplotypes separately for their role in AD risk in the combined UCSF+ADGC cohort. Of 202 analyzed class I haplotypes, ten two-allele haplotypes were significantly associated with AD (p < 0.05), including A*03:01~B*07:02 (p = 0.03, OR = 1.1 [1.0–1.2]) (Table 5). Only one three-allele class II haplotype (out of 30 analyzed) was associated with AD risk, DR15 (p = 0.025, OR = 1.1 [1.0–1.2]). Together, these two separate haplotypes represent the most strongly associated five-allele haplotype identified in the combined analysis.
Tab. 5. Separate class I and class II haplotypes with significant risk associations in combined cohort. 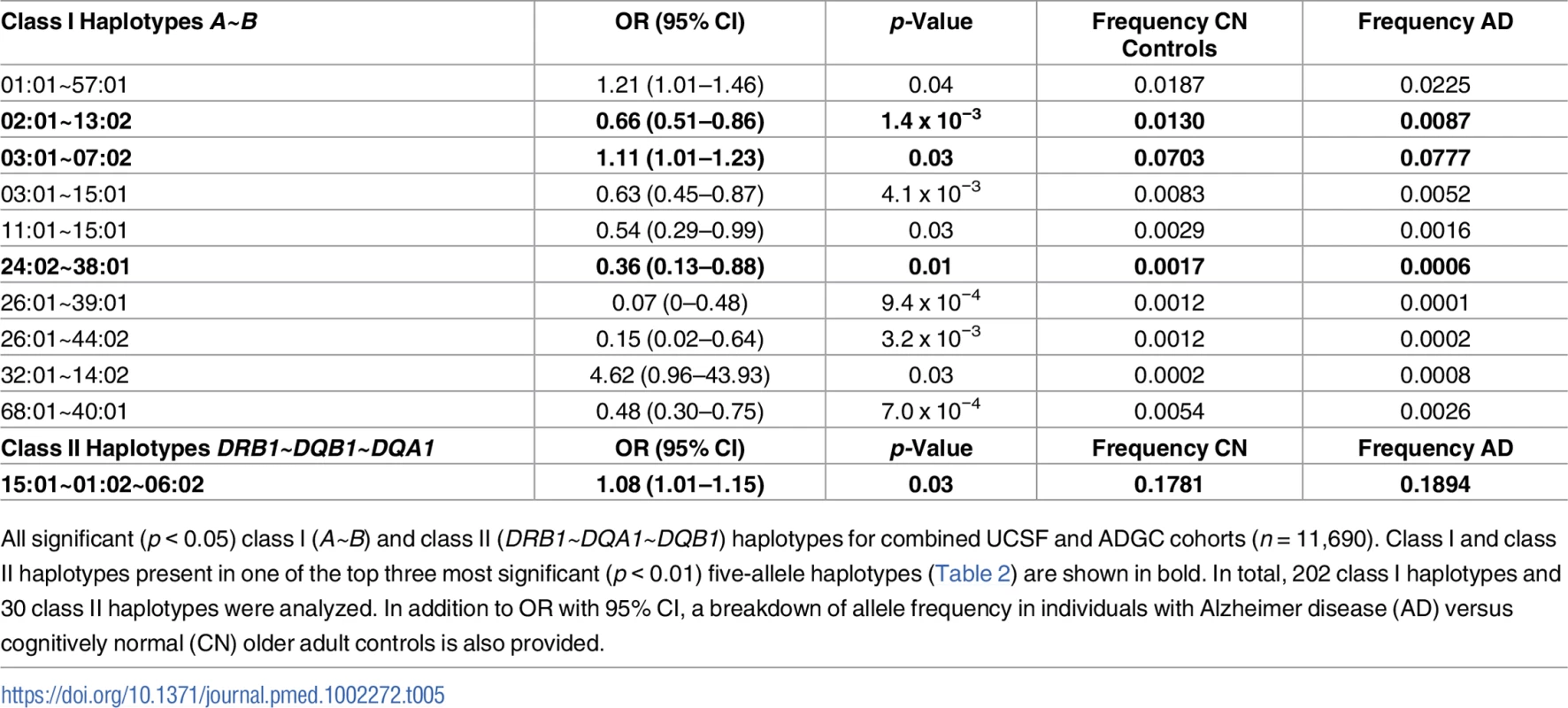
All significant (p < 0.05) class I (A~B) and class II (DRB1~DQA1~DQB1) haplotypes for combined UCSF and ADGC cohorts (n = 11,690). Class I and class II haplotypes present in one of the top three most significant (p < 0.01) five-allele haplotypes (Table 2) are shown in bold. In total, 202 class I haplotypes and 30 class II haplotypes were analyzed. In addition to OR with 95% CI, a breakdown of allele frequency in individuals with Alzheimer disease (AD) versus cognitively normal (CN) older adult controls is also provided. Class I haplotype A*03:01~B*07:02 was associated with baseline CSF amyloid levels
We utilized a subset of the ADNI cohort with genetic and cognitive data available to assess the disease-specific relevance of the class I A*03:01~B*07:02 and class II DR15 haplotypes across the AD spectrum, including cognitively normal controls, individuals with MCI, and those with AD. We analyzed the two haplotypes separately to assess whether class I and class II risk-associated haplotypes were correlated with similar or different clinical measures of AD. The cohort was balanced with respect to age, sex, and haplotype distributions (Table 6). The cohort was significantly different with respect to education and number of time points and showed expected differences in CDR-SB baseline score, APOE ɛ4 carrier status, ADAS baseline score, and RAVLT forgetting baseline score.
Tab. 6. Summary statistics for ADNI participants with longitudinal cognitive measures. 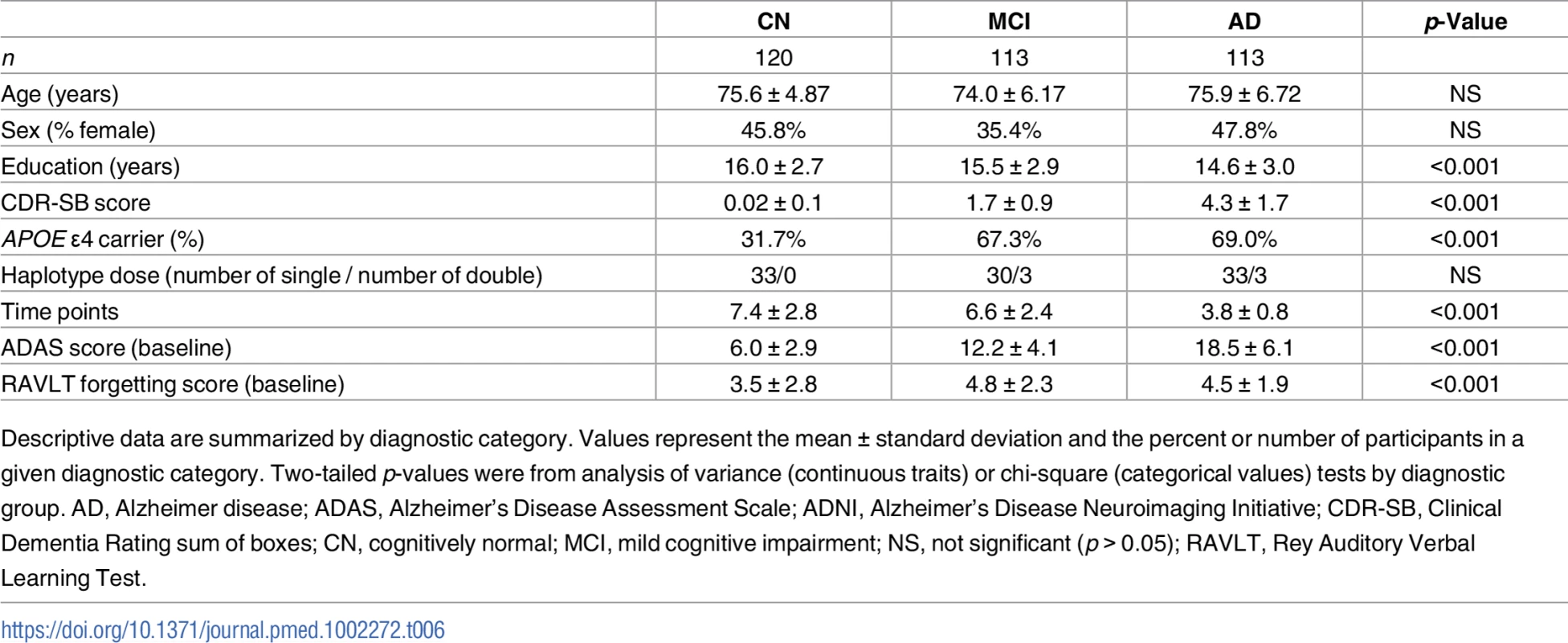
Descriptive data are summarized by diagnostic category. Values represent the mean ± standard deviation and the percent or number of participants in a given diagnostic category. Two-tailed p-values were from analysis of variance (continuous traits) or chi-square (categorical values) tests by diagnostic group. AD, Alzheimer disease; ADAS, Alzheimer’s Disease Assessment Scale; ADNI, Alzheimer’s Disease Neuroimaging Initiative; CDR-SB, Clinical Dementia Rating sum of boxes; CN, cognitively normal; MCI, mild cognitive impairment; NS, not significant (p > 0.05); RAVLT, Rey Auditory Verbal Learning Test. Carrying A*03:01~B*07:02 was associated with higher baseline levels of amyloid β as measured in CSF (Fig 1, p = 0.01). Traditionally, CSF amyloid levels are inversely correlated with amyloid burden in the brain; our results suggest that carrying A*03:01~B*07:02 is correlated with lower amyloid levels in the brain [43]. This is observed despite the fact that there were no statistically significant differences in baseline clinical or biomarker measures in patients with versus without the risk haplotype (S3 Table, S2 Fig). A*03:01~B*07:02 was not associated with any other baseline measures and was not associated with change in longitudinal measures over time.
Fig. 1. Carrying the A*03:01~B*07:02 risk haplotype was associated with CSF (cerebrospinal fluid) amyloid β. 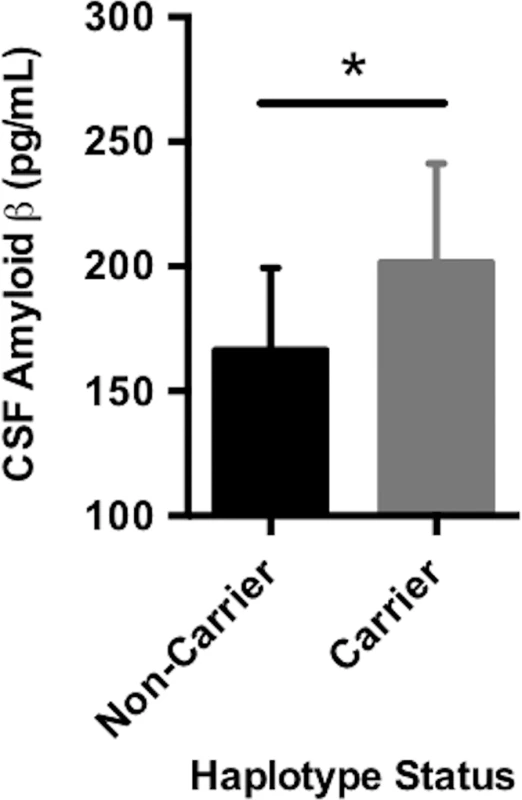
CSF amyloid β levels were on average higher in carriers of the A*03:01~B*07:02 haplotype, suggesting that haplotype carriers may have lower average intracranial amyloid pathological burden compared to noncarriers. The plotted points are best linear unbiased predictions from a multiple regression model, which controlled for age, sex, education, CDR-SB score, and APOE ɛ4 status. Data shown are the mean ± standard deviation (SD). DR15 risk haplotype correlated with worse cognitive decline and greater baseline inflammation across the AD spectrum
Longitudinal analysis of cognitive data identified a statistically significant association between the number of alleles of the DR15 risk haplotype and ADAS cognitive scores (p = 0.03), as well as with RAVLT forgetting scores (p = 0.02, S4 Table). The DR15 haplotype was associated with worse decline over time on both measures, corresponding to increasing longitudinal ADAS cognitive scores and decreasing longitudinal RAVLT forgetting scores over time (shown relative to noncarriers in Figs 2 and 3). DR15-associated changes in cognitive trajectory occurred despite the fact that there were no baseline differences in clinical severity or cognitive function in patients with AD based on DR15 carrier status (S5 Table). In addition, baseline biomarker measures most relevant to AD were similar in both patients with AD who are DR15 carriers and those who are noncarriers (S5 Table, S3 Fig), indicating that all patients had equivalent baseline disease severity.
Fig. 2. DR15 haplotype carriers showed greater change over time on the ADAS cognitive assessment when compared to noncarriers. 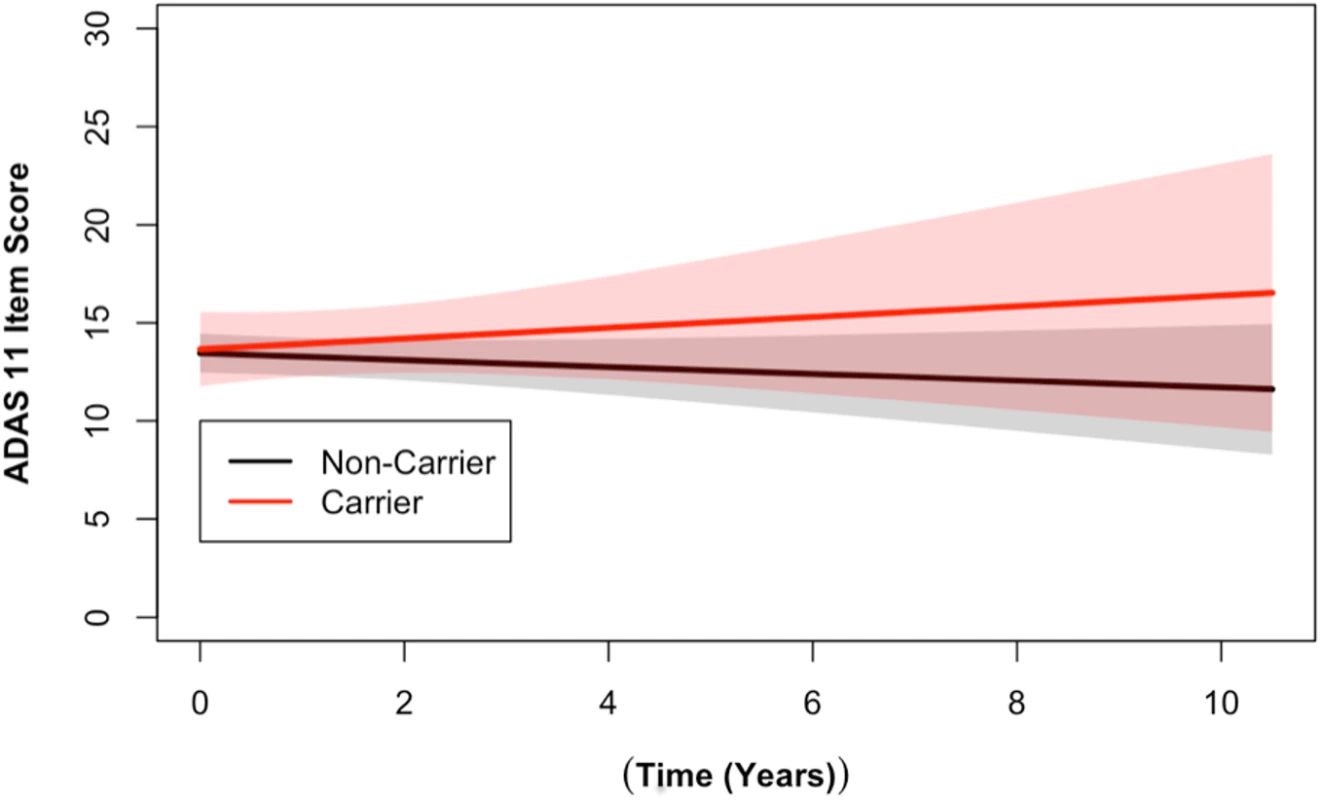
Longitudinal ADAS 11-item cognitive subscale scores from the ADNI cohort are shown. The ADAS broadly measures cognitive functions impaired in AD [34], with higher scores representing more cognitive impairment. DR15 haplotype carriers (in red) showed worse cognitive function over time when compared to noncarriers (in black) (p = 0.03). The plotted data represent the best linear unbiased prediction results from the regression model specified (see Methods) with 95% CIs (shaded regions). Fig. 3. DR15 haplotype carriers declined more on the RAVLT forgetting score when compared to noncarriers. 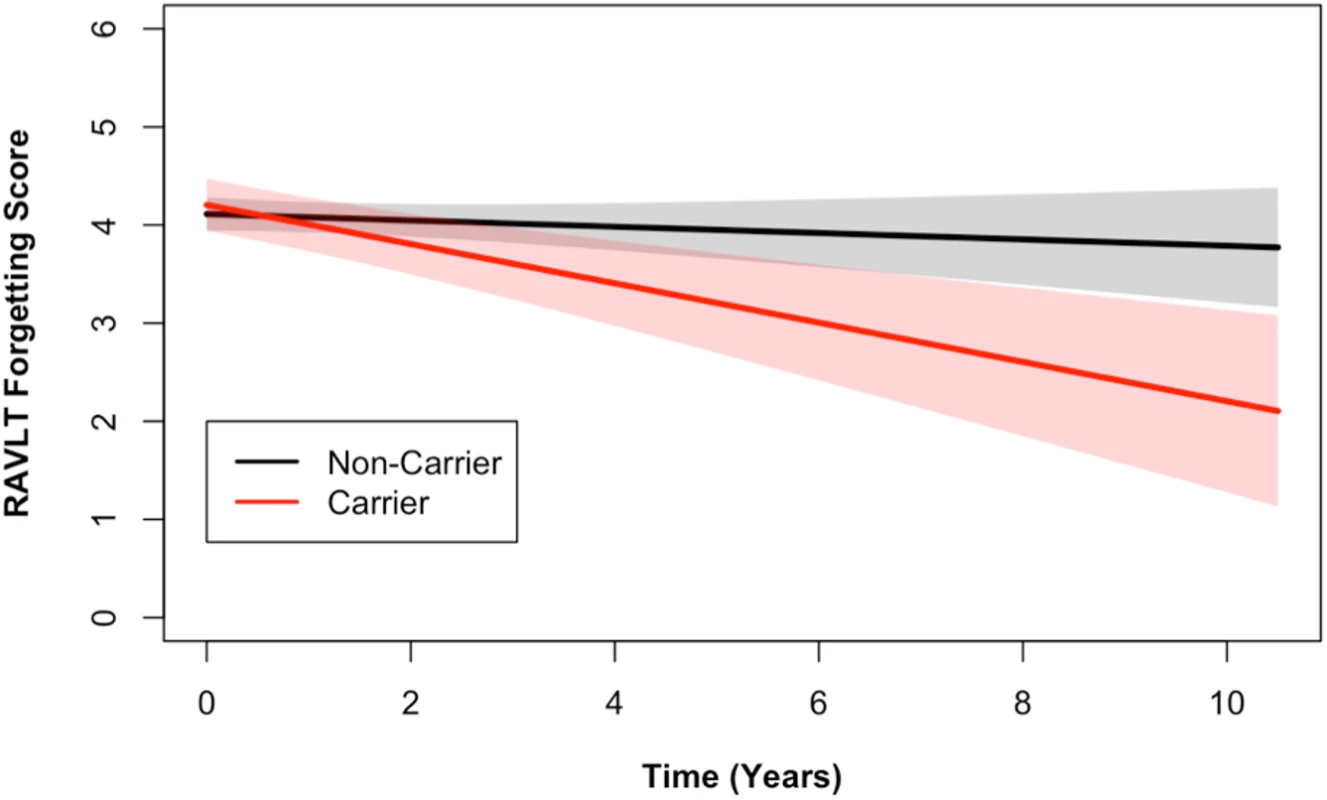
Longitudinal RAVLT measurements from the ADNI cohort are shown. The RAVLT forgetting score is defined as the difference between the delayed recall and immediate recall scores on the RAVLT and represents a measure of memory consolidation. Over time, DR15 risk haplotype carriers showed more change on the forgetting score (i.e., more forgetting) than noncarriers. The plotted data represent the best linear unbiased prediction results from the regression model specified (see Methods) with 95% CIs (shaded regions). In a subset of individuals who also had baseline CSF data available (S2 Table), we tested whether the DR15 risk haplotype altered any biomarker measures of immunological function and inflammation. We tested 28 analyte levels related to immune function and inflammation (S1 List). At baseline, there was an association between chemokine CC-4 (CC4) and age (p = 0.02, S4 Fig), as well as CC4 with dose of DR15 risk haplotype (p = 5.18 x 10−3, Fig 4, S6 Table). Although not reaching strict statistical significance after adjustment for the 28 biomarkers tested (at Bonferroni adjusted p < 1.79 x 10−3), this analysis provides suggestive biomarker evidence of heightened baseline inflammation in individuals carrying the DR15 risk haplotype.
Fig. 4. DR15 dosage was associated with higher baseline levels of chemokine CC4. 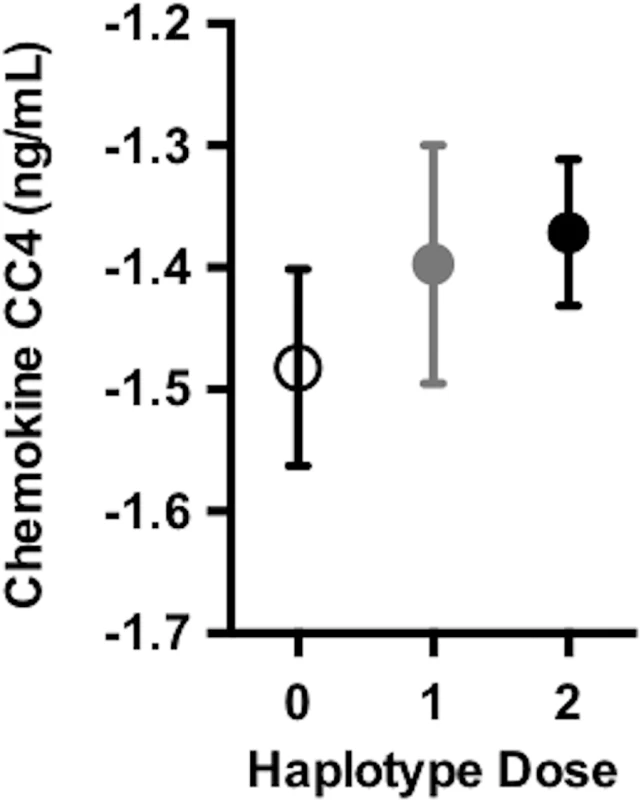
As the number of DR15 risk haplotype alleles increases, there were higher average levels of chemokine CC4, suggesting higher levels of inflammation at baseline. Chemokine CC4 levels are quality controlled and transformed as described in S1 Methods. The plotted points are partial residuals with 95% confidence bands provided in shading. HLA haplotype risk effects differed by sex
Given previous reports of greater risk effects of DR15 in female patients with multiple sclerosis (MS) [44] and the stronger effect of APOE ɛ4 in females [45], we assessed whether men versus women showed similar or different HLA haplotype associations with AD risk. When split by sex, two of the three most significant five-allele haplotypes from the combined sex analysis were significant in an individual sex. The five-allele haplotype A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 was significant only in men (OR 1.31 [1.09–1.58], p = 0.0035) (S7 Table). However, A*02:01~B*13:02~DRB1*07:01~DQA1*02:01~DQB1*02:02 was significant only in women (OR 0.68 [0.46–0.99], p = 0.034) (S7 Table). Similar findings appeared in separate class I and class II haplotype analyses. Only men showed significant associations with class I haplotype A*03:01~B*07:02 (p = 0.027), and only women showed significant associations with A*02:01~B*13:02 (p = 0.0049) (S8 Table). Finally, class II haplotype DR15 was only significantly associated with AD risk in men (p = 0.01) (S8 Table). Locus-level analyses were consistent, with only men showing significant associations with ten alleles, including B*07:02 (p = 0.013), DRB1*15:01 (p = 0.0096), DQA1*01:02 (p = 0.029), and DQB1*06:02 (p = 0.01) (S9 Table). There were four individual alleles associated with AD risk in women, none of which were components of any of the top three significant five-allele haplotypes in the combined sex analysis (S9 Table).
Iterative subanalyses corroborate role of HLA-A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 in AD
To attempt to alleviate concern over possible Type I error in this analysis, we randomly split the combined ADGC+UCSF cohort ten times (maintaining the same proportion of AD:controls) and reran the five-allele haplotype analysis in the 20 resulting (smaller) cohorts. Two of the top-associated five-allele haplotypes showed p-values < 0.05 in over half of the randomly split analyses (S10 Table), which was more than any of the other “top” haplotypes from the original analysis. This included the one we focused on in this study (A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02), which showed significance in 11 iterations of the randomly split analysis, with p-values from 0.026–0.0001 and ORs of 1.21–1.40, further corroborating the contributions of this haplotype and its subcomponents to AD risk.
Discussion
In a total of over 11,000 individuals, we found evidence suggesting that the five-allele HLA haplotype A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 is a risk factor for AD and that this effect may be driven by men who do not carry the major AD risk factor, APOE ɛ4. Locus-level analysis further confirmed AD associations of the individual alleles B*07:02, DRB1*15:01, DQA1*01:02, and DQB1*06:02. In separate class I and class II haplotype analyses, the class I A*03:01~B*07:02 haplotype and the class II DRB1*15:01~DQA1*01:02~DQB1*06:02 (DR15) haplotype were both significantly associated with risk for AD. We assessed the clinical relevance of each of these haplotypes separately in a smaller cohort representing the spectrum of cognitively normal controls and individuals with MCI and AD. Carrying the MHC class I haplotype A*03:01~B*07:02 was associated with higher CSF amyloid levels, suggesting lower levels of amyloid in the brains of haplotype carriers across the AD spectrum. The class II haplotype DR15 was associated with greater rate of decline on two different measures of cognitive function relevant to AD in a dose-dependent manner. In a subset of the same cohort, carrying the DR15 risk haplotype was also associated with higher baseline levels of CC4, a biomarker of AD-related inflammation [46]. Taking these findings together, this study provides evidence for the contribution of the A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype and its components, A*03:01~B*07:02 and DR15, to risk of AD.
Over 30 y of research into HLA alleles and risk of AD has yielded mixed conclusions due in part to limitations in mapping alleles within this complicated genomic region. Early studies mapped risk of AD to the HLA region of chromosome 6 [47], and the studies that followed differed significantly in their methodological approach, the identities and resolution of the alleles studied, the ethnicity of the study cohorts, and the inferences drawn from the data. MHC class I molecule HLA-A*02 has been shown to either be associated with increased risk of AD or to have no effect in nearly 15 different studies [48–61]. Given that only B*07:02, DRB1*15:01, DQA1*01:02, and DQB1*06:02 showed significant locus-level associations with AD, our findings are consistent with an ambiguous role of HLA-A*02 in AD. In terms of class II alleles, one study by Mansouri and colleagues demonstrated a link between DRB1*15:01~DQB1*06:02 and AD in a small cohort of Tunisians [62], consistent with our findings. Previous GWASs have found that AD risk is associated with a SNP in DRB5 [63]. As there is strong LD between DRB5*01 and DRB1*15:01~DQB1*06:02 [64], it is possible that the AD association we have detected with DR15 is due in part or wholly to DRB5. Finally, our finding that HLA associations with AD are stronger in APOE ɛ4-negative individuals is consistent with prior work for different HLA alleles [65,66].
The HLA region has been studied to a varying extent for its contributions to neurological disease, and many of the risk alleles implicated in the present studies have also been linked to other disorders. Most notably, the class II DR15 haplotype is the most consistently replicated genetic finding in MS [67–69]. DR15 also correlates with worse clinical progression in women with relapsing-onset MS (e.g., younger age at onset and more subcortical atrophy) [44]. Class I allele B*07 has also been associated with MS risk, particularly in those also carrying DRB1*15 [70]. In one Parkinson disease (PD) study, four alleles identified in a risk haplotype overlapped with our top five-allele haplotype association [71]. Similar to AD, other studies have also implicated the HLA-DRB5 region in PD risk [72]. In one small autism study, the class I allele B*07 and class II allele DQB1*06:02 were both associated with disease risk [73]. Finally, class II allele DQB1*06:02 has been associated with marked increased risk for [74,75], and worsened severity of [76,77], narcolepsy. These findings are consistent with our study, in which we identified a dose-dependent association between DR15 and greater cognitive decline in individuals representing the AD spectrum.
Participants who carried at least one copy of the class I haplotype A*03:01~B*07:02 on average had higher baseline CSF amyloid levels, suggesting lower amyloid burden in the brains of these participants. Similar findings have been observed in APOE ɛ4-negative individuals when compared to APOE ɛ4-positive individuals across the phenotypic spectrum of cognitively normal to early MCI, suggesting higher brain amyloid β in carriers [78]. This finding raises the possibility that there could be a tau-mediated effect on AD clinical symptoms, as the AD group did not differ in clinical measures by haplotype carrier status. On the other hand, DR15 haplotype carriers demonstrated subtle differences in baseline inflammatory biomarker levels, as well as a worse cognitive trajectory over time, suggesting a disease-modifying effect that could be mediated by changes in immune function.
Our study benefited from several strengths. The primary discovery cohort was a well-characterized sample of patients who received extensive clinical evaluations at the UCSF Memory and Aging Center. The replication dataset from the ADGC of over 11,000 AD and cognitively normal control individuals is the largest dataset to date used to explore immunogenetic contributions to AD risk. Lastly, longitudinal data from ADNI allowed us to probe the potential clinical relevance of haplotype findings across the AD spectrum. Our study also has caveats that are important to consider. Our imputation program predicted accuracy for individuals with European ancestry that was likely higher than 94.8%–99.2% based on our more stringent call threshold in comparison to other studies [38]. Imputed HLA alleles have been shown to be reliable classification tools in studies with similar methodologies [38,79,80]; however, future studies would benefit from direct sequencing of HLA alleles to avoid potential imputation inaccuracies. Because of limitations in the imputation package selected for HLA allele calling, we were only able to impute genotypes for a subset of MHC class I and II genes. For example, the imputed genes available did not include DRB5, which was indicated in previous studies to be associated with AD risk or pathological processes [9,24,81]. DRB5 is on the DR15 haplotype, so it is likely that the association we identified reflects these previous results. However, we can neither directly confirm nor refute this possibility in the present study. The DR15 risk haplotype is most common in Europeans, and to minimize genetic heterogeneity in population substructure, we limited the present analysis to white individuals of non-Hispanic descent. Additional studies are required to assess the identified HLA risk haplotypes and component alleles for their contribution to AD in more diverse populations where patterns of LD differ and may uncouple alleles that were tightly linked in our study population, though the initial study identifying class II associations with AD in Tunisians suggests this may be a generalized risk phenomenon. Although p < 0.05 may be considered lenient based on the number of total alleles tested, it is also true that all of these alleles represent only five genes within one genomic region that has been previously linked to AD risk. Despite reduced statistical power due to low frequency of HLA haplotypes imparted by the extraordinary diversity of this region, we feel that this study is an important first step in elucidating the underlying contribution of the HLA to AD risk given the medical implications of ultimately identifying immune-related therapies as a means of modifying a complex, common disease. We have greater confidence in our findings due to corroborating clinical validity as identified in the ADNI cohort. Iterative subanalyses of the combined study cohort further support a role of our top five-allele haplotypes in AD risk. In addition, two of the main alleles of interest we identified, DRB1*15:01 and DQB1*06:02, have been linked to AD risk in two prior studies, further supporting our results. We also identified several other risk haplotypes in our analyses beyond the ones we focused on in this study; the clinical relevance of these additional haplotypes and alleles requires further investigation. Future work is also required to test whether these findings extend to early-onset and atypical clinical syndromes with underlying AD pathology.
In summary, we present evidence for a role of the HLA class I A*03:01~B*07:02 haplotype and the HLA class II DR15 haplotype in AD risk. Our study also suggests that these risk haplotypes may be associated with CSF AD biomarker levels (class I) and greater decline in cognition over time, as well as higher levels of inflammation across aging (class II). The results of our study indicate that the broad A*03:01~B*07:02~DRB1*15:01~DQA1*01:02~DQB1*06:02 haplotype may contribute genetic risk to AD beyond that contributed by the established risk factor APOE ɛ4, particularly in men. As components of this haplotype are well-established risk factors in MS, PD, autism, and narcolepsy, we propose that they may contribute to underlying biological risk mechanisms in multiple neurological diseases. Future work is required to establish the precise molecular processes underlying this risk association, as well as to expand this finding to broader, diverse populations of AD and potentially even other neurodegenerative conditions.
Supporting Information
Zdroje
1. Prince M, Wimo A, Guerchet M, Ali G-C, Wu Y-T, Prina M, et al. World Alzheimer Report 2015. The global impact of dementia: An analysis of prevalence, incidence, cost and trends. 2015. https://www.alz.co.uk/research/world-report-2015
2. Zetzsche T, Rujescu D, Hardy J, Hampel H. Advances and perspectives from genetic research: development of biological markers in Alzheimer’s disease. Expert Rev Mol Diagn. 2010;10 : 667–690. doi: 10.1586/erm.10.48 20629514
3. Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E, et al. TREM2 Variants in Alzheimer’s Disease. N Engl J Med. 2013;368(2): 117–127. doi: 10.1056/NEJMoa1211851 23150934
4. Jonsson T, Stefansson H, Steinberg S, Jonsdottir I, Jonsson P, Snaedal J, et al. Variant of TREM2 Associated with the Risk of Alzheimer’s Disease. N Engl J Med. 2013;368(2): 107–116. doi: 10.1056/NEJMoa1211103 23150908
5. Gonzalez Murcia JD, Schmutz C, Munger C, Perkes A, Gustin A, Peterson M, et al. Assessment Of Trem2 Rs75932628 Association With Alzheimer’s Disease In A Population-Based Sample: The Cache County Study. Neurobiol Aging. 2013;34 : 2889.e11–2889.e13.
6. Sirkis DW, Bonham LW, Aparicio RE, Geier EG, Ramos EM, Wang Q, et al. Rare TREM2 variants associated with Alzheimer’s disease display reduced cell surface expression. Acta Neuropathol Commun. 2016;4(1):98. doi: 10.1186/s40478-016-0367-7 27589997
7. Benitez BA, Cooper B, Pastor P, Jin SC, Lorenzo E, Cervantes S, et al. TREM2 is associated with risk of Alzheimer disease in Spanish population. Neurobiol Aging. 2013;34 : 1711.e15–1711.e17.
8. Coppola G, Chinnathambi S, Lee JJ, Dombroski BA, Baker MC, Soto-Ortolaza AI, et al. Evidence for a role of the rare p.A152T variant in MAPT in increasing the risk for FTD-spectrum and Alzheimer’s diseases. Hum Mol Genet. 2012;21 : 3500–3512. doi: 10.1093/hmg/dds161 22556362
9. Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45 : 1452–8. doi: 10.1038/ng.2802 24162737
10. Naj AC, Jun G, Beecham GW, Wang L, Vardarajan BN, Buros J, et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat Genet. 2011;43 : 436–441. doi: 10.1038/ng.801 21460841
11. Karch CM, Goate AM. Alzheimer’s disease risk genes and mechanisms of disease pathogenesis. Biol Psychiatry. 2015;77 : 43–51. doi: 10.1016/j.biopsych.2014.05.006 24951455
12. Hensley K. Neuroinflammation in Alzheimer’s disease: mechanisms, pathologic consequences, and potential for therapeutic manipulation. J Alzheimers Dis. 2010;21 : 1–14. doi: 10.3233/JAD-2010-1414 20182045
13. Wes PD, Holtman IR, Boddeke EWGM, Möller T, Eggen BJL. Next generation transcriptomics and genomics elucidate biological complexity of microglia in health and disease. Glia. 2016;64 : 197–213. doi: 10.1002/glia.22866 26040959
14. Gan L, Ye S, Chu A, Anton K, Yi S, Vincent VA, et al. Identification of Cathepsin B as a Mediator of Neuronal Death Induced by Aβ-activated Microglial Cells Using a Functional Genomics Approach. J Biol Chem. 2004;279 : 5565–5572. doi: 10.1074/jbc.M306183200 14612454
15. Hong S, Beja-Glasser VF, Nfonoyim BM, Frouin A, Li S, Ramakrishnan S, et al. Complement and microglia mediate early synapse loss in Alzheimer mouse models. Science. 2016;352 : 712–6. doi: 10.1126/science.aad8373 27033548
16. Eikelenboom P, Hoozemans JJ, Veerhuis R, van Exel E, Rozemuller AJ, van Gool WA. Whether, when and how chronic inflammation increases the risk of developing late-onset Alzheimer’s disease. Alzheimers Res Ther. 2012;4 : 15. doi: 10.1186/alzrt118 22647384
17. Schmidt R, Schmidt H, Curb JD, Masaki K, White LR, Launer LJ. Early inflammation and dementia: A 25-year follow-up of the Honolulu-Asia Aging Study. Ann Neurol. 2002;52 : 168–174. doi: 10.1002/ana.10265 12210786
18. Mrak RE, Griffin WST. Potential inflammatory biomarkers in Alzheimer’s disease. J Alzheimers Dis. 2005;8 : 369–75. 16556968
19. Wood JA, Wood PL, Ryan R, Graff-Radford NR, Pilapil C, Robitaille Y, et al. Cytokine indices in Alzheimer’s temporal cortex: no changes in mature IL-1 beta or IL-1RA but increases in the associated acute phase proteins IL-6, alpha 2-macroglobulin and C-reactive protein. Brain Res. 1993;629 : 245–52. 7509248
20. Shi Q, Colodner KJ, Matousek SB, Merry K, Hong X, Kenison JE, et al. Complement C3-Deficient Mice Fail to Display Age-Related Hippocampal Decline. J Neurosci. 2015;35 : 13029–13042. doi: 10.1523/JNEUROSCI.1698-15.2015 26400934
21. Stephan AH, Madison D V, Mateos JM, Fraser DA, Lovelett EA, Coutellier L, et al. A Dramatic Increase of C1q Protein in the CNS during Normal Aging. J Neurosci. 2013;33 : 13460–13474. doi: 10.1523/JNEUROSCI.1333-13.2013 23946404
22. Dorman JS, Bunker CH. HLA-DQ locus of the human leukocyte antigen complex and type 1 diabetes mellitus: a HuGE review. Epidemiol Rev. 2000;22 : 218–27. 11218373
23. Allen M, Kachadoorian M, Carrasquillo MM, Karhade A, Manly L, Burgess JD, et al. Late-onset Alzheimer disease risk variants mark brain regulatory loci. Neurol Genet. 2015;1: e15. doi: 10.1212/NXG.0000000000000012 27066552
24. Yokoyama JS, Wang Y, Schork AJ, Thompson WK, Karch CM, Cruchaga C, et al. Association Between Genetic Traits for Immune-Mediated Diseases and Alzheimer Disease. JAMA Neurol. 2016;94158 : 1–7.
25. Dubois B, Feldman H, Jacova C. Advancing research diagnostic criteria for Alzheimer’s disease: the IWG-2 criteria. Lancet Neurol. 2014;13 : 614–29. doi: 10.1016/S1474-4422(14)70090-0 24849862
26. Boehme KL, Young B, Mukherjee S, Crane PK, Kauwe J, Young B. ADGC 1000 Genomes combined data workflow. 2014; 1–12. http://kauwelab.byu.edu/Portals/22/adgc_combined_1000G_12032014.pdf
27. Bonham LW, Desikan RS, Yokoyama JS. The relationship between complement factor C3, APOE ε4, amyloid and tau in Alzheimer’s disease. Acta Neuropathol Commun. 2016;4 : 1–7.
28. Desikan RS, Thompson WK, Holland D, Hess CP, Brewer JB, Zetterberg H, et al. The role of clusterin in amyloid-β-associated neurodegeneration. JAMA Neurol. 2014;71 : 180–7. doi: 10.1001/jamaneurol.2013.4560 24378367
29. Desikan RS, Thompson WK, Holland D, Hess CP, Brewer JB, Zetterberg H, et al. Heart fatty acid binding protein and Aβ-associated Alzheimer’s neurodegeneration. Mol Neurodegener. 2013;8 : 39. doi: 10.1186/1750-1326-8-39 24088526
30. Morris JC. The Clinical Dementia Rating (CDR): current version and scoring rules. Neurology. 1993;43 : 2412–2414.
31. Saykin AJ, Shen L, Foroud TM, Potkin SG, Swaminathan S, Kim S, et al. Alzheimer’s Disease Neuroimaging Initiative biomarkers as quantitative phenotypes: Genetics core aims, progress, and plans. Alzheimers Dement. 2010;6(3): 265–273. doi: 10.1016/j.jalz.2010.03.013 20451875
32. Shaw LM, Vanderstichele H, Knapik-Czajka M, Clark CM, Aisen PS, Petersen RC, et al. Cerebrospinal fluid biomarker signature in alzheimer’s disease neuroimaging initiative subjects. Ann Neurol. 2009;65 : 403–413. doi: 10.1002/ana.21610 19296504
33. Rey A. L’examen clinique en psychologie. Lexamen clinique en psychologie. 1958.
34. Rosen WG, Mohs RC, Davis KL. A new rating scale for Alzheimer’s disease. Am J Psychiatry. 1984;141(11): 1356–1364. doi: 10.1176/ajp.141.11.1356 6496779
35. Mohs RC CL. Alzheimer’s Disease Assessment Scale (ADAS). Psychopharmacol Bull. 1988;24 : 627–8. 3249763
36. Hughes CP, Berg L, Danziger WL. A new clinical scale for the staging of dementia. Br J Psychiatry. 1982;140 : 566–572. 7104545
37. Khor S-S, Yang W, Kawashima M, Kamitsuji S, Zheng X, Nishida N, et al. High-accuracy imputation for HLA class I and II genes based on high-resolution SNP data of population-specific references. Pharmacogenomics J. 2015;15 : 530–7. doi: 10.1038/tpj.2015.4 25707395
38. Zheng X, Shen J, Cox C, Wakefield JC, Ehm MG, Nelson MR, et al. HIBAG—HLA genotype imputation with attribute bagging. Pharmacogenomics J. 2014;14 : 192–200. doi: 10.1038/tpj.2013.18 23712092
39. Pappas DJ, Marin W, Hollenbach JA, Mack SJ. Bridging ImmunoGenomic Data Analysis Workflow Gaps (BIGDAWG): An integrated case-control analysis pipeline. Hum Immunol. 2016;77 : 283–7. doi: 10.1016/j.humimm.2015.12.006 26708359
40. Van Deerlin VM, Sleiman PM, Martinez-Lage M, Chen-Plotkin A, Wang L-S, Graff-Radford NR, et al. Common variants at 7p21 are associated with frontotemporal lobar degeneration with TDP-43 inclusions. Nat Genet. 2010;42 : 234–9. doi: 10.1038/ng.536 20154673
41. Hollenbach JA, Mack SJ, Thomson G, Gourraud PA. Analytical methods for disease association studies with immunogenetic data. Methods Mol Biol. 2012;882 : 245–266. doi: 10.1007/978-1-61779-842-9_14 22665238
42. Maiers M., Gragert L., Klitz W. High resolution HLA alleles and haplotypes in the US population. Human Immunology (2007) 68, 779–788. doi: 10.1016/j.humimm.2007.04.005 17869653
43. Tapiola T, Pirttilä T, Mehta PD, Alafuzof I, Lehtovirta M, Soininen H. Relationship between apoE genotype and CSF β-amyloid (1–42) and tau in patients with probable and definite Alzheimer’s disease. Neurobiol Aging. 2000;21 : 735–740. 11016543
44. Isobe N, Keshavan A, Gourraud P-A, Zhu AH, Datta E, Schlaeger R, et al. Association of HLA Genetic Risk Burden With Disease Phenotypes in Multiple Sclerosis. JAMA Neurol. 2016;73 : 795–802. doi: 10.1001/jamaneurol.2016.0980 27244296
45. Farrer LA, Cupples LA, Haines JL, Hyman B, Kukull WA, Mayeux R, et al. Effects of Age, Sex, and Ethnicity on the Association Between Apolipoprotein E Genotype and Alzheimer Disease: A Meta-analysis. JAMA. 1997;278(16): 1349–1356. 9343467
46. Kessler H, Pajonk F-G, Meisser P, Schneider-Axmann T, Hoffmann K-H, Supprian T, et al. Cerebrospinal fluid diagnostic markers correlate with lower plasma copper and ceruloplasmin in patients with Alzheimer’s disease. J Neural Transm. 2006;113 : 1763–9. doi: 10.1007/s00702-006-0485-7 16736242
47. Weitkamp LR, Nee L, Keats B, Polinsky RJ, Guttormsen S. Alzheimer disease: evidence for susceptibility loci on chromosomes 6 and 14. Am J Hum Genet. 1983;35 : 443–53. 6859040
48. Payami H, Schellenberg GD, Zareparsi S, Kaye J, Sexton GJ, Head MA, et al. Evidence for association of HLA-A2 allele with onset age of Alzheimer’s disease. Neurology. 1997;49 : 512–8. 9270587
49. Payami H, Kaye J, Becker W, Norman D, Wetzsteon P. HLA-A2, or a closely linked gene, confers susceptibility to early-onset sporadic Alzheimer’s disease in men. Neurology. 1991;41 : 1544–8. 1922793
50. Combarros O, Escribano J, Sánchez-Velasco P, Leyva-Cobián F, Oterino A, Leno C, et al. Association of the HLA-A2 allele with an earlier age of onset of Alzheimer’s disease. Acta Neurol Scand. 1998;98 : 140–1. 9724015
51. Guerini FR, Calabrese E, Agliardi C, Zanzottera M, Franceschi M, Grimaldi LM, et al. Association study of the HLA-A2 allele in Italian Alzheimer disease patients. Neurobiol Aging. 2009;30 : 2082–2083. doi: 10.1016/j.neurobiolaging.2008.02.001 18359130
52. Listì F, Candore G, Balistreri CR, Grimaldi MP, Orlando V, Vasto S, et al. Association between the HLA-A2 allele and Alzheimer disease. Rejuvenation Res. 2006;9 : 99–101. doi: 10.1089/rej.2006.9.99 16608404
53. Ma SL, Tang NLS, Tam CWC, Lui VWC, Suen EWC, Chiu HFK, et al. Association between HLA-A alleles and Alzheimer’s disease in a southern Chinese community. Dement Geriatr Cogn Disord. 2008;26 : 391–7. doi: 10.1159/000164275 18936542
54. Araria-Goumidi L, Lambert JC, Cottel D, Amouyel P, Chartier-Harlin MC. No association of the HLA-A2 allele with Alzheimer’s disease. Neurosci Lett. 2002;335 : 75–8. 12459502
55. Zareparsi S, James DM, Kaye JA, Bird TD, Schellenberg GD, Payami H. HLA-A2 homozygosity but not heterozygosity is associated with Alzheimer disease. Neurology. 2002;58 : 973–5. 11914421
56. Harris JM, Cumming AM, Craddock N, St Clair D, Lendon CL. Human leucocyte antigen-A2 increases risk of Alzheimer’s disease but does not affect age of onset in a Scottish population. Neurosci Lett. 2000;294 : 37–40. 11044581
57. Small GW, Scott WK, Komo S, Yamaoka LH, Farrer LA, Auerbach SH, et al. No association between the HLA-A2 allele and Alzheimer disease. Neurogenetics. 1999;2 : 177–82. 10541592
58. Small GW, Ebeling SC, Matsuyama SS, Heyman A, Reisner EG, Renvoize EB, et al. Variable association of HLA-A2 in men with early-onset Alzheimer disease. Neurobiol Aging. 1991;12 : 375–7. 1961374
59. Ballerini C, Nacmias B, Rombolà G, Marcon G, Massacesi L, Sorbi S. HLA A2 allele is associated with age at onset of Alzheimer’s disease. Ann Neurol. 1999;45 : 397–400. 10072057
60. Guerini FR, Tinelli C, Calabrese E, Agliardi C, Zanzottera M, De Silvestri A, et al. HLA-A*01 is associated with late onset of Alzheimer’s disease in Italian patients. Int J Immunopathol Pharmacol. 2009;22 : 991–9. doi: 10.1177/039463200902200414 20074462
61. Middleton D, Mawhinney H, Curran MD, Edwardson JA, Perry R, McKeith I, et al. Frequency of HLA-A and B alleles in early and late-onset Alzheimer’s disease. Neurosci Lett. 1999;262 : 140–2. 10203251
62. Mansouri L, Messalmani M, Klai S, Bedoui I, Derbali H, Gritli N, et al. Association of HLA-DR/DQ polymorphism with Alzheimer’s disease. Am J Med Sci. 2015;349 : 334–7. doi: 10.1097/MAJ.0000000000000416 25651370
63. Yokoyama JS, Wang Y, Schork AJ, Thompson WK, Karch CM, Cruchaga C, et al. Association between genetic traits for immune-mediated diseases and Alzheimer disease. JAMA Neurol. 2016;73.
64. Miretti MM, Walsh EC, Ke X, Delgado M, Griffiths M, Hunt S, et al. A High-Resolution Linkage-Disequilibrium Map of the Human Major Histocompatibility Complex and First Generation of Tag Single-Nucleotide Polymorphisms. Am J Hum Genet. 2005;76 : 634–646. doi: 10.1086/429393 15747258
65. Lehmann DJ, Wiebusch H, Marshall SE, Johnston C, Warden DR, Morgan K, et al. HLA class I, II & III genes in confirmed late-onset Alzheimer’s disease. Neurobiol Aging. 2001;22 : 71–77. 11164278
66. Neill D, Curran MD, Middleton D, Mawhinney H, Edwardson JA, McKeith I, et al. Risk for Alzheimer’s disease in older late-onset cases is associated with HLA-DRB1*03. Neurosci Lett. 1999;275 : 137–40. 10568518
67. Schmidt H, Williamson D, Ashley-Koch A. HLA-DR15 haplotype and multiple sclerosis: a HuGE review. Am J Epidemiol. 2007;165 : 1097–109. doi: 10.1093/aje/kwk118 17329717
68. International Multiple Sclerosis Genetics Consortium, Wellcome Trust Case Control Consortium 2, Sawcer S, Hellenthal G, Pirinen M, Spencer CCA, et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature. 2011;476 : 214–9. doi: 10.1038/nature10251 21833088
69. Patsopoulos NA, Barcellos LF, Hintzen RQ, Schaefer C, van Duijn CM, Noble JA, et al. Fine-mapping the genetic association of the major histocompatibility complex in multiple sclerosis: HLA and non-HLA effects. PLoS Genet. 2013;9: e1003926. doi: 10.1371/journal.pgen.1003926 24278027
70. Lysandropoulos AP, Racapé J, Holovska V, Toungouz M. Human leucocyte antigen (HLA) class I and II typing in Belgian multiple sclerosis patients. Acta Neurol Belg. 2016;
71. Wissemann WT, Hill-Burns EM, Zabetian CP, Factor SA, Patsopoulos N, Hoglund B, et al. Association of Parkinson disease with structural and regulatory variants in the HLA region. Am J Hum Genet. 2013;93 : 984–993. doi: 10.1016/j.ajhg.2013.10.009 24183452
72. Nalls MA, Plagnol V, Hernandez DG, Sharma M, Sheerin UM, Saad M, et al. Imputation of sequence variants for identification of genetic risks for Parkinson’s disease: A meta-analysis of genome-wide association studies. Lancet. 2011;377 : 641–649. doi: 10.1016/S0140-6736(10)62345-8 21292315
73. Al-Hakbany M, Awadallah S, AL-Ayadhi L. The Relationship of HLA Class I and II Alleles and Haplotypes with Autism: A Case Control Study. Autism Res Treat. 2014;2014 : 1–6.
74. Tafti M, Hor H, Dauvilliers Y, Lammers GJ, Overeem S, Mayer G, et al. DQB1 Locus Alone Explains Most of the Risk and Protection in Narcolepsy with Cataplexy in Europe. Sleep. 2014;37 : 19–25. doi: 10.5665/sleep.3300 24381371
75. Chabas D, Taheri S, Renier C, Mignot E. The genetics of narcolepsy. Annu Rev Genomics Hum Genet. 2003;4 : 459–83. doi: 10.1146/annurev.genom.4.070802.110432 14527309
76. Mignot E, Hayduk R, Black J, Grumet FC, Guilleminault C. HLA DQB1*0602 is associated with cataplexy in 509 narcoleptic patients. Sleep. 1997;20 : 1012–20. 9456467
77. Watson NF, Ton TGN, Koepsell TD, Gersuk VH, Longstreth WT Jr. Does narcolepsy symptom severity vary according to HLA-DQB1*0602 allele status? Sleep. 2010;33 : 29–35. 20120618
78. Risacher SL, Kim S, Nho K, Foroud T, Shen L, Petersen RC, et al. APOE effect on Alzheimer’s disease biomarkers in older adults with significant memory concern. Alzheimer’s Dement. 2015;11 : 1417–1429.
79. Hsieh A-R, Chang S-W, Chen P-L, Chu C-C, Hsiao C-L, Yang W-S, et al. Predicting HLA genotypes using unphased and flanking single-nucleotide polymorphisms in Han Chinese population. BMC Genomics. 2014;15 : 81. doi: 10.1186/1471-2164-15-81 24476119
80. Dilthey A, Leslie S, Moutsianas L, Shen J, Cox C, Nelson MR, et al. Multi-Population Classical HLA Type Imputation. PLoS Comput Biol. 2013;9.
81. Yu L, Chibnik LB, Srivastava GP, Pochet N, Yang J, Xu J, et al. Association of Brain DNA methylation in SORL1, ABCA7, HLA-DRB5, SLC24A4, and BIN1 with pathological diagnosis of Alzheimer disease. JAMA Neurol. 2015;72 : 15–24. doi: 10.1001/jamaneurol.2014.3049 25365775
Štítky
Interní lékařství
Článek vyšel v časopisePLOS Medicine
Nejčtenější tento týden
2017 Číslo 3- U pacientů s traumatem je častěji dosaženo adekvátních hladin anti-Xa při dávkování enoxaparinu podle hmotnosti
- Riziko rozvoje závažné infekce po císařském řezu lze snížit provedením antiseptického výplachu vaginy
- Spolehlivý systém skórování jizev k hodnocení fotografií popálenin
- Intermitentní hladovění v prevenci a léčbě (nejen) civilizačních chorob
- Stárnutí populace: problém, či výzva pro české zdravotnictví a medicínu?
-
Všechny články tohoto čísla
- 2016 Reviewer and Editorial Board Thank You
- , , and mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases
- Differential associations of plasma lipids with incident dementia and dementia subtypes in the 3C Study: A longitudinal, population-based prospective cohort study
- Mixed pathologies and neural reserve: Implications of complexity for Alzheimer disease drug discovery
- -related risk of mild cognitive impairment and dementia for prevention trials: An analysis of four cohorts
- Neuropathological diagnoses and clinical correlates in older adults in Brazil: A cross-sectional study
- Early diagnosis of mild cognitive impairment and mild dementia through basic and instrumental activities of daily living: Development of a new evaluation tool
- Potentially modifiable lifestyle factors, cognitive reserve, and cognitive function in later life: A cross-sectional study
- Association between fatty acid metabolism in the brain and Alzheimer disease neuropathology and cognitive performance: A nontargeted metabolomic study
- Fine-mapping of the human leukocyte antigen locus as a risk factor for Alzheimer disease: A case–control study
- What’s the “Take Home” from Research on Dementia Trends?
- Cultural representations of dementia
- Dementia and aging populations—A global priority for contextualized research and health policy
- Dementia in the oldest old: Beyond Alzheimer disease
- Rehabilitation for people living with dementia: A practical framework of positive support
- Dementia in low-income and middle-income countries: Different realities mandate tailored solutions
- Challenges and opportunities in understanding dementia and delirium in the acute hospital
- Dementia incidence trend over 1992-2014 in the Netherlands: Analysis of primary care data
- Association between delirium superimposed on dementia and mortality in hospitalized older adults: A prospective cohort study
- Development of an adaptive, personalized, and scalable dementia care program: Early findings from the Care Ecosystem
- Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score
- Age-related cognitive decline and associations with sex, education and apolipoprotein E genotype across ethnocultural groups and geographic regions: a collaborative cohort study
- The impact of individual Cognitive Stimulation Therapy (iCST) on cognition, quality of life, caregiver health, and family relationships in dementia: A randomised controlled trial
- Effectiveness of an intervention to facilitate prompt referral to memory clinics in the United Kingdom: Cluster randomised controlled trial
- Subjective and objective cognitive function among older adults with a history of traumatic brain injury: A population-based cohort study
- Association of lifelong exposure to cognitive reserve-enhancing factors with dementia risk: A community-based cohort study
- Multimorbidity and healthcare utilization among home care clients with dementia in Ontario, Canada: A retrospective analysis of a population-based cohort
- PLOS Medicine
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Effectiveness of an intervention to facilitate prompt referral to memory clinics in the United Kingdom: Cluster randomised controlled trial
- , , and mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases
- Challenges and opportunities in understanding dementia and delirium in the acute hospital
- Early diagnosis of mild cognitive impairment and mild dementia through basic and instrumental activities of daily living: Development of a new evaluation tool
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: MUDr. Petr Výborný, CSc., FEBO
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání





