-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariérní portál
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Combined Bioinformatics and Literature Based Approach for Identification of Long Non-coding RNAs That Modulate Vitamin D Receptor Signaling in Breast Cancer
Kombinovaný bioinformatický a literární přístup k identifikaci dlouhých nekódujících molekul RNA, které modulují signalizaci přes receptor vitaminu D u karcinomu prsu
Úvod:
Bylo prokázáno, že dlouhé nekódující RNA (lncRNA) jako důležitá frakce lidského transkriptomu hrají zásadní roli při regulaci signálních drah, které se podílejí na karcinogenezi. Mezi nimi je signalizace receptoru vitaminu D (VDR), jejíž účast na různých nádorech vč. nádoru prsu (breast cancer – BC) je patrná. Navzdory přítomnosti několika důkazů účasti lncRNA, stejně jako signalizace VDR v patogenezi BC, žádná souhrnná studie nehodnotila vztah mezi dysregulací lncRNA a signalizací VDR u BC.Cíl:
Zavést bioinformatický přístup k identifikaci lncRNA, které modulují signalizaci VDR u BC. Tento přístup zahrnuje koexpresní analýzu, in silico identifikaci lncRNAs, které jsou zaměřeny na VDR a literární vyhledávání. Závěr: Předpokládá se, že desítky lncRNA ovlivní signalizaci VDR. Mezi nimi jsou některé lncRNA, jako je MALAT1, který má významnou roli v patogenezi BC. Identifikace lncRNA, které ovlivňují expresi genu VDR, je možná pomocí in silico analýzy. Vzhledem k prominentní roli VDR v patogenezi BC a dostupnosti modulačních činidel VDR je hodnocení VDR signalizační dráhy a souvisejících sítí praktického významu a nástroje bioinformatiky by měly usnadnit tuto činnost.Klíčová slova:
receptor vitaminu D – dlouhé nekódující molekuly RNA – koexprese – bioinformatika – receptor kalcitriolu – výpočetní biologieTento článek je výňatkem z práce Vahida Kholghi Oskooei z lékařské fakulty, Univerzity Shahid Beheshti (registrační číslo 46).
Autoři deklarují, že v souvislosti s předmětem studie nemají žádné komerční zájmy.
Redakční rada potvrzuje, že rukopis práce splnil ICMJE kritéria pro publikace zasílané do biomedicínských časopisů.
Obdrženo: 17. 3. 2018
Accepted: 6. 5. 2018
Authors: Kholghi Oskooei Vahid; Ghafouri-Fard Soudeh; Omrani Mir Davood
Authors place of work: Department of Medical Genetics, Shahid Beheshti University of Medical Sciences, Tehran, Iran
Published in the journal: Klin Onkol 2018; 31(4): 264-269
Category: Přehled
doi: https://doi.org/10.14735/amko2018264Summary
Background:
Long non-coding RNAs (lncRNAs) as an important fraction of human transcriptome have been shown to exert fundamental role in regulation of signaling pathways implicated in carcinogenesis. Among them is vitamin D receptor (VDR) signaling whose participation in various cancers including breast cancer (BC) is evident. In spite of the presence of several evidences for participation of lncRNAs as well as VDR signaling in BC pathogenesis, no comprehensive study has evaluated the link between lncRNA dysregulation and VDR signaling in BC.Aim:
To introduce a bioinformatics approach for identification of lncRNAs that modulate VDR signaling in BC. This approach includes co-expression analysis, in silico identification of lncRNAs that target VDR and literature search. Conclusions: Tens of lncRNAs are predicted to affect VDR signaling. Among them are some lncRNAs such as MALAT1 which has prominent role in BC pathogenesis. Identification of the lncRNAs that influence VDR gene expression is possible through in silico analysis. Considering the prominent role of VDR in BC pathogenesis as well as availability of VDR modulating agents, evaluation of VDR signaling pathway and related networks are of practical significance and bioinformatics tools are expected to facilitate such action.Key words:
vitamin D receptor – long non-coding RNAs – co-expression – bioinformatics – calcitriol receptor – computational biologyIntroduction
Breast cancer (BC) as the most common women’s malignancy is regarded as an important health problem [1]. Several researchers have identified biomarkers for early detection or prognosis evaluation of BC patients [2–4]. Among pathways which involvement in BC pathogenesis has been well studied is vitamin D receptor (VDR) signaling pathway. The VDR is a member of the nuclear class II receptor family and a ligand transcription factor that facilitates the roles of 1,25-dihydroxyvitamin D3 in cell growth and differentiation [5]. In BC samples, VDR expression has been negatively associated with aggressive tumor features, such as large tumor size, hormonal receptor negativity, and triple-negative subtype [6]. In addition, elevated expression of VDR in breast tumors has been associated with a lower risk of cancer-associated mortality [7,8]. The protective effect of vitamin D against other types of human cancers, such as skin cancer, has also been documented [9]. Such effect has been associated with alterations in the expression of certain transcripts including long non-coding RNAs (lncRNAs) in a way that these lncRNAs have been suggested as skin cancer biomarkers which are secreted into the blood or urine via exosomes [10]. In general, lncRNAs comprise an important portion of human transcriptome with fundamental roles in virtually every aspect of cell physiology [11]; their aberrant expression has been associated with pathologic conditions such as cancer [12–18]. More specifically, VDR-regulated lncRNAs has been shown to participate in imprinting, tumor suppression and invasion/metastasis which implies their involvement in the protective effect of VDR signaling against skin cancer [10]. LncRNAs expression in skin cells changes in response to vitamin D in a way that diminishes their oncogenic activity while increasing their tumor suppressive role [9]. In BC, there is also evidence for the presence of a link between VDR signaling and lncRNAs. For instance, H19 has been shown to be up-regulated in a significant proportion of BC tissues [19]. On the other hand, H19 has been shown to suppress VDR expression via microRNA 675-5p (miR-675-5p) in colon cancer. Besides, H19 up-regulation leads to vitamin D resistance both in vitro and in vivo [20]. Recently, we have introduced a bioinformatics approach for identification of miRNAs implicated in BC [21].
In spite of the presence of evidence supporting the individual role of lncRNAs as well as VDR signaling in BC pathogenesis, no comprehensive study has evaluated the link between lncRNA dysregulation and VDR signaling in BC. Consequently, in the present study, we introduce a bioinformatics approach for identification of lncRNAs that modulate VDR signaling in BC.
Material and Methods
Co-expression analysis
In order to find lncRNAs, which are co-expressed in breast tissues with VDR, we used co-LncRNA. This web-based computational tool facilitates detection of Gene ontology (GO) annotations and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways that are influenced by co-expressed proteincoding genes and lncRNAs. Co-expression pattern of lncRNAs and protein-coding genes has been retrieved from publicly available human RNA-Seq datasets, comprising 241 datasets from 6 560 total datasets which exemplify 28 tissue types/cell lines. Subsequently, lncRNA combinatorial influence on particular GO annotations or KEGG pathways is analyzed [22]. Spearman rank order correlation analysis was used to define the relationship between expression of VDR and certain lncRNAs. For such purpose, a Spearman correlation rank order correlation coefficient higher than 0.5 and p value less than 0.001 were regarded as statistically significant.
In silico identification of lncRNAs that target VDR
LncRNA2Target tool was used for identification of lncRNAs that function upstream of VDR. This database provides the list of lncRNA targets, which have been identified through lncRNA knockdown or overexpression experiments [23].
Identification of lncRNAs with genomic changes in breast cancer
Subsequently, we used the cBioPortal for Cancer Genomics tool and the Catalogue of Somatic Mutations in Cancer (COSMIC) to find lncRNAs that harbor genomic alteration in BC tissues. The cBioPortal for Cancer Genomics facilitates discovering, envisioning, and analyzing multidimensional cancer genomics data at genetic, epigenetic, gene expression, and proteomic levels [24]. COSMIC is regarded as the most all-inclusive source for discovering the effect of somatic mutations in human cancer [25].
In silico functional analysis of lncRNAs
The functional interactions of selected lncRNAs with tumor suppressor and oncogenes (including both mRNA coding and miRNA genes) which are implicated in BC have been evaluated using two online tools. AnnoLnc is an online tool which provides lncRNAs annotations including their interactions with miRNA and proteins [26]. The miRcode has provided a map of possible miRNA target sites across the complete GENCODE annotated transcriptome, including more than 10 000 lncRNA genes so facilitates the identification of miRNA-lncRNA interactions [27]. Schema 1 shows the pipeline used for identification of VDR related lncRNAs in BC.
Schema 1. The pipeline used for identification of VDR related lncRNAs in breast cancer. 
VDR – vitamin D receptor, lncRNA – long non-coding RNA, CNV – copy number variation Network construction
Pathway studio software [28] was used for construction of a network among VDR, miRNAs lncRNAs and mRNA coding genes.
Results
By using co-lncRNA tool, we could identify 304 lncRNAs which are co-expressed with VDR in BC tissues. Then, we used LncRNA2Target tool to identify VDR upstream lncRNAs which led to identification of four lncRNAs including metastasis associated lung adenocarcinoma transcript 1 (MALAT1), lincFOXF1, lincTNS1 and DA125942. Application of cBioPortal and COSMIC resulted in identification of 26 lncRNAs among total 304 lncRNAs which have been the subjects of amplification, deletion or mutation in BC tissues. Finally, we have shown that these lncRNAs have interactions with transcription factors, such as MYC, GATA3, ZNF217, TP53 and ESR1 as well as BC related tumor suppressors or oncogenes (RB1, PTEN, CCND1, ERBB2, FGFR1, MAP3K1, MUC16, PIK3CA and TTN). In addition, the interactions of the selected lncRNAs with oncomiRs (miR-125b, miR-205, miR-17-92,miR-206, miR-200, miR-146b, miR-126, miR-335 and miR-31) as well as tumor suppressor miRNAs (miR-10b, miR-21, miR-155, miR-373 and miR-520c) have been demonstrated. Tab. 1 shows the summary of lncRNAs that possibly regulate VDR signaling in BC. Using Pathway studio, we designed a network among VDR, miRNAs, lncRNAs and mRNA coding genes. By using “regulation”, “direct regulation”, “binding”, “promoter binding” and “expression” filters, 829 genes (including mRNA coding, lncRNAs and miRNAs) were retrieved that interact with VDR. Subsequently, we narrowed the search by exclusive inclusion of cancer related genes which led to construction of the desired network.
Tab. 1. Summary of lncRNAs that possibly regulate VDR signaling in breast cancer. 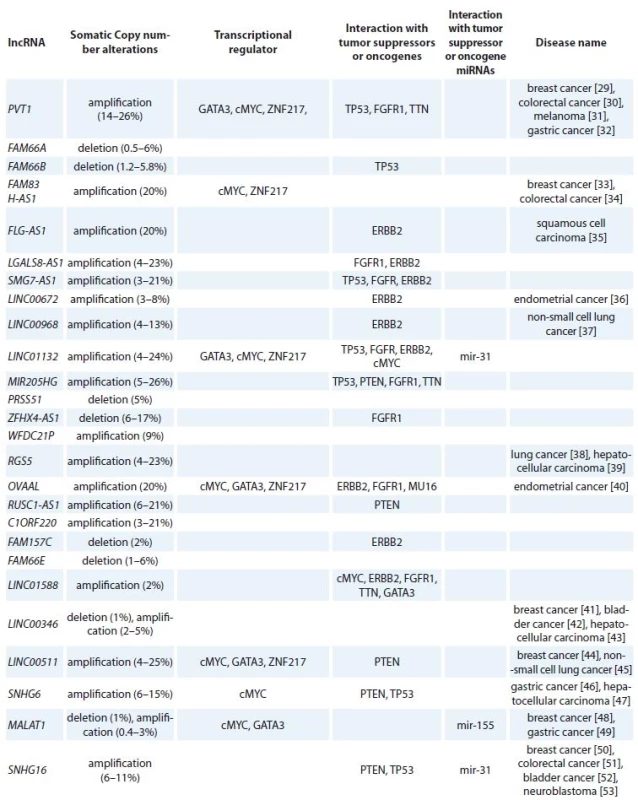
lncRNA – long non-coding RNA, VDR – vitamin D receptor, Discussion
In addition to protein coding mRNAs which have been documented to be regulated by VDR through application of microarray based methods, miRNAs and lncRNAs have been shown to be regulated by this signaling pathway as well. Several patterns of co-expression, co-regulation and interactions have been revealed through integrated analyses of mRNA, miRNAs and lncRNAs [54]. Animal studies have shown the effect of VDR on lncRNA expression in a way that in VDR null mouse epidermis, mHOTAIR, Malat1 and SRA are up-regulated while Foxn2-as, Gtl2-as and H19-as are down-regulated [55]. However, VDR transcriptome puzzle has many gaps which is mostly originated from the scarcity of RNA-Seq data focused on VDR function [54]. With the purpose of filling such knowledge gap, in the present study, we aimed at identification of the complex network between lncRNA expression, VDR signaling and BC using a novel bioinformatics approach. Previously, several bioinformatics approaches have been suggested for identification of disease or phenotype related VDR downstream networks –combination of VDR and chromatin immunoprecipitation (ChIP) -Seq studies with genome-wide association studies or combination of VDR ChIP-Seq with Cancer Genome Atlas (TCGA) data to evaluate the influence of VDR target genes in tumorigenesis process [56]. In the present study, using co-LncRNA tool, we assessed lncRNAs that are co-expressed with VDR in BC tissues. To find more clinically relevant candidates, we used other tools to identify those with genomic alterations in BC tissues and assess their interactions with known tumor suppressor genes and oncogenes. The final lncRNA list provided by this approach offers researchers potential candidates for functional or expression analyses. Finally, we demonstrated the interaction network between mRNA coding genes, lncRNAs and VDR. As demonstrated in this network, linc00261 is among lncRNAs which bind to VDR. Linc00261 is a tumor suppressor which decreases the stability of Slug proteins leading to inhibition of epithelial-mesenchymal transition [57]. SLUG has been previously shown to bind to the E2-box sequences of the VDR gene promoter leading to suppression of VDR gene expression through chromatin remodeling [58]. MALAT1 as a well-known lncRNA in VDR signaling pathway has been demonstrated to interact with several genes in this network. However, due to scarcity of experimental data regarding the interactions between lncRNAs and VDR, this network does not include many of putative lncRNAs. Future experimental studies would help in enrichment of this network.
Identification of the lncRNAs that influence VDR gene expression is possible through genome-wide or individual gene expression analysis followingsilencing or overexpressing each lncRNA. However, such experiments are time-consuming and need prior identification of potential candidates which are expected to influence or be influenced by VDR signaling pathway. Considering the prominent role of VDR in BC pathogenesis as well as availability of VDR modulating agents, evaluation of VDR signaling pathway and related networks are of practical significance and bioinformatics tools are expected to facilitate such action.
This article has been extracted from the thesis written by Vahid Kholghi Oskooei in School of Medicine, Shahid Beheshti University of Medical Sciences (Registration No: 46).
The authors declare they have no potential conflicts of interest concerning drugs, products, or services used in the study.
The Editorial Board declares that the manuscript met the ICMJE recommendation for biomedical papers.
Soudeh Ghafouri-Fard, MD, PhD
Department of Medical Genetics
Shahid Beheshti University of Medical Sciences
Bldg No.2 SBUMS Arabi Ave, Daneshjoo Blvd, Velenjak Tehran, Iran
e-mail: s.ghafourifard@sbmu.ac.ir
Submitted: 17. 3. 2018
Accepted: 6. 5. 2018
Zdroje
1. Dianatpour M, Mehdipour P, Nayernia K et al. Expression of testis specific genes TSGA10, TEX101 and ODF3 in breast cancer. Iran Red Crescent Med J 2012; 14 (11): 722–726. doi: 10.5812/ircmj.3611.
2. Sarrafzadeh S, Geranpayeh L, Ghafouri-Fard S. Expression analysis of long non-coding PCAT-1in breast cancer. Int J Hematol Oncol Stem Cell Res 2017; 11 (3): 185–191.
3. Seifi-Alan M, Shamsi R, Ghafouri-Fard S et al. Expression analysis of two cancer-testis genes, FBXO39 and TDRD4, in breast cancer tissues and cell lines. Asian Pac J Cancer Prev 2014; 14 (11): 6625–6629.
4. Kazemi-Oula G, Ghafouri-Fard S, Mobasheri MB et al. Upregulation of RHOXF2 and ODF4 expression in breast cancer tissues. Cell J 2015; 17 (3): 471–477.
5. Ditsch N, Toth B, Mayr D et al. The Association between vitamin D receptor expression and prolonged overall survival in breast cancer. J Histochem Cytochem 2012; 60 (2): 121–129.
6. Al-Azhri J, Zhang YL, Bshara W et al. Tumor expression of vitamin D receptor and breast cancer histopathological characteristics and prognosis. Clin Cancer Res 2017; 23 (1): 97–103. doi: 10.1158/1078-0432.CCR-16-0075.
7. Almlof JC, Lundmark P, Lundmark A et al. Single nucleotide polymorphisms with cis-regulatory effects on long non-coding transcripts in human primary monocytes. PLoS One 2014; 9 (7): e102612. doi: 10.1371/journal.pone.0102612.
8. Ditsch N, Toth B, Mayr D et al. The association between vitamin D receptor expression and prolonged overall survival in breast cancer. J Histochem Cytochem 2012; 60 (2): 121–129. doi: 10.1369/0022155411429155.
9. Bikle DD, Jiang Y. The protective role of vitamin D signaling in non-melanoma skin cancer. Cancers 2013; 5 (4): 1426–1438. doi: 10.3390/cancers5041426.
10. Jiang YJ, Bikle DD. LncRNA: a new player in 1α, 25 (OH) 2 vitamin D3/VDR protection against skin cancer formation. Exp Dermatol 2014; 23 (3): 147–150. doi: 10.1111/exd.12341.
11. Dianatpour A, Ghafouri-Fard S. The role of long non coding RNAs in the repair of DNA double strand breaks. Int J Mol 2017; 6 (1): 1–12.
12. Dianatpour A, Ghafouri-Fard S. Long non coding RNA expression intersecting cancer and spermatogenesis: a systematic review. Asian Pac J Cancer Prev 2017; 18 (10): 2601–2610. doi: 10.22034/APJCP.2017.18.10.2601.
13. Nikpayam E, Soudyab M, Tasharrofi B et al. Expression analysis of long non-coding ATB and its putative target in breast cancer. Breast Dis 2017; 37 (1): 11–20. doi: 10.3233/BD-160264.
14. Sarrafzadeh S, Geranpayeh L, Tasharrofi B et al. Expression study and clinical correlations of MYC and CCAT2 in breast cancer patients. Iran Biomed J 2017; 21 (5): 303–311.
15. Iranpour M, Soudyab M, Geranpayeh L et al. Expression analysis of four long noncoding RNAs in breast cancer. Tumour Biol 2016; 37 (3): 2933–2940. doi: 10.1007/s13277-015-4135-2.
16. Soudyab M, Iranpour M, Ghafouri-Fard S. The role of long non-coding RNAs in breast cancer. Arch Iran Med 2016; 19 (7): 508–517. doi: 0161907/AIM.0011.
17. Nikpayam E, Tasharrofi B, Sarrafzadeh S et al. The role of long non-coding RNAs in ovarian cancer. Iranian biomedical journal 2017; 21 (1): 3–15. doi:
18. Tasharrofi B, Soudyab M, Nikpayam E et al. Comparative expression analysis of hypoxia-inducible factor-alpha and its natural occurring antisense in breast cancer tissues and adjacent noncancerous tissues. Cell Biochem Funct 2016; 34 (8): 572–578. doi: 10.1002/cbf.3230.
19. Collette J, Le Bourhis X, Adriaenssens E. Regulation of human breast cancer by the long non-coding RNA H19. Int J Mol Sci 2017; 18 (11). Pii: E2319. doi: 10.3390/ijms18112319.
20. Chen S, Bu D, Ma Y et al. H19 Overexpression induces resistance to 1, 25 (OH) 2D3 by targeting VDR through miR-675-5p in colon cancer cells. Neoplasia 2017; 19 (3): 226–236. doi: 10.1016/j.neo.2016.10.007.
21. Shamsi R, Seifi-Alan M, Behmanesh A et al. A bioinformatics approach for identification of miR-100 targets implicated in breast cancer. Cell Mol Biol 2017; 63 (10): 99–105. doi: 10.14715/cmb/2017.63.10.16.
22. Zhao Z, Bai J, Wu A et al. Co-LncRNA: investigating the lncRNA combinatorial effects in GO annotations and KEGG pathways based on human RNA-Seq data. Database 2015; 2015. pii: bav082. doi: 10.1093/database/bav082.
23. Jiang Q, Wang J, Wu X et al. LncRNA2Target: a database for differentially expressed genes after lncRNA knockdown or overexpression. Nucleic Acids Res 2015; 43: D193–D196. doi: 10.1093/nar/gku1173.
24. Gao J, Aksoy BA, Dogrusoz U et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013; 6 (269): pl1. doi: 10.1126/scisignal.2004088.
25. Forbes SA, Beare D, Boutselakis H et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 2017; 45 (D1): D777–D83. doi: 10.1093/nar/gkw1121.
26. Hou M, Tang X, Tian F et al. AnnoLnc: a web server for systematically annotating novel human lncRNAs. BMC Genomics 2016; 17 (1): 931. doi: 10.1186/s12864-016-3287-9.
27. Jeggari A, Marks DS, Larsson E. miRcode: a map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 2012; 28 (15): 2062–2063. doi: 10.1093/bioinformatics/bts344.
28. Nikitin A, Egorov S, Daraselia N et al. Pathway studio-the analysis and navigation of molecular networks. Bioinformatics 2003; 19 (16): 2155–2157.
29. Conte F, Fiscon G, Chiara M et al. Role of the long non-coding RNA PVT1 in the dysregulation of the ceRNA-ceRNA network in human breast cancer. PLoS One 2017; 12 (2): e0171661. doi: 10.1371/journal.pone.0171661.
30. Guo K, Yao J, Yu Q et al. The expression pattern of long non-coding RNA PVT1 in tumor tissues and in extracellular vesicles of colorectal cancer correlates with cancer progression. Tumour Biol 2017; 39 (4): 1010428317699122. doi: 10.1177/1010428317699122.
31. Chen X, Gao G, Liu S et al. Long noncoding RNA PVT1 as a novel diagnostic biomarker and therapeutic target for melanoma. Biomed Res Int 2017; 2017 : 7038579. doi: 10.1155/2017/7038579.
32. Zhang XW, Bu P, Liu L et al. Overexpression of long non-coding RNA PVT1 in gastric cancer cells promotes the development of multidrug resistance. Biochem Biophys Res Commun 2015; 462 (3): 227–232. doi: 10.1016/j.bbrc.2015.04.121.
33. Zhang J, Feng S, Su W et al. Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer. Sci Rep 2017; 7 : 42819. doi: 10.1038/srep42819.
34. Yang L, Xu L, Wang Q et al. Dysregulation of long non-coding RNA profiles in human colorectal cancer and its association with overall survival. Oncol Lett 2016; 12 (5): 4068–4074. doi: 10.3892/ol.2016.5138.
35. Feng L, Houck JR, Lohavanichbutr P et al. Transcriptome analysis reveals differentially expressed lncRNAs between oral squamous cell carcinoma and healthy oral mucosa. Oncotarget 2017; 8 (19): 31521–31531. doi: 10.18632/oncotarget.16358.
36. Li W, Li H, Zhang L et al. Long non-coding RNA LINC00672 contributes to p53 protein-mediated gene suppression and promotes endometrial cancer chemosensitivity. J Biol Chem 2017; 292 (14): 5801–5813. doi: 10.1074/jbc.M116.758508.
37. Wang Y, Zhou J, Xu YJ et al. Long non-coding RNA LINC00968 acts as oncogene in NSCLC by activating the Wnt signaling pathway. J Cell Physiol 2018; 233 (4): 3397–3406. doi: 10.1002/jcp.26186.
38. Huang G, Song H, Wang R et al. The relationship between RGS5 expression and cancer differentiation and metastasis in non-small cell lung cancer. J Surg Oncol 2012; 105 (4): 420–424. doi: 10.1002/jso.22033.
39. Hu M, Chen X, Zhang J et al. Over-expression of regulator of G protein signaling 5 promotes tumor metastasis by inducing epithelial-mesenchymal transition in hepatocellular carcinoma cells. J Surg Oncol 2013; 108 (3): 192–196. doi: 10.1002/jso.23367.
40. Takenaka K, Chen BJ, Modesitt SC et al. The emerging role of long non-coding RNAs in endometrial cancer. Cancer Genet 2016; 209 (10): 445–455. doi: 10.1016/j.cancergen.2016.09.005.
41. Liu H, Li J, Koirala P et al. Long non-coding RNAs as prognostic markers in human breast cancer. Oncotarget 2016; 7 (15): 20584–20596. doi: 10.18632/oncotarget.7828.
42. Ye T, Ding W, Wang N et al. Long noncoding RNA linc00346 promotes the malignant phenotypes of bladder cancer. Biochem Biophys Res Commun 2017; 491 (1): 79–84. doi: 10.1016/j.bbrc.2017.07.045.
43. Zhang J, Fan D, Jian Z et al. Cancer specific long noncoding RNAs show differential expression patterns and competing endogenous RNA potential in hepatocellular carcinoma. PLoS One 2015; 10 (10): e0141042. doi: 10.1371/journal.pone.0141042.
44. Cabanski CR, White NM, Dang HX et al. Pan-cancer transcriptome analysis reveals long noncoding RNAs with conserved function. RNA Biol 2015; 12 (6): 628–642. doi: 10.1080/15476286.2015.1038012.
45. Sun CC, Li SJ, Li G et al. Long intergenic noncoding RNA 00511 acts as an oncogene in non-small-cell lung cancer by binding to EZH2 and suppressing p57. Mol Ther Nucleic Acids 2016; 5 (11): e385. doi: 10.1038/mtna.2016.94.
46. Yan K, Tian J, Shi W et al. LncRNA SNHG6 is associated with poor prognosis of gastric cancer and promotes cell proliferation and EMT through epigenetically silencing p27 and sponging miR-101-3p. Cell Physiol Biochem 2017; 42 (3): 999–1012. doi: 10.1159/000478682.
47. Birgani MT, Hajjari M, Shahrisa A et al. Long non-coding RNA SNHG6 as a potential biomarker for hepatocellular carcinoma. Pathol Oncol Res 2018; 24 (2): 329–337. doi: 10.1007/s12253-017-0241-3.
48. Jadaliha M, Zong X, Malakar P et al. Functional and prognostic significance of long non-coding RNA MALAT1 as a metastasis driver in ER negative lymph node negative breast cancer. Oncotarget 2016; 7 (26): 40418–40436. doi: 10.18632/oncotarget.9622.
49. Li J, Gao J, Tian W et al. Long non-coding RNA MALAT1 drives gastric cancer progression by regulating HMGB2 modulating the miR-1297. Cancer Cell Int 2017; 17 : 44. doi: 10.1186/s12935-017-0408-8.
50. Cai C, Huo Q, Wang X et al. SNHG16 contributes to breast cancer cell migration by competitively binding miR-98 with E2F5. Biochem Biophys Res Commun 2017; 485 (2): 272–278. doi: 10.1016/j.bbrc.2017.02.094.
51. Christensen LL, True K, Hamilton MP net al. SNHG16 is regulated by the Wnt pathway in colorectal cancer and affects genes involved in lipid metabolism. Mol Oncol 2016; 10 (8): 1266–1282. doi: 10.1016/j.molonc.2016.06.003.
52. Zhu Y, Yu M, Li Z et al. ncRAN, a newly identified long noncoding RNA, enhances human bladder tumor growth, invasion, and survival. Urology 2011; 77 (2): 510 e1–e5. doi: 10.1016/j.urology.2010.09.022.
53. Yu M, Ohira M, Li Y et al. High expression of ncRAN, a novel non-coding RNA mapped to chromosome 17q25.1, is associated with poor prognosis in neuroblastoma. Int J Oncol 2009; 34 (4): 931–938.
54. Campbell MJ. Vitamin D and the RNA transcriptome: more than mRNA regulation. Front Physiol 2014; 5.
55. Jiang YJ, Bikle DD. LncRNA profiling reveals new mechanism for VDR protection against skin cancer formation. J Steroid Biochem Mol Biol 2014; 144 Pt A: 87–90. doi: 10.1016/j.jsbmb.2013.11.018.
56. Campbell MJ. Bioinformatic approaches to interrogating vitamin D receptor signaling. Mol Cell Endocrinol 2017; 453 : 3–13. doi: 10.1016/j.mce.2017.03.011.
57. Yu Y, Li L, Zheng Z et al. Long non-coding RNA linc00261 suppresses gastric cancer progression via promoting Slug degradation. J Cell Mol Med 2017; 21 (5): 955–967. doi: 10.1111/jcmm.13035.
58. Mittal MK, Myers JN, Misra S at al. Chaudhuri G. In vivo binding to and functional repression of the VDR gene promoter by SLUG in human breast cells. Biochem Biophys Res Commun 2008; 372 (1): 30–34. doi: 10.1016/j.bbrc.2008.04.187.
Štítky
Dětská onkologie Chirurgie všeobecná Onkologie
Článek Precizovaná onkologie
Článek vyšel v časopiseKlinická onkologie
Nejčtenější tento týden
2018 Číslo 4- Prognostický vliv hodnoty CRP před léčbou u pacientů s metastatickým karcinomem ledviny léčených inhibitory tyrozinkinázy
- I „pouhé“ doporučení znamená velkou pomoc. Nasměrujte své pacienty pod křídla Dobrých andělů
- Když se ve střevech děje něco nepatřičného...
- Renesance hirudoterapie aneb léčba pomocí krevsajících pijavic včera, dnes a zítra
- Perorální antivirotika jako vysoce efektivní nástroj prevence hospitalizací kvůli COVID-19 − otázky a odpovědi pro praxi
-
Všechny články tohoto čísla
- Precizovaná onkologie
- Use of Salivary MicroRNAs for Diagnosis of Solid Cancers
- Expression Analysis of OIP5-AS1 in Non-Small Cell Lung Cancer
- A Combined Bioinformatics and Literature Based Approach for Identification of Long Non-coding RNAs That Modulate Vitamin D Receptor Signaling in Breast Cancer
- Monoclonal Gammopathy of Undetermined Significance (MGUS)Monoclonal Gammopathy of Undetermined Significance (MGUS)
- Carcinoma of Unknown Primary in Head and Neck Region
- Occurrence and Antibiotic Resistance of Enterobacteriaceae in Acute Leukemia Patients
- Malignant Melanomas of the Skin Arising on the Feet
- Metastases of a Breast Cancer to Skull Base
- Primary Branchiogenic Carcinoma
- Use of Fludarabine for the Treatment of Indolent Lymphoma with Chylothorax
- The Therapeutic Effects of Specific CDK4/6-inhibitors in Treating HR-positive, HER2-negative Advanced Breast Cancer
- Nové techniky IGRT – sledování povrchu těla pacienta (SIGRT)
- Klinická onkologie
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Monoclonal Gammopathy of Undetermined Significance (MGUS)Monoclonal Gammopathy of Undetermined Significance (MGUS)
- Carcinoma of Unknown Primary in Head and Neck Region
- Occurrence and Antibiotic Resistance of Enterobacteriaceae in Acute Leukemia Patients
- Use of Salivary MicroRNAs for Diagnosis of Solid Cancers
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



