-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaWhole Exome Sequencing in Atrial Fibrillation
Atrial fibrillation is a common and morbid cardiac arrhythmia. Atrial fibrillation is heritable, and numerous genome-wide susceptibility loci have been identified, predominantly in non-coding regions. Over 35 genes also have been implicated in atrial fibrillation pathogenesis mostly through prior smaller scale candidate gene association studies, which generally did not have robust replication to support the associations. Therefore, the role of coding variation in the biology of atrial fibrillation is unclear. We examined whole exome sequencing data from 1,734 individuals with and 9,423 without atrial fibrillation, and did not observe any significant associations between coding variation and the arrhythmia. Furthermore, we did not observe any enrichment for association in previously implicated atrial fibrillation genes. In aggregate, our findings suggest that large effect coding variation is unlikely to be a predominant mechanism of common forms of atrial fibrillation encountered in the community.
Published in the journal: . PLoS Genet 12(9): e32767. doi:10.1371/journal.pgen.1006284
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1006284Summary
Atrial fibrillation is a common and morbid cardiac arrhythmia. Atrial fibrillation is heritable, and numerous genome-wide susceptibility loci have been identified, predominantly in non-coding regions. Over 35 genes also have been implicated in atrial fibrillation pathogenesis mostly through prior smaller scale candidate gene association studies, which generally did not have robust replication to support the associations. Therefore, the role of coding variation in the biology of atrial fibrillation is unclear. We examined whole exome sequencing data from 1,734 individuals with and 9,423 without atrial fibrillation, and did not observe any significant associations between coding variation and the arrhythmia. Furthermore, we did not observe any enrichment for association in previously implicated atrial fibrillation genes. In aggregate, our findings suggest that large effect coding variation is unlikely to be a predominant mechanism of common forms of atrial fibrillation encountered in the community.
Introduction
Atrial fibrillation (AF) is a common [1, 2] arrhythmia associated with substantial morbidity [3–7]. Current treatments for AF have limited efficacy and can cause significant adverse effects [8, 9]. AF is heritable and approximately one in four individuals with AF has a first-degree relative with the condition [10].
In recent years a large number of genes have been implicated in AF risk using both genome-wide association studies and candidate gene screening approaches. Large-scale genome-wide association studies have identified multiple AF susceptibility loci [11–15], and the top variants at discovered loci have largely been localized to noncoding regions of the genome. In contrast, there have been over 35 genes implicated in AF in candidate gene studies [16]. These studies have had a number of limitations including small sample sizes, consideration of only one or a small number of genes, and the lack of suitable control populations. To date, large-scale studies to determine whether these genes are truly related to AF have not been performed.
Since the discovery of genes causally related to AF may enable a better understanding of AF pathogenesis and potentially inform the development of therapies for AF, there is a critical need to systematically identify the genetic basis of AF. We therefore sought to assess the relations between coding variation and AF in a large sample of individuals who underwent whole exome sequencing. We further sought to determine whether coding variation in genes implicated in AF was enriched among AF cases.
Results
The current analysis included 6,737 participants of European ancestry (n = 1,155 AF events) and 1,246 participants of African ancestry (n = 246 AF cases) from a Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) exome sequencing effort and 1,919 participants of European ancestry (n = 233 cases) and 1,255 participants of African ancestry (n = 100 AF events) from the NHLBI-GO Exome Sequencing Project (ESP). The clinical characteristics of studied participants are listed in Table 1. Sequencing coverage for the subset of AF genes is provided in S2 Table.
Tab. 1. Baseline characteristics of the participating studies. 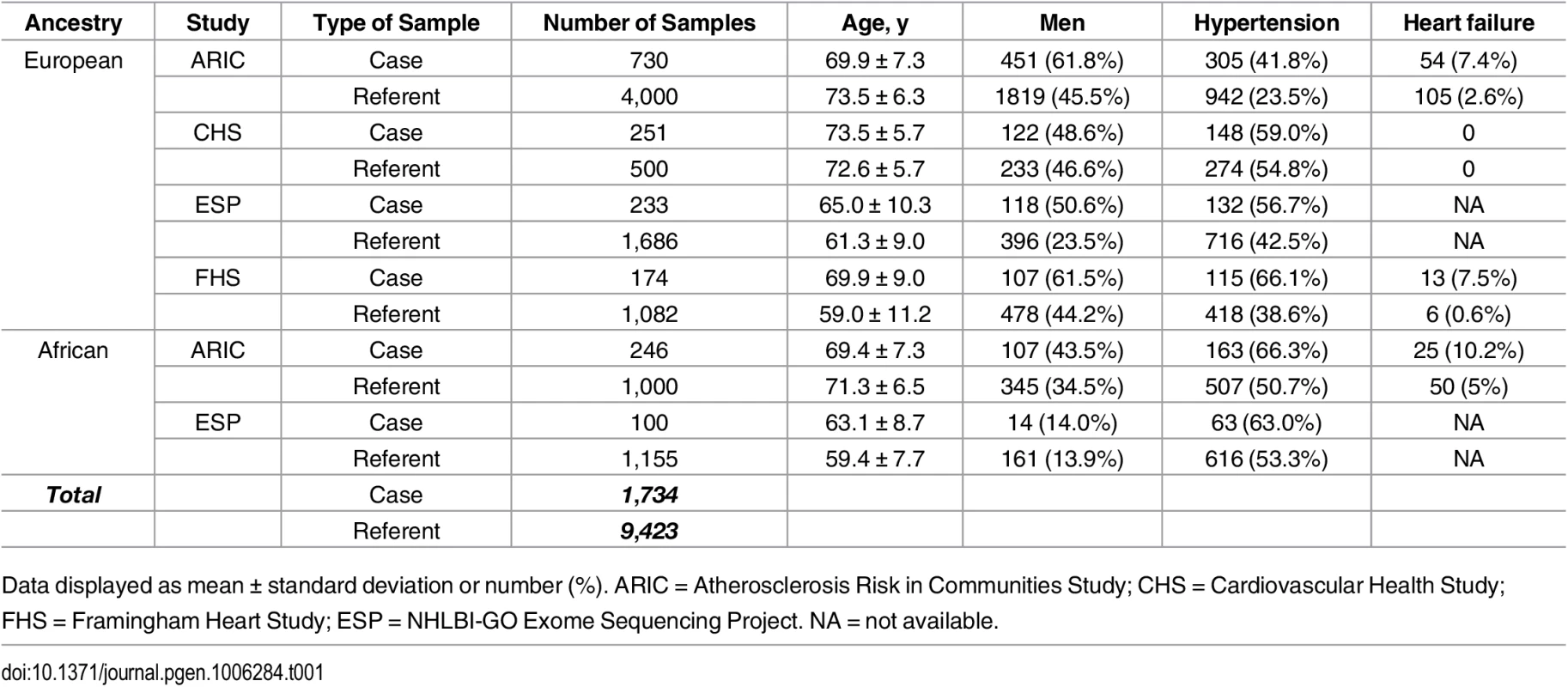
Data displayed as mean ± standard deviation or number (%). ARIC = Atherosclerosis Risk in Communities Study; CHS = Cardiovascular Health Study; FHS = Framingham Heart Study; ESP = NHLBI-GO Exome Sequencing Project. NA = not available. Association of common variants with AF
A total of 99,404 common variants (MAF≥0.01) were included in our study. Approximately 99.7% of the variants were already reported in dbSNP (version 142) or the 1000 Genomes Project. The Manhattan plot representing the primary pooled ancestry analysis is displayed in Fig 1 and the QQ plot is shown in S1 Fig. No inflation of Type I error was observed (genomic control λ = 0.91).
Fig. 1. Manhattan plot of common variant associations with atrial fibrillation. 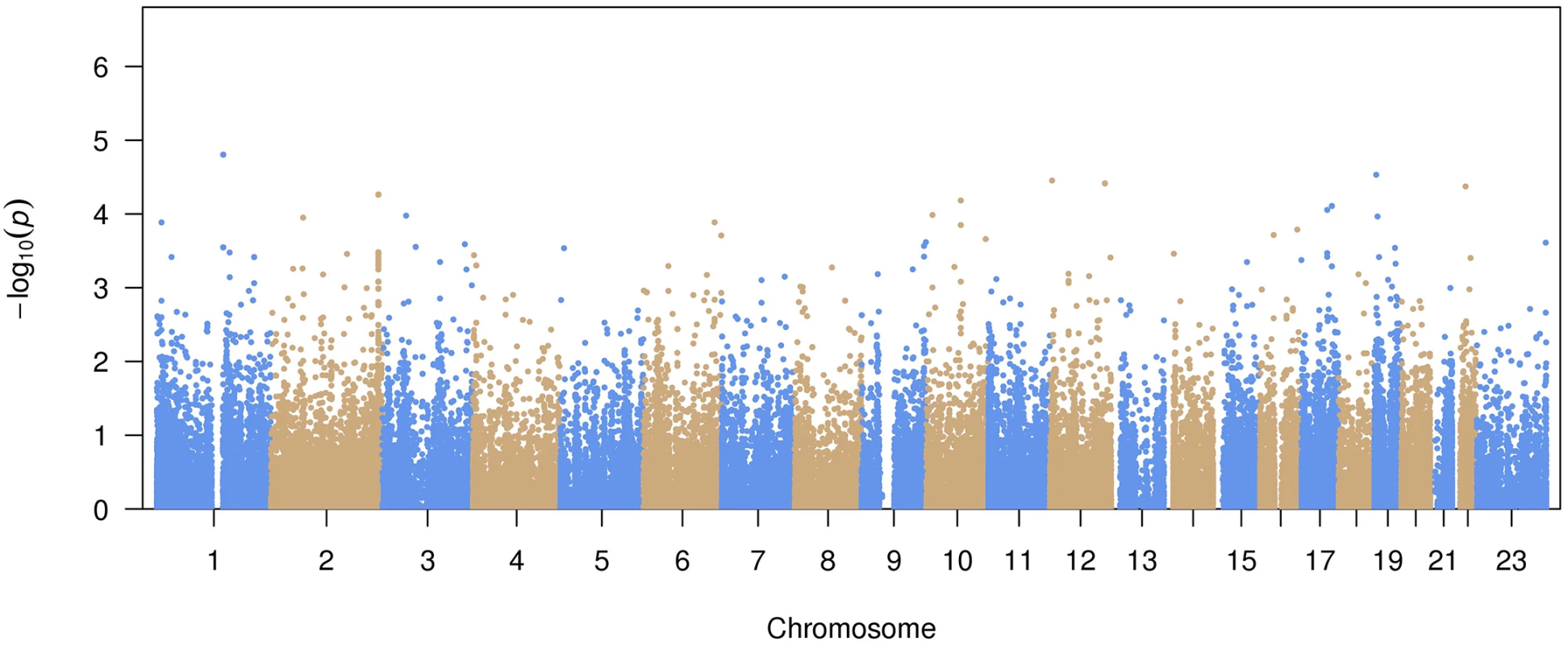
The top 15 variants most significantly associated with AF are listed in Table 2. No common variants were significantly associated with AF after Bonferroni correction for multiple testing (all P>0.05/99,404 = 5.0x10-7). The most significantly associated variant was rs56025621 (P = 1.6x10-5), which is located in the first intron of HFE2, a gene encoding the hemochromatosis type 2 peptide. The SNP was not genotyped in HapMap phase II, thus the association between rs56025621 and AF was not assessed in previous genome-wide association studies.
Tab. 2. Most significant common variants associated with atrial fibrillation. 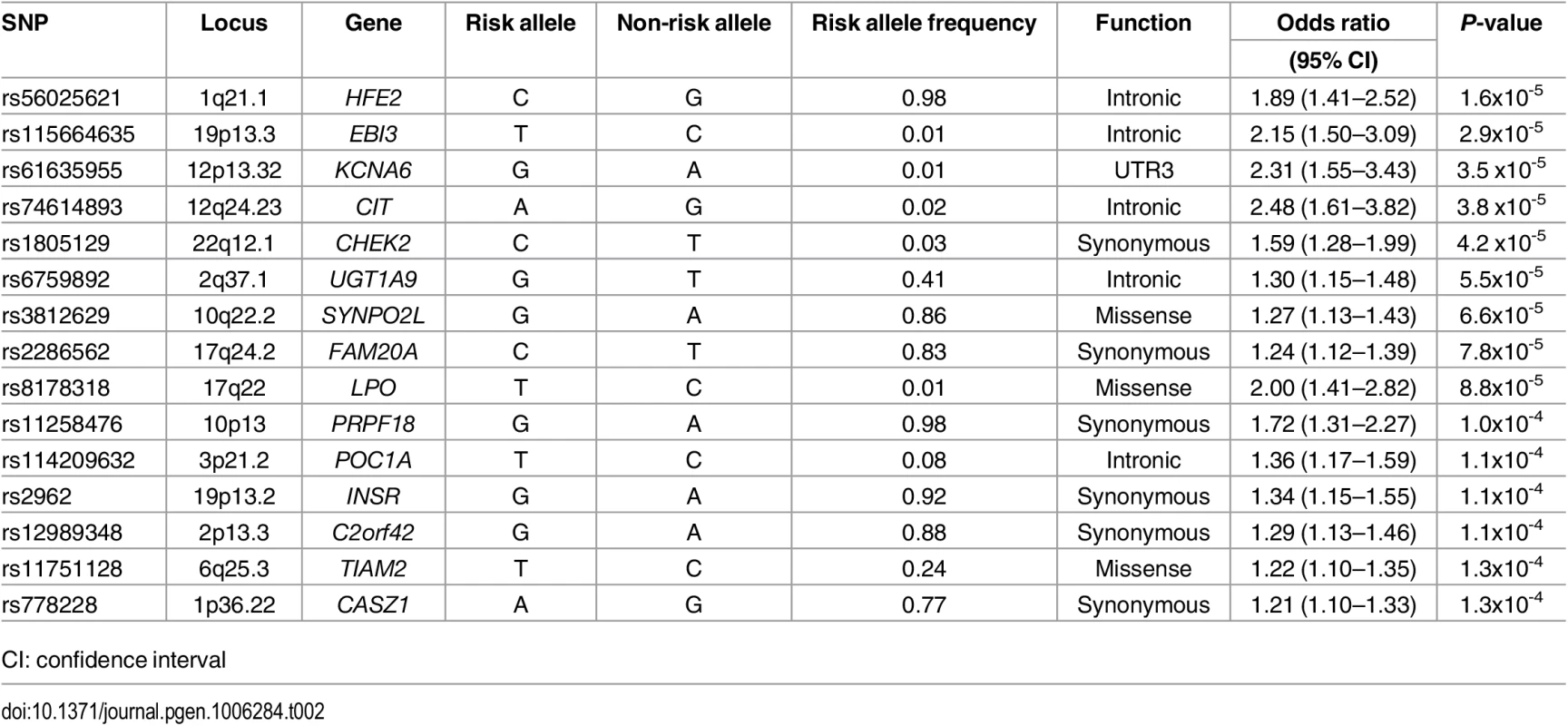
CI: confidence interval The SNP rs3812629, a missense variant encoding a proline to leucine amino acid substitution at amino acid 707 of SYNPO2L, occurs at a genome-wide significant disease susceptibility locus for AF [15]. The variant is in moderate linkage disequilibrium (r2 = 0.69 European ancestry, 1000 Genomes Project) with the top SNP (rs10824026) associated with AF at the locus in a prior genome-wide association study [15]. In the subset of individuals with both genome-wide genotyping data and exome sequence data available from ARIC (n = 6,630), CHS (n = 671), and FHS (n = 1,256) we examined associations between the top noncoding SNP (rs10824026) and the p.Pro707Leu (rs3812629) variant with AF after adjustment for one another (S3 Table). Adjustment for the coding variant attenuated the signal of the lead GWAS SNP in the analysis, and vice versa, suggesting the two variants represent the same AF susceptibility signal.
Previously reported top SNPs for AF derived from genome-wide association studies [15, 17], which are located in noncoding regions, were not assayed using the capture arrays in this study. As such, they were not analyzed in the current analysis.
Association of rare variants with AF
We collapsed rare variants (MAF<1%) into gene regions and performed association testing between each gene region with AF. Our primary analysis was restricted to nonsynonymous and splice-site variants. We excluded gene regions with a cumulative MAF less than 1%. In total, we tested 8,879 gene regions. None of the gene regions were significantly associated with AF after adjusting for multiple testing (all p>0.05/8,879 = 5.6x10-6). The most significantly associated gene region was IL17REL (p = 1.3x10-5), a gene encoding interleukin 17 receptor E-like (S4 Table). The most significant single variant in IL17REL in this analysis was rs200958270 (OR 6.92, 95% CI 3.38–14.15, p = 1.2x10-7), a missense (p.Glu151Gly) variant that has a minor allele frequency of 0.004%. The variant did not meet our prespecified significance criteria for association. In a secondary analysis restricted to damaging variants, no specific gene regions were significantly associated with AF (S5 Table). Again, the most significantly associated gene in the damaging analysis was again IL17REL (p = 1.9x10-6). Variants in IL17REL have been implicated in inflammatory bowel disease [18, 19] though the relations between variation in IL17REL and cardiac function are unclear.
We also examined the associations between rare coding variants and AF within reported AF-susceptibility genes (Table 3). None were significantly associated with AF after adjusting for multiple testing.
Tab. 3. Genes previously implicated in atrial fibrillation pathogenesis. 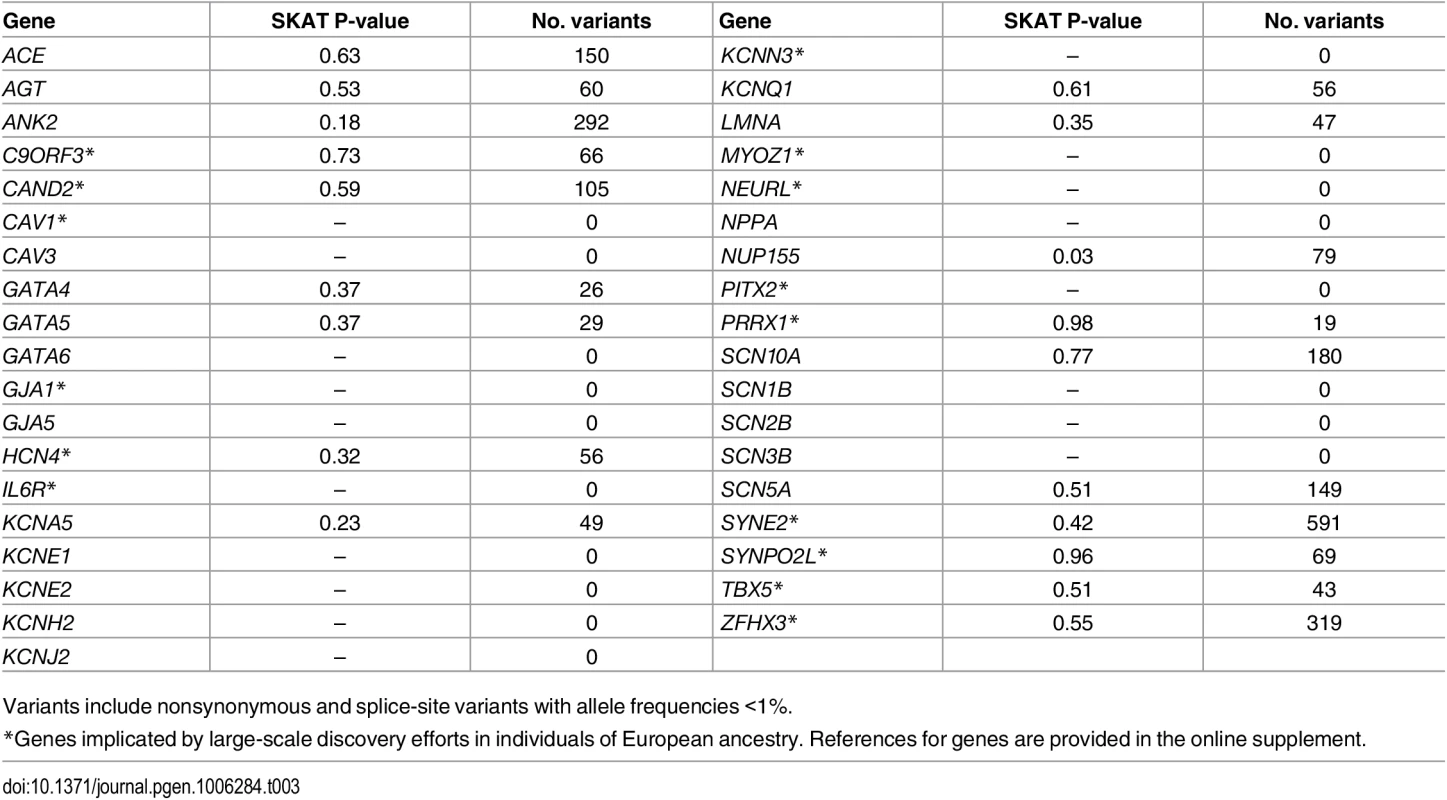
Variants include nonsynonymous and splice-site variants with allele frequencies <1%. In a post-hoc exploratory analyses, we included all rare variants (<1%) within each gene region, irrespective of annotation, and tested them for association with AF using an adjusted significance threshold of p = 2.5x10-6 (0.05/19913 genes). The results are summarized in S6 Table. The lead gene associated with AF was ACY3 (p = 2.2x10-7), which encodes aminoacylase 3. No relation between ACY3 and cardiac function or arrhythmias has been described previously.
With the current sample size, we estimated the statistical power to identify genetic variants with α = 5x10-7, assuming 100,000 independent tests. As shown in Fig 2, we had limited statistical power to identify genetic variants with allele frequencies as low as 1% unless the genetic relative risk was higher than two. In contrast, the statistical power increased significantly for relatively common variants with allele frequencies of at least 5%.
Fig. 2. Statistical power with current sample size and α = 5x10<sup>-7</sup>. 
Pathway enrichment analysis
We subsequently assessed whether genetic variation in pre-specified gene sets was enriched among individuals with AF. We did not observe enrichment for common (FDR = 0.38) or rare (FDR = 0.91) variation in reported AF-related genes among individuals with AF (Table 4).
Tab. 4. Assessment of variant enrichment in genes purportedly implicated in atrial fibrillation. 
Rare variants were defined as those with minor allele frequencies < 1%, and common as those with frequencies ≥ 1%. Discussion
In our sample of 1,734 individuals with and 9,423 without AF who underwent whole exome sequencing, we did not observe any rare coding variation significantly associated with AF. Our observations suggest that coding variation with large effect sizes is unlikely to be the predominant mechanism underlying common forms of AF.
Our results extend prior literature focusing on coding variation underlying AF. Numerous reports propose coding variation as a mechanism underlying AF (S1 Table). However, much of the prior literature was generated via candidate gene association studies. Such discoveries have not been routinely replicated, and the studies were of small size, potentially favoring spurious results. Indeed, we previously observed that most findings from prior AF candidate gene association studies were not replicated when tested in additional study samples [17].
In the context of prior literature and our sample size, our study has two major implications for understanding AF pathogenesis. First, the lack of observed association between coding variation and AF implies that large effect coding variation is not likely to be common in typical forms of AF. In contrast, both noncoding variation, and coding variation with smaller effect sizes, may contribute to AF pathogenesis. Genome-wide association studies have identified highly associated common genetic variants near ion channels, cardiac and pulmonary transcription factors, and other genes in individuals with AF [11–15], underscoring the polygenic nature of AF. Nevertheless, the causal variants and genes underlying the arrhythmia remain unknown. Future whole-genome sequencing efforts may help to clarify the genetic contributions to AF.
Second, our findings suggest that efforts to identify potential therapeutic targets for AF through exome sequencing analyses will require much larger sample sizes or populations enriched for large genetic effects. Such populations might include those with early onset AF or consanguineous populations with the propensity to homozygous loss of function alleles in genes. Nevertheless, the additional cost required to sequence such large populations must be balanced against the potentially more cost-efficient approach of performing GWAS genotyping, imputation, and subsequent functional characterization for genetic discovery. The lack of observation of any prominent coding variation underlying AF is consistent with other whole exome sequencing efforts of complex diseases such as coronary disease and diabetes [20], which generally have not identified coding variation as the major mechanisms underlying these conditions.
Our study should be interpreted in the context of the study design. Our study was predominantly comprised of individuals of European ancestry, and therefore the findings may not be generalizable to other ancestral groups. The individuals with AF may have had multiple etiologies for the condition, and may not have been enriched for genetic forms of the arrhythmia. We cannot exclude that AF may have been misclassified, especially since AF may be paroxysmal and asymptomatic at times. Such misclassification is expected to bias the results toward the null. Furthermore, our study had limited power to assess the role of many coding variants, particularly because classifying missense variants as pathogenic or not remains challenging despite the routine use of bioinformatic algorithms. An earlier report of whole exome sequencing in 6 families with AF has summarized some of the bioinformatics challenges of utilizing whole exome sequencing data [21]. The size of our study sample limited our ability to detect potentially functional rare variants. Additionally, we utilized a Bonferroni significance threshold, which may be overly conservative for genetic discovery.
In conclusion, we observed that coding variation is not a major contributor to AF in a sample of individuals predominantly of European ancestry. Efforts to identify coding variation underlying AF will require much larger study samples. Future analyses that integrate coding and noncoding variation, such as whole genome sequencing, are warranted.
Materials and Methods
Study participants
The current study included participants from three population-based cohorts that participated in the CHARGE exome sequencing effort (N = 15,459 individuals of either European or African ancestry): the Atherosclerosis Risk in Communities study (ARIC), Cardiovascular Health Study (CHS), and Framingham Heart Study (FHS). In ARIC, a random subset of 4000 European ancestry control subjects and 1000 African ancestry subjects were chosen without regard for age or sex matching. Each cohort has been described in detail previously [22–25].
We also included individuals from ESP (N = 6823 individuals of European or African ancestry) in whom AF data were ascertained (cohorts included ARIC, CHS, FHS and the Women's Health Initiative) [26]. We omitted from analysis samples for whom phenotypic data for AF were missing (N = 2593 CHARGE, N = 3689 ESP). Individuals in ESP that overlapped with individuals from the CHARGE effort (n = 40) were omitted to avoid duplicate individuals in analyses. Institutional Review Boards or Ethics Committees approved each contributing study. All participants provided written informed consent to participate in genetic research on cardiovascular disease.
Exome sequencing
We performed a combined analysis of exome sequencing conducted in the CHARGE consortium [27] and ESP [26]. In CHARGE, the exome was captured using NimbleGen SeqCap EZ VCRome (Roche, Basel, Switzerland). The enriched library was then sequenced by Illumina HiSeq platform at Human Genome Sequencing Center at Baylor College of Medicine. The Mercury pipeline [28] was used to process sequencing data, whereas the raw short reads were aligned to the reference human genome (NCBI Genome Build 37, 2009) by Burrows-Wheeler Aligner [29], and the variants were called by Atlas [30]. The mean read depth was 92x, and more than 92% of target regions were covered by at least 20 unique reads. Rigorous quality control was performed to exclude low-quality variants or samples. We excluded variants that were multi-allelic or monomorphic, had a missing rate higher than 20%, had mean depth higher than 500, or had Hardy-Weinberg equilibrium p-value less than 5x10-6 within ancestry groups. For individual samples, we calculated four quality metrics: mean depth, transition to transversion (Ti/Tv) ratio, number of singletons, and heterozygote to homozygote ratio. Samples with any metric exceeding 6 standard deviations in the respective study were omitted from analyses.
ESP included samples from 6823 individuals of European or African ancestry. The details of library construction, sequencing and alignment have been described previously [31–33]. Briefly, the exome was captured using either Agilent SureSelect Human All Exon 50Mb (Agilent, Santa Clara, CA) or NimbleGen SeqCap EZ VCRome (Roche, Basel, Switzerland). The sequencing was performed at the University of Washington and at the Broad Institute of MIT and Harvard. The mean depth was 127x. Variants with mean depth greater than 500, or with missing rate greater than 20% were excluded.
AF ascertainment
Ascertainment of AF in each cohort has been described previously [15]. Briefly, ascertainment of AF was standardized at each participating study and included the presence of either atrial fibrillation or flutter observed on a study electrocardiogram, within obtained medical encounters, or indicated by billing codes. Both incident and prevalent AF were treated together as AF cases for the purposes of this analysis. For ESP, AF information was obtained from the phenotype file (“ESP6800_Phenotype_Update_061212_final.xlsx”), from which individual level phenotypic data was provided.
Statistical analyses
Each cohort from CHARGE performed separate analyses and shared results for downstream meta-analysis. For ESP, samples from all cohorts were treated as a single sample for analyses, and adjusted for study sites and capture kits.
For common variants with minor allele frequency (MAF) at least 1%, the association of variants with AF was tested by multivariable logistic regression (ARIC, CHS, and ESP) or logistic generalized estimating equation to account for familial correlation (FHS). In common variant association analyses, we also included noncoding variants in regions flanking exons that were captured by the exome arrays. For rare variants (MAF<1%), we pooled all rare variants based on RefSeq gene regions, and jointly tested their associations with AF with the Sequence Kernel Association Test (SKAT) [34]. To circumvent the dilution of signals by variants with unknown functions, our primary analysis of rare variants focused on nonsynonymous and splice-site variants. In secondary analyses, we limited the analysis to damaging variants, defined as nonsense variants or variants predicted to be damaging by PolyPhen [35] or SIFT [36].
For both common and rare variant analyses, models adjusted for age and sex, and stratified by ancestry (European or African American). ARIC and CHS additionally adjusted for their clinical sites, FHS accounted for family structure. The association analyses were performed using the R package seqMeta (http://cran.r-project.org/web/packages/seqMeta/). Each cohort provided single variant score tests as well as genotype covariance matrices for all variants. We meta-analyzed the individual-cohort results using the inverse-variance weighted fixed effects model in seqMeta. Bonferroni correction was used to adjust for multiple testing, and the significance was defined as 0.05/N, where N is the total number of tests.
Pathway analyses
Pathway analyses were used to investigate the collective effects of multiple genetic variants on AF risk. Each common variant was assigned a score to indicate its association with AF. The score was calculated as –log10(P-value), where the P-value was derived from the common variant test described above. The genetic variant was then mapped back to RefSeq genes (August 23, 2015). A gene score was defined as the highest score of variants within 110kb upstream and 40kb downstream of the gene’s most extreme transcript boundaries, which was anticipated to include the majority of cis-regulatory gene elements [37]. For rare variants, each gene was assigned a score equivalent to –log10(P-value), in which the P-value was derived from the SKAT test described previously.
We examined the enrichment of AF-related variants in an AF gene set comprised of 37 genes previously implicated in AF (S1 Table). Genes identified on the basis of GWAS results were selected on the basis of proximity to the AF susceptibility signal, biological literature supporting a putative functional role in AF pathogenesis, or using GRAIL [38]. Gene set enrichment analysis [39] was used to estimate the enrichment, and the significant gene sets were defined as those with P-value less than 0.05/3 = 0.017.
Supporting Information
Zdroje
1. Miyasaka Y, Barnes ME, Gersh BJ, Cha SS, Bailey KR, Abhayaratna WP, et al. Secular trends in incidence of atrial fibrillation in Olmsted County, Minnesota, 1980 to 2000, and implications on the projections for future prevalence. Circulation. 2006;114(2):119–25. 16818816.
2. Go AS, Hylek EM, Phillips KA, Chang Y, Henault LE, Selby JV, et al. Prevalence of diagnosed atrial fibrillation in adults: national implications for rhythm management and stroke prevention: the AnTicoagulation and Risk Factors in Atrial Fibrillation (ATRIA) Study. JAMA. 2001;285(18):2370–5. 11343485.
3. Kannel WB, Wolf PA, Benjamin EJ, Levy D. Prevalence, incidence, prognosis, and predisposing conditions for atrial fibrillation: population-based estimates. Am J Cardiol. 1998;82(8A):2N–9N. 9809895.
4. Ott A, Breteler MM, de Bruyne MC, van Harskamp F, Grobbee DE, Hofman A. Atrial fibrillation and dementia in a population-based study. The Rotterdam Study. Stroke. 1997;28(2):316–21. 9040682.
5. Wang TJ, Larson MG, Levy D, Vasan RS, Leip EP, Wolf PA, et al. Temporal relations of atrial fibrillation and congestive heart failure and their joint influence on mortality: the Framingham Heart Study. Circulation. 2003;107(23):2920–5. 12771006.
6. Krahn AD, Manfreda J, Tate RB, Mathewson FA, Cuddy TE. The natural history of atrial fibrillation: incidence, risk factors, and prognosis in the Manitoba Follow-Up Study. Am J Med. 1995;98(5):476–84. 7733127.
7. Stewart S, Hart CL, Hole DJ, McMurray JJ. A population-based study of the long-term risks associated with atrial fibrillation: 20-year follow-up of the Renfrew/Paisley study. Am J Med. 2002;113(5):359–64. 12401529.
8. Cappato R, Calkins H, Chen SA, Davies W, Iesaka Y, Kalman J, et al. Updated worldwide survey on the methods, efficacy, and safety of catheter ablation for human atrial fibrillation. CircArrhythmElectrophysiol. 2010;3(1):32–8. CIRCEP.109.859116 [pii];doi: 10.1161/CIRCEP.109.859116 19995881.
9. January CT, Wann LS, Alpert JS, Calkins H, Cleveland JC Jr., Cigarroa JE, et al. 2014 AHA/ACC/HRS Guideline for the Management of Patients With Atrial Fibrillation: A Report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines and the Heart Rhythm Society. Circulation. 2014. doi: 10.1161/CIR.0000000000000041 24682347.
10. Lubitz SA, Yin X, Fontes JD, Magnani JW, Rienstra M, Pai M, et al. Association between familial atrial fibrillation and risk of new-onset atrial fibrillation. JAMA. 2010;304(20):2263–9. Epub 2010/11/16. jama.2010.1690 [pii] doi: 10.1001/jama.2010.1690 21076174.
11. Gudbjartsson DF, Arnar DO, Helgadottir A, Gretarsdottir S, Holm H, Sigurdsson A, et al. Variants conferring risk of atrial fibrillation on chromosome 4q25. Nature. 2007;448(7151):353–7. 17603472.
12. Benjamin EJ, Rice KM, Arking DE, Pfeufer A, van Noord C, Smith AV, et al. Variants in ZFHX3 are associated with atrial fibrillation in individuals of European ancestry. Nat Genet. 2009;41(8):879–81. Epub 2009/07/15. ng.416 [pii] doi: 10.1038/ng.416 19597492.
13. Gudbjartsson DF, Holm H, Gretarsdottir S, Thorleifsson G, Walters GB, Thorgeirsson G, et al. A sequence variant in ZFHX3 on 16q22 associates with atrial fibrillation and ischemic stroke. Nat Genet. 2009;41(8):876–8. Epub 2009/07/15. ng.417 [pii] doi: 10.1038/ng.417 19597491.
14. Ellinor PT, Lunetta KL, G N.L., Pfeufer A, Alonso A, Chung MK, et al. Common Variants in KCNN3 are Associated with Lone Atrial Fibrillation Nat Genet. 2010;42(4):240–4. Epub 2010/02/23. ng.537 [pii] doi: 10.1038/ng.537 20173747.
15. Ellinor PT, Lunetta KL, Albert CM, Glazer NL, Ritchie MD, Smith AV, et al. Meta-analysis identifies six new susceptibility loci for atrial fibrillation. Nat Genet. 2012;44(6):670–5. Epub 2012/05/01. doi: 10.1038/ng.2261 22544366; PubMed Central PMCID: PMC3366038.
16. Tucker NR, Ellinor PT. Emerging directions in the genetics of atrial fibrillation. Circ Res. 2014;114(9):1469–82. doi: 10.1161/CIRCRESAHA.114.302225 24763465; PubMed Central PMCID: PMCPMC4040146.
17. Sinner MF, Lubitz SA, Pfeufer A, Makino S, Beckmann BM, Lunetta KL, et al. Lack of replication in polymorphisms reported to be associated with atrial fibrillation. Heart rhythm: the official journal of the Heart Rhythm Society. 2011;8(3):403–9. Epub 2010/11/09. doi: 10.1016/j.hrthm.2010.11.003 21056700; PubMed Central PMCID: PMC3068750.
18. Franke A, Balschun T, Sina C, Ellinghaus D, Hasler R, Mayr G, et al. Genome-wide association study for ulcerative colitis identifies risk loci at 7q22 and 22q13 (IL17REL). Nat Genet. 2010;42(4):292–4. doi: 10.1038/ng.553 20228798.
19. Sasaki MM, Skol AD, Hungate EA, Bao R, Huang L, Kahn SA, et al. Whole-exome Sequence Analysis Implicates Rare Il17REL Variants in Familial and Sporadic Inflammatory Bowel Disease. Inflamm Bowel Dis. 2016;22(1):20–7. doi: 10.1097/MIB.0000000000000610 26480299; PubMed Central PMCID: PMCPMC4679526.
20. Lohmueller KE, Sparso T, Li Q, Andersson E, Korneliussen T, Albrechtsen A, et al. Whole-exome sequencing of 2,000 Danish individuals and the role of rare coding variants in type 2 diabetes. Am J Hum Genet. 2013;93(6):1072–86. doi: 10.1016/j.ajhg.2013.11.005 24290377; PubMed Central PMCID: PMC3852935.
21. Weeke P, Muhammad R, Delaney JT, Shaffer C, Mosley JD, Blair M, et al. Whole-exome sequencing in familial atrial fibrillation. Eur Heart J. 2014;35(36):2477–83. doi: 10.1093/eurheartj/ehu156 24727801; PubMed Central PMCID: PMC4169871.
22. The Atherosclerosis Risk in Communities (ARIC) Study: design and objectives. The ARIC investigators. Am J Epidemiol. 1989;129(4):687–702. Epub 1989/04/01. 2646917.
23. Fried LP, Borhani NO, Enright P, Furberg CD, Gardin JM, Kronmal RA, et al. The Cardiovascular Health Study: design and rationale. Annals of epidemiology. 1991;1(3):263–76. Epub 1991/02/01. 1669507.
24. Feinleib M, Kannel WB, Garrison RJ, McNamara PM, Castelli WP. The Framingham Offspring Study. Design and preliminary data. Prev Med. 1975;4(4):518–25. Epub 1975/12/01. 1208363.
25. Kannel WB, Dawber TR, Kagan A, Revotskie N, Stokes J 3rd. Factors of risk in the development of coronary heart disease—six year follow-up experience. The Framingham Study. Ann Intern Med. 1961;55 : 33–50. Epub 1961/07/01. 13751193.
26. Fu W, O'Connor TD, Jun G, Kang HMAbecasis G, Leal SM, et al. Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature. 2013;493(7431):216–20. doi: 10.1038/nature11690 23201682; PubMed Central PMCID: PMC3676746.
27. Psaty BM, O'Donnell CJ, Gudnason V, Lunetta KL, Folsom AR, Rotter JI, et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: Design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ Cardiovasc Genet. 2009;2(1):73–80. doi: 10.1161/CIRCGENETICS.108.829747 20031568; PubMed Central PMCID: PMC2875693.
28. Reid JG, Carroll A, Veeraraghavan N, Dahdouli M, Sundquist A, English A, et al. Launching genomics into the cloud: deployment of Mercury, a next generation sequence analysis pipeline. BMC bioinformatics. 2014;15 : 30. Epub 2014/01/31. doi: 10.1186/1471-2105-15-30 24475911; PubMed Central PMCID: PMC3922167.
29. Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–60. doi: 10.1093/bioinformatics/btp324 19451168; PubMed Central PMCID: PMC2705234.
30. Challis D, Yu J, Evani US, Jackson AR, Paithankar S, Coarfa C, et al. An integrative variant analysis suite for whole exome next-generation sequencing data. BMC Bioinformatics. 2012;13 : 8. doi: 10.1186/1471-2105-13-8 22239737; PubMed Central PMCID: PMC3292476.
31. Reiner AP, Beleza S, Franceschini N, Auer PL, Robinson JG, Kooperberg C, et al. Genome-wide association and population genetic analysis of C-reactive protein in African American and Hispanic American women. Am J Hum Genet. 2012;91(3):502–12. doi: 10.1016/j.ajhg.2012.07.023 22939635; PubMed Central PMCID: PMC3511984.
32. Tennessen JA, Bigham AW, O'Connor TD, Fu W, Kenny EE, Gravel S, et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science. 2012;337(6090):64–9. doi: 10.1126/science.1219240 22604720.
33. Lange LA, Hu Y, Zhang H, Xue C, Schmidt EM, Tang ZZ, et al. Whole-exome sequencing identifies rare and low-frequency coding variants associated with LDL cholesterol. Am J Hum Genet. 2014;94(2):233–45. doi: 10.1016/j.ajhg.2014.01.010 24507775; PubMed Central PMCID: PMC3928660.
34. Wu MC, Lee S, Cai T, Li Y, Boehnke M, Lin X. Rare-variant association testing for sequencing data with the sequence kernel association test. American journal of human genetics. 2011;89(1):82–93. Epub 2011/07/09. doi: 10.1016/j.ajhg.2011.05.029 21737059; PubMed Central PMCID: PMC3135811.
35. Adzhubei I, Jordan DM, Sunyaev SR. Predicting functional effect of human missense mutations using PolyPhen-2. Current protocols in human genetics / editorial board, Jonathan L Haines [et al]. 2013;Chapter 7:Unit7 20. doi: 10.1002/0471142905.hg0720s76 23315928; PubMed Central PMCID: PMC4480630.
36. Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4(7):1073–81. Epub 2009/06/30. nprot.2009.86 [pii] doi: 10.1038/nprot.2009.86 19561590.
37. Segre AV, Consortium D, investigators M, Groop L, Mootha VK, Daly MJ, et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 2010;6(8). Epub 2010/08/18. doi: 10.1371/journal.pgen.1001058 20714348; PubMed Central PMCID: PMC2920848.
38. Raychaudhuri S, Plenge RM, Rossin EJ, Ng AC, International Schizophrenia C, Purcell SM, et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 2009;5(6):e1000534. doi: 10.1371/journal.pgen.1000534 19557189; PubMed Central PMCID: PMCPMC2694358.
39. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–50. Epub 2005/10/04. 0506580102 [pii] doi: 10.1073/pnas.0506580102 16199517; PubMed Central PMCID: PMC1239896.
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2016 Číslo 9- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
Nejčtenější v tomto čísle- Reactive Oxygen Species: From Harmful Molecules to Fine-Tuning Regulators of Stem Cell Niche Maintenance
- The Clock Is Ticking: Countdown to Metastases
- Whole Exome Sequencing in Atrial Fibrillation
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



