-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Transposable Elements Re-Wire and Fine-Tune the Transcriptome
What good are transposable elements (TEs)? Although their activity can be harmful to host genomes and can cause disease, they nevertheless represent an important source of genetic variation that has helped shape genomes. In this review, we examine the impact of TEs, collectively referred to as the mobilome, on the transcriptome. We explore how TEs—particularly retrotransposons—contribute to transcript diversity and consider their potential significance as a source of small RNAs that regulate host gene transcription. We also discuss a critical role for the mobilome in engineering transcriptional networks, permitting coordinated gene expression, and facilitating the evolution of novel physiological processes.
Published in the journal: . PLoS Genet 9(1): e32767. doi:10.1371/journal.pgen.1003234
Category: Review
doi: https://doi.org/10.1371/journal.pgen.1003234Summary
What good are transposable elements (TEs)? Although their activity can be harmful to host genomes and can cause disease, they nevertheless represent an important source of genetic variation that has helped shape genomes. In this review, we examine the impact of TEs, collectively referred to as the mobilome, on the transcriptome. We explore how TEs—particularly retrotransposons—contribute to transcript diversity and consider their potential significance as a source of small RNAs that regulate host gene transcription. We also discuss a critical role for the mobilome in engineering transcriptional networks, permitting coordinated gene expression, and facilitating the evolution of novel physiological processes.
Introduction
The 1983 Nobel Prize for Physiology or Medicine was awarded to Barbara McClintock for her seminal discovery of transposable elements (TEs). McClintock's studies of colour patterns in maize kernels led her to conclude that “controlling elements” that could jump around the genome regulate gene expression (reviewed in [1]). Although the response of the scientific community to her work was initially cautious, the discovery of similar elements in flies, bacteria, and yeast underlined its significance. Today, TEs are recognised as important components of genomes that have helped shape their evolution.
Approximately half of the human genome is derived from TEs [2], although recent work suggests this may be closer to two-thirds [3]. Most human TEs are retrotransposons, and some are still active today (Box 1 and Figure 1). Consequently, TEs represent a significant source of genetic variation [4]–[7].
Fig. 1. Active human retrotransposons. 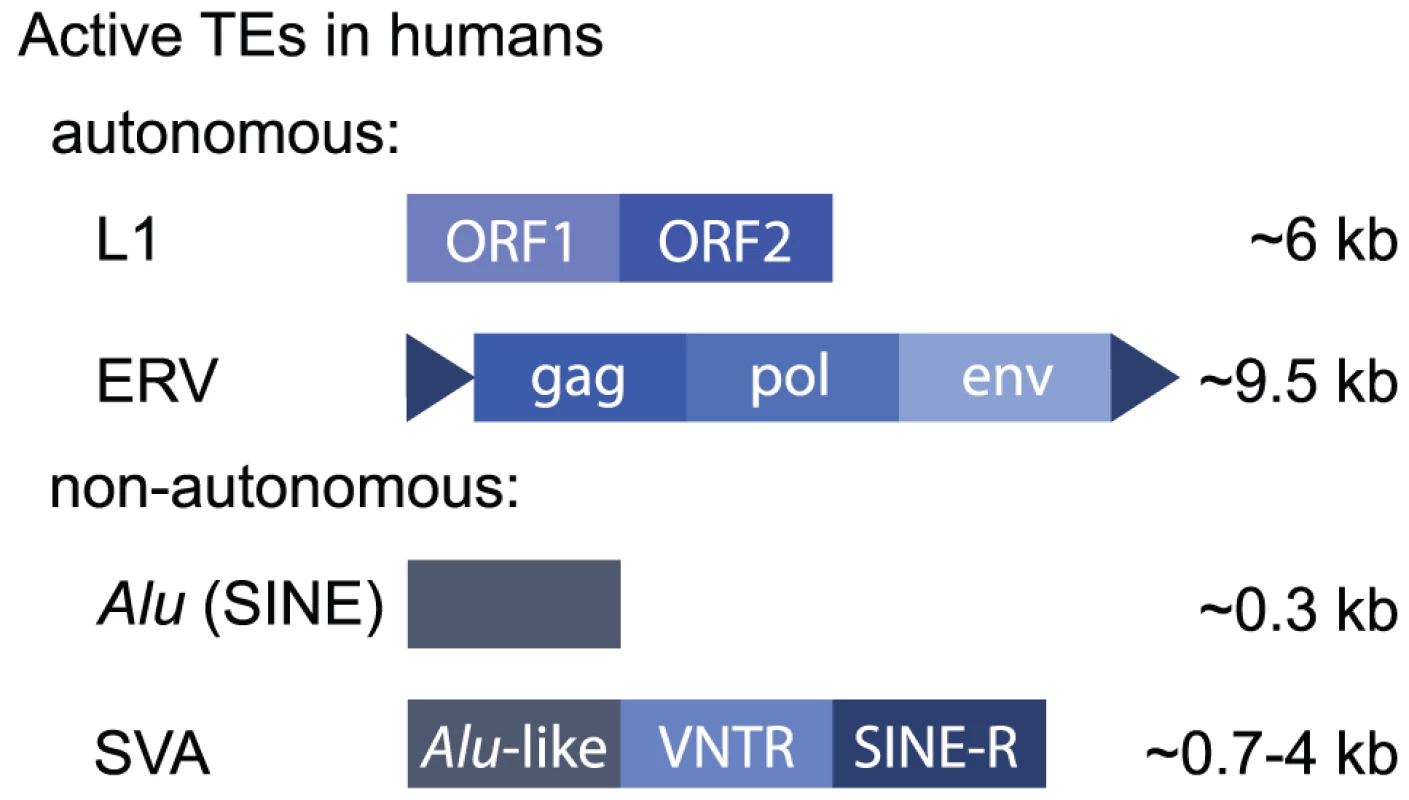
Box 1. The Human Mobilome
Human TEs may be classified as retrotransposons, which replicate using an mRNA intermediate to “copy and paste,” or DNA transposons, which transpose using a DNA intermediate. Retrotransposons constitute ∼42% of the human genome [2], and some elements are still active today, meaning they are still capable of retrotransposing. DNA transposons represent ∼3% of the human genome but are no longer transposition-competent. Retrotransposons can be further classified based on their structure. LTR elements are characterised by long terminal repeats and include endogenous retroviruses (ERVs) that encode gag, pol, and env genes. These evolved as a consequence of retroviral infection of germ cells, so that they are inherited through generations (reviewed in [77]). Other LTRs are “solo” LTRs, meaning they exist alone. These result from a recombination event that deletes the intervening retroviral genes [78]. Non-LTR retrotransposons include long and short interspersed repeat elements (LINEs and SINEs) and SVA elements that are a composite of sequences derived from other repeats (SINE, VNTR [variable number tandem repeat], and Alu). There are only three highly active elements in the human genome: (1) a subset of L1 LINEs, (2) Alu elements that are a family of primate-specific SINEs, and (3) SVA elements (Figure 1). Alu elements are the most active, with approximately one de novo germline insertion per 20 births [79]. De novo insertions of L1 and SVA elements occur at the rate of approximately 1 in 108 births and 1 in 916 births, respectively [4], [80]. Some ERVs may still be active in humans [81], although the vast majority are nonfunctional, in contrast to their relatively high levels of activity in mice. Only L1 elements encode the enzymes required for retrotransposition, and these preferentially (but not exclusively) recognise L1 mRNA molecules. Alu and SVA elements co-opt the L1 machinery to retrotranspose. The mechanism of retrotransposition has been expertly reviewed elsewhere recently [82], [83].
How might TEs influence gene expression? It is easy to imagine how an insertion into a gene might disrupt an open reading frame (ORF), preventing the synthesis of a protein (Figure 2). Indeed, examples of human diseases caused in this manner have been reported [8], [9]. However, the impact of an insertion may not be so dramatic or deleterious. TEs can influence host genes by providing novel promoters, splice sites, or polyadenylation signals (Figure 2). An important consequence is the generation of transcript diversity. There are many more different mRNA molecules in the human transcriptome than the 20,000 protein-coding genes in the genome, and this transcript diversity is thought to be key for promoting phenotypic diversity in higher eukaryotes [10], [11]. Additionally, genome-scale studies have revealed the importance of TEs in dispersing transcription factor binding sites, linking genes in transcriptional networks (e.g., [12], [13]), and facilitating the evolution of novel traits.
Fig. 2. How the mobilome can impact the transcriptome. 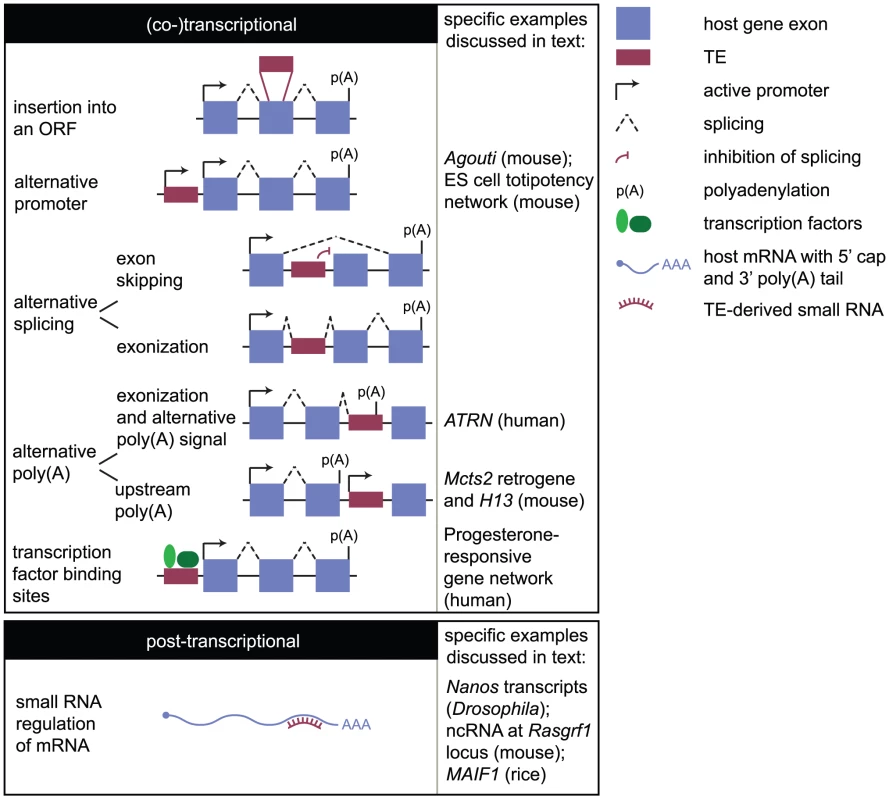
Impacts on the transcriptome may be considered transcriptional (or co-transcriptional) and posttranscriptional. The former mechanisms include insertion of a TE into an ORF; provision of an alternative promoter that may be tissue- or stage-specific in its activity; promotion of alternative splicing either through prevention of the splicing machinery from recognising a splice acceptor site in an endogenous exon (exon skipping) or through incorporation of the TE into the mature transcript (exonization); promotion of alternative polyadenylation (poly(A)) either by providing an alternative polyadenylation signal or by promoter activity interfering with host gene transcription and causing upstream polyadenylation; and by introducing transcription factor binding sites that may confer tissue- or stage-specific expression, or link a gene into a transcriptional network. Posttranscriptional regulation involves TE-derived small RNAs binding to host transcripts. In the case of Drosophila Nanos transcripts, small RNAs destabilise the transcript by recruiting the deadenylation machinery. In the case of murine Rasgrf1, the binding of small RNAs to an ncRNA associated with one allele results in the recruitment of the de novo methylation machinery to that allele, causing allele-specific Rasgrf1 expression. The events occurring downstream of small RNA binding are therefore diverse and locus-specific. In this review, we consider how TEs, collectively referred to as the mobilome, have impacted the transcriptome. This includes elements active today as well as those no longer transposition-competent. Although not nearly an exhaustive account, we draw on specific examples from a range of organisms to illustrate the variety of mechanisms through which this can occur. We aim to highlight the importance of the mobilome in shaping both the diversity and regulation of the transcriptome.
Generating Transcriptome Diversity
One surprising finding from sequencing the human genome was that humans have a similar number of genes to the model nematode Caenorhabditis elegans: about 20,000 and 19,000, respectively. This was unexpected because of the apparent complexity of humans relative to nematodes; indeed, earlier estimates for the number of human genes ranged from 60,000–150,000. Several factors may account for this discrepancy [14], but one important consideration is alternative mRNA processing. This includes alternative splicing and polyadenylation, enabling multiple mRNA species to be generated from a single gene. These mRNA isoforms can encode proteins with different functions or may be differentially regulated. More than 95% of human multi-exonic genes are alternatively spliced [15], while this is around 25% in C. elegans [16].
TEs, particularly L1 and Alu elements, can introduce novel splice sites [17], [18]. Indeed, Alu elements inserted into a gene can provide both splice acceptor and donor sites, creating new exons [19], [20]. Moreover, most Alu-derived exons are alternatively spliced [21], contributing to transcript diversity. They are enriched in the 5′ untranslated regions of human genes, where they regulate mRNA translation [22]. Furthermore, many alternative splicing events of Alu-derived exons are tissue-specific, suggesting TEs contribute to the transcriptome differences that define cell types [23]–[25]. Polyadenylation stabilises mRNA transcripts and influences nuclear export and translation efficiency. The majority of human genes utilise alternative polyadenylation sites [26], and the signals for some of these are embedded in TEs [27], suggesting TEs can influence the 3′ end processing of host gene transcripts.
The human ATRN gene provides a good example of how TE-induced alternative mRNA processing can enable functional diversification of one gene. A subset of ATRN transcripts are cleaved and polyadenylated within an L1 element that has retrotransposed into an intron (Figure 3A) [28]. Other transcripts splice around the L1 element and incorporate an additional five exons. Transcripts polyadenylated within the L1 element encode a soluble form of Attractin that is released by activated T lymphocytes as part of the basic inflammatory response [29]. The alternative transcripts encode a protein with transmembrane and cytoplasmic domains that is membrane-bound. This isoform is similar to murine Atrn, which functions as a receptor involved in pigmentation and energy metabolism [28], [30], [31]. This is a clear example of how a single retrotransposition event can increase transcript diversity with direct consequences on cellular function.
Fig. 3. Retrotransposition can influence mRNA processing. 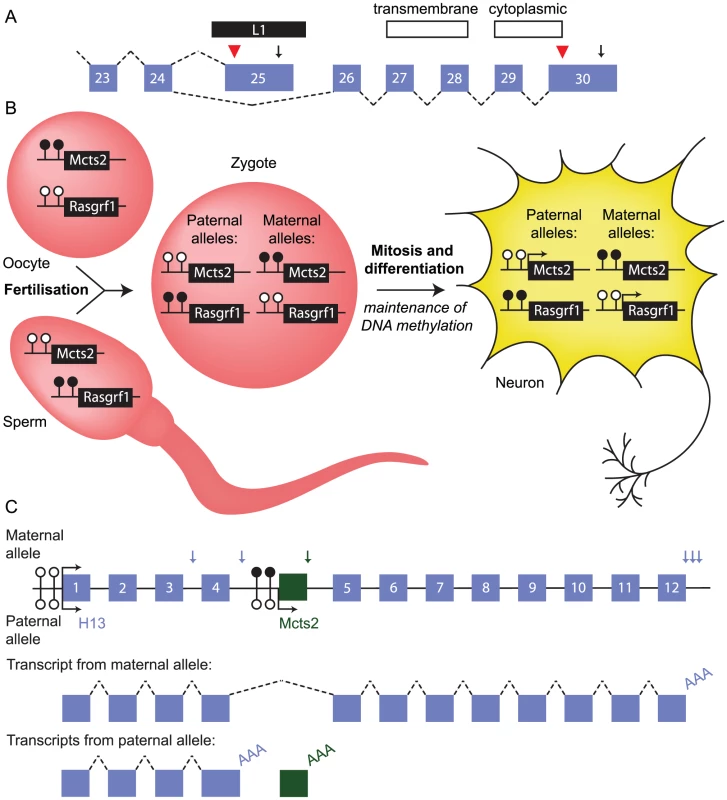
(A) Schematic of the 3′ end of the human ATRN gene. An L1 element (black bar) inserted between exons 24 and 26 (numbered boxes) provides a terminal exon, translation termination site (red arrowhead), and polyadenylation signal (arrow) for a subset of transcripts. Alternative splicing produces an mRNA isoform that is polyadenylated in exon 30; only this isoform encodes transmembrane and cytoplasmic domains. Dashed lines, splicing event. (B) Inheritance of DNA methylation at the imprinted Mcts2 and Rasgrf1 genes in mouse. The promoter of Mcts2 is methylated (filled lollipops) in the oocyte and unmethylated (empty lollipops) in sperm. This is opposite to the Rasgrf1 promoter. After fertilisation, these differences persist, marking the origin of the parental alleles even in terminally differentiated cell types, where the unmethylated promoters are transcriptionally active (arrows). (C) Relationship between the retrogene Mcts2 and the gene H13. (Top) Locus structure. Mcts2 (green box) is situated between exons 4 and 5 of H13. Allele-specific differences in methylation at the Mcts2 promoter result in expression of Mcts2 from the paternal allele only. The H13 promoter is unmethylated and active on both alleles. H13 transcripts use alternative polyadenylation sites (vertical blue arrows). Vertical green arrow, single Mcts2 polyadenylation site. (Middle) Representative transcript produced from transcription of the maternal allele. H13 transcripts splice around Mcts2 and use one of three downstream polyadenylation signals (one transcript is shown for clarity). (Bottom) Representative transcripts produced from transcription of the paternal allele. Mcts2 is transcribed and the mRNA is polyadenylated (AAA). H13 transcripts use one of two upstream polyadenylation signals (one transcript is shown for clarity). Transcription of the retrogene Mcts2 is associated with upstream polyadenylation of H13 transcripts. mRNA processing can also be impacted by TEs indirectly. Although the L1-encoded retrotransposition enzymes preferentially recognise L1 mRNA molecules, host protein-coding mRNAs can also be retrotransposed by the L1 machinery, creating a copy of the original gene [32]. In most cases the copy is nonfunctional, but it can potentially evolve into a retrogene with a novel function or expression pattern. About 120 retroposed sequences have evolved into bona fide genes in the human genome [33]. Retrogenes embedded in introns of other genes can influence transcription of that gene, causing upstream transcript polyadenylation. This is a mechanism through which TEs can indirectly influence mRNA processing, further promoting transcript diversity.
In some cases, the impact of the retrogene on host mRNA processing is specific to only one of the two parental alleles [34], [35]. For example, the retrogene Mcts2 is embedded in an intron of H13. Mcts2 is subject to genomic imprinting. It is expressed exclusively from the allele inherited through the paternal line because its promoter is silenced on the maternally inherited allele by DNA methylation (Figure 3B). Silencing of Mcts2 on the maternally inherited allele permits transcription of H13 to continue through the retrogene, which is spliced out of mature transcripts, and downstream polyadenylation sites are used (Figure 3C). Mcts2 transcription on the paternally inherited allele is associated with H13 transcripts using upstream polyadenylation sites. This is not a consequence of introducing alternative polyadenylation signals, but may involve the elongation complexes that are transcribing H13 “crashing” into those at the transcribing retrogene, a process termed “transcriptional interference” [36]. This interference may promote H13 transcript cleavage and polyadenylation. Intronic ERVs may influence transcription in a similar manner at many loci, impacting on the levels of protein produced from the endogenous gene [37], [38].
TEs further promote transcript diversity by providing alternative promoters for host genes. Perhaps one of the most elegant examples of this can be found in viable yellow agouti (Avy) mice, in which an ERV intracisternal A particle (IAP) upstream of the Agouti gene provides an alternative promoter than can drive ectopic Agouti expression, producing yellow fur [39]. Although the overt phenotype means this is a well-studied specific example, high-throughput approaches have demonstrated widespread use of TEs as alternative gene promoters in normal tissues, and these contribute to tissue-specific expression profiles [24], [40]. Inappropriate activation of promoters embedded in TEs, perhaps resulting from the relaxation of repressive epigenetic marks, can drive ectopic gene expression, and this mechanism has been implicated in human diseases, including cancer [24], [41].
These examples illustrate the impacts of retrotransposition events that occurred in the germline and are stably inherited. Recently there has been much debate about the extent and significance of somatic retrotransposition. L1 expression increases as neural stem cells commit to a neuronal lineage, and this was reported to be associated with elevated L1 retrotransposition [42]. The proposed outcome of this is increased transcriptome heterogeneity among neurons, contributing to interindividual variation [43]. However, the true extent of this mechanism in vivo is debated, with recent estimates ranging from ∼80 to fewer than 0.6 somatic L1 insertions per neuron in the human brain [44], [45], depending on the experimental approach used. Additional work will be required to confirm the true significance of this mechanism for promoting somatic transcriptome diversity. However, the generation of transcript diversity by alternative mRNA processing and the utilisation of alternative promoters in the genome inherited through the germline is likely to be a key factor in the evolution of higher eukaryotes. The mobilome has played a significant role in this process, both directly, by introducing regulatory sequences, and indirectly, by interfering with host gene transcription.
The Mobilome as a Source of Small Regulatory RNAs
Mcts2 is one example of ∼150 genes that are subject to genomic imprinting in the mouse. Although the mechanisms for regulating imprinting vary between loci, silencing of one allele by DNA methylation is a common theme. DNA methylation is likely to have evolved initially as a host defence mechanism against TE expression. Indeed, male mouse germ cells lacking the de novo methyltransferase Dnmt3L exhibit elevated expression of L1 elements and IAPs, resulting in meiotic catastrophe [46]. Thus, the need to minimise the impacts of retrotransposons may have driven the evolution of a novel mode of gene regulation—genomic imprinting—that is critical for mammalian development ([47] and reviewed in [48], [49]).
In addition to DNA methylation, small regulatory RNAs, including PIWI-interacting RNAs (piRNAs) and small interfering RNAs (siRNAs), may also have evolved as a host defence mechanism, repressing the translation of TE transcripts or promoting their decay. Like DNA methylation, this mechanism has been adopted by the host to regulate endogenous genes. Indeed, TEs are major players in the production of small RNAs that regulate host gene transcripts. For example, TE-encoded piRNAs are required to establish a gradient of maternal Nanos mRNA transcripts in the early Drosophila embryo [50]. This is achieved by the binding of piRNAs to a specific sequence in the 3′ untranslated region of Nanos transcripts, which promotes removal of the polyadenylate tail and transcript degradation. This process is essential for establishing correct anterior-posterior patterning in the embryo.
Mammals also utilise TE-encoded small RNAs to regulate host gene expression. In mice, the imprinted gene Rasgrf1, like Mcts2, is under the control of differential DNA methylation on the two alleles. At this locus, DNA methylation is established in the paternal germline (Figure 3B), opposite to that for Mcts2, and this requires TE-encoded piRNAs [51]. During spermatogenesis, these piRNAs bind to a noncoding (nc)RNA transcribed from the locus; specifically, they recognise an LTR-type retrotransposon RMER4B embedded within the ncRNA. Targeting of this ncRNA results in recruitment of the de novo methylation machinery, through an unknown mechanism. Disruption of the piRNA pathway or expression of the ncRNA perturbs methylation and imprinting of Rasgrf1, and mice defective for Rasgrf1 imprinting exhibit impaired postnatal growth [52].
These examples from Drosophila and mice demonstrate the importance of TEs as a source of small RNAs for regulating host transcripts. As such, the host depends upon TEs to provide these regulatory molecules, illustrating their intimate relationship. This relationship is not exclusive to animals, with plants utilising the same system to fine-tune gene expression. For example, in rice, siRNAs originating from the miniature inverted-repeat TE (MITE) Stowaway1 regulate tolerance to abiotic stress [53]. These siRNAs may function by targeting transcripts of the growth regulator MAIF1 and stalling growth, a common physiological response in plants to abiotic stresses.
A defensive response to TEs is critical to guard against uncontrolled transposition. Hosts have evolved several mechanisms for tackling this, including transcriptional repression by DNA methylation and posttranscriptional repression by small RNAs. The evolution of these systems has dramatically impacted the transcriptome because they have been adopted for more general gene regulation.
TEs as Engineers of Transcriptional Networks
TEs can influence alternative mRNA processing or generate small regulatory RNAs, but these effects on the transcriptome are locus-specific. How is transcription regulated on a global scale, say in response to an environmental cue? In the yeast Saccharomyces cerevisiae, genes involved in common metabolic pathways are physically clustered in the genome [54], [55], permitting co-ordinated expression [56] and tight gene regulation in response to a stimulus. However, not all genes that must be co-ordinately expressed are physically clustered. In higher eukaryotes, the binding of transcription factors can co-ordinately activate the expression of genes dispersed throughout the genome. Genes linked in this manner can be considered part of a single transcriptional network. The mobilome has been vital for linking genes in this manner. Regulatory elements required for TE expression can be co-opted by endogenous genes, or the TE may harbour transcription factor binding sites (TFBSs; Figure 2) [12], [13], [57], [58]. TE-derived TFBSs evolve rapidly relative to non-repeat-derived sites [59], suggesting they are important drivers in conferring species-specific gene expression profiles.
A good example of the importance of TEs in linking genes in a network can be found in embryonic stem (ES) cells. ES cells are pluripotent but can enter a transient phase of totipotency from which they can generate both embryonic and extra-embryonic lineages [60]. This switch depends on the activation of a network of transcripts that initiate within ERV LTRs and is controlled by epigenetic modifications. In the pluripotent state, ERVs are transcriptionally repressed, in part by histone 3 lysine 9 methylation [61]. This is established by the histone methyltransferase SETDB1 that is recruited to ERVs by KAP1 [62], [63]. ES cells deficient for Kap1 switch more readily to the totipotent state, consistent with the idea that relaxation of ERV repression drives network activation [60]. These studies highlight a critical role for ERVs in contributing to host cell fate decisions by activating a transcriptional network. This is mediated by epigenetic marks that are established and removed by endogenous cellular machinery.
Earlier, we discussed how a TE could contribute to the evolution of novel functions at a single gene, such as human ATRN. However, by re-wiring networks, the mobilome can facilitate the evolution of complex physiological processes involving gene expression on a global scale. The evolution of pregnancy, the trait that defines mammals, is an intriguing example of this. The hormone progesterone triggers the differentiation of endometrial stromal cells to form the decidua, the maternal component of the placenta [64]. This relies on the activation of a network of transcripts linked by MER20 elements that provide binding sites for progesterone-responsive signalling molecules [65].
An important progesterone-responsive gene is prolactin. In addition to being linked in this network by MER20, the promoter for human prolactin is derived from an independent TE, MER39 [66]. This TE is primate-specific, yet other mammals activate prolactin expression during pregnancy. Emera et al. [67] demonstrated that the endometrial stromal cell-specific promoters of human, mouse, and elephant prolactin are all distinct and are all derived from different TEs (MER77 for mouse, L1 for elephant), suggesting TEs can contribute to convergent evolution. Similarly, the syncytin genes, essential for formation of the syncytiotrophoblast layer that mediates fetal-maternal exchange, are derived from ERV env genes and have been independently acquired in the human, mouse, and rabbit genomes [68].
Other aspects of the physiological changes required for pregnancy may have evolved by TEs re-wiring networks, such as the tolerance of the maternal immune system to a fetus expressing paternal antigens [69]. Together, these examples illustrate the requirement for TEs in pregnancy: engineering a transcriptional network, providing cell type-specific promoters, and contributing to gene function. Thus, the impact of TEs can extend well beyond single-locus effects, making vital contributions to the evolution of complex physiological processes. This role is not confined to animals; TEs in plants have had similar impacts [70].
Conclusions and Future Perspectives
The impacts of TEs on the host transcriptome are diverse. At the single locus level, a transposition event may result in an alternatively processed transcript that can evolve a new function. At the genome level, TEs may disperse regulatory elements that rewire transcriptional networks. The significance of TEs is becoming increasingly apparent with more widespread application of next-generation sequencing technologies. At first, repetitive elements were a nuisance in the analysis of genome-wide datasets; now new experimental protocols and computational pipelines are being utilised to ask questions specifically about TE distribution [71], [72]. The 1000 Genomes Project will provide a valuable tool for interrogating the extent and functional importance of insertional polymorphisms in humans, and indeed has already yielded some intriguing findings; for example, the insertion rates of TEs differ between populations [73].
Many important questions remain unanswered. For example, what is the extent and biological significance of somatic retrotransposition? What is the contribution of this mechanism to the transcriptome differences between neurons, and how does this influence behaviour? Additionally, inappropriate activation of TEs has been associated with somatic cancers [71]. It is too early to say if this mechanism is truly causative, but the current data are provocative. Another exciting area with important outstanding questions is the influence of epigenetic silencing of TEs on the host. For example, could methylated TEs act as “messengers,” transmitting epigenetic information between generations? During primordial germ cell development, most DNA methylation is erased and reset, but a small fraction of the genome is resistant to erasure. This includes IAPs, suggesting these are candidates for mediating transgenerational epigenetic inheritance [74], [75]. Indeed, such a role has been demonstrated for at least two specific IAPs [39], [76], but whether this represents a more global mechanism is undetermined.
Are the evolutionary benefits conferred by TEs purely accidental? Most point mutations arising in the germline have a deleterious or neutral effect on the host, but some do introduce innovative changes that are beneficial. Likewise, TE insertions may be deleterious but can also provide opportunities for increasing transcript diversity or rewiring the transcriptome. Such advantageous insertion events can be selected for and fixed in a population. This “fine-tuning” of the transcriptome could explain why organisms have evolved mechanisms to regulate TE activity without completely silencing all types.
Our understanding of the diverse impacts of the mobilome on the transcriptome has come a long way since the finding that TEs could cause insertional mutations leading to disease. TEs have been fundamental players in evolution and are intimately associated with the regulation of host gene transcription.
Zdroje
1. FedoroffN (2001) How jumping genes were discovered. Nat Struct Biol 8 : 300–301.
2. LanderES, LintonLM, BirrenB, NusbaumC, ZodyMC, et al. (2001) Initial sequencing and analysis of the human genome. Nature 409 : 860–921.
3. de KoningAP, GuW, CastoeTA, BatzerMA, PollockDD (2011) Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet 7: e1002384 doi:10.1371/journal.pgen.1002384.
4. HuangCR, SchneiderAM, LuY, NiranjanT, ShenP, et al. (2010) Mobile interspersed repeats are major structural variants in the human genome. Cell 141 : 1171–1182.
5. BeckCR, CollierP, MacfarlaneC, MaligM, KiddJM, et al. (2010) LINE-1 retrotransposition activity in human genomes. Cell 141 : 1159–1170.
6. EwingAD, KazazianHHJr (2010) High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res 20 : 1262–1270.
7. IskowRC, McCabeMT, MillsRE, ToreneS, PittardWS, et al. (2010) Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 141 : 1253–1261.
8. KazazianHHJr, WongC, YoussoufianH, ScottAF, PhillipsDG, et al. (1988) Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 332 : 164–166.
9. ChenJM, StensonPD, CooperDN, FerecC (2005) A systematic analysis of LINE-1 endonuclease-dependent retrotranspositional events causing human genetic disease. Hum Genet 117 : 411–427.
10. GraveleyBR (2001) Alternative splicing: increasing diversity in the proteomic world. Trends Genet 17 : 100–107.
11. NilsenTW, GraveleyBR (2010) Expansion of the eukaryotic proteome by alternative splicing. Nature 463 : 457–463.
12. BourqueG, LeongB, VegaVB, ChenX, LeeYL, et al. (2008) Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res 18 : 1752–1762.
13. SchmidtD, SchwaliePC, WilsonMD, BallesterB, GoncalvesA, et al. (2012) Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell 148 : 335–348.
14. HodgkinJ (2001) What does a worm want with 20,000 genes? Genome Biol 2: COMMENT2008.
15. PanQ, ShaiO, LeeLJ, FreyBJ, BlencoweBJ (2008) Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet 40 : 1413–1415.
16. RamaniAK, CalarcoJA, PanQ, MavandadiS, WangY, et al. (2011) Genome-wide analysis of alternative splicing in Caenorhabditis elegans. Genome Res 21 : 342–348.
17. LiWH, GuZ, WangH, NekrutenkoA (2001) Evolutionary analyses of the human genome. Nature 409 : 847–849.
18. NekrutenkoA, LiWH (2001) Transposable elements are found in a large number of human protein-coding genes. Trends Genet 17 : 619–621.
19. Lev-MaorG, SorekR, ShomronN, AstG (2003) The birth of an alternatively spliced exon: 3′ splice-site selection in Alu exons. Science 300 : 1288–1291.
20. SelaN, MerschB, Gal-MarkN, Lev-MaorG, Hotz-WagenblattA, et al. (2007) Comparative analysis of transposed element insertion within human and mouse genomes reveals Alu's unique role in shaping the human transcriptome. Genome Biol 8: R127.
21. SorekR, AstG, GraurD (2002) Alu-containing exons are alternatively spliced. Genome Res 12 : 1060–1067.
22. ShenS, LinL, CaiJJ, JiangP, KenkelEJ, et al. (2011) Widespread establishment and regulatory impact of Alu exons in human genes. Proc Natl Acad Sci U S A 108 : 2837–2842.
23. LinL, ShenS, TyeA, CaiJJ, JiangP, et al. (2008) Diverse splicing patterns of exonized Alu elements in human tissues. PLoS Genet 4: e1000225 doi:10.1371/journal.pgen.1000225.
24. FaulknerGJ, KimuraY, DaubCO, WaniS, PlessyC, et al. (2009) The regulated retrotransposon transcriptome of mammalian cells. Nat Genet 41 : 563–571.
25. DjebaliS, DavisCA, MerkelA, DobinA, LassmannT, et al. (2012) Landscape of transcription in human cells. Nature 489 : 101–108.
26. DertiA, Garrett-EngeleP, MacisaacKD, StevensRC, SriramS, et al. (2012) A quantitative atlas of polyadenylation in five mammals. Genome Res 22 : 1173–1183.
27. LeeJY, JiZ, TianB (2008) Phylogenetic analysis of mRNA polyadenylation sites reveals a role of transposable elements in evolution of the 3′-end of genes. Nucleic Acids Res 36 : 5581–5590.
28. TangW, GunnTM, McLaughlinDF, BarshGS, SchlossmanSF, et al. (2000) Secreted and membrane attractin result from alternative splicing of the human ATRN gene. Proc Natl Acad Sci U S A 97 : 6025–6030.
29. Duke-CohanJS, GuJ, McLaughlinDF, XuY, FreemanGJ, et al. (1998) Attractin (DPPT-L), a member of the CUB family of cell adhesion and guidance proteins, is secreted by activated human T lymphocytes and modulates immune cell interactions. Proc Natl Acad Sci U S A 95 : 11336–11341.
30. NagleDL, McGrailSH, VitaleJ, WoolfEA, DussaultBJJr, et al. (1999) The mahogany protein is a receptor involved in suppression of obesity. Nature 398 : 148–152.
31. GunnTM, MillerKA, HeL, HymanRW, DavisRW, et al. (1999) The mouse mahogany locus encodes a transmembrane form of human attractin. Nature 398 : 152–156.
32. EsnaultC, MaestreJ, HeidmannT (2000) Human LINE retrotransposons generate processed pseudogenes. Nat Genet 24 : 363–367.
33. VinckenboschN, DupanloupI, KaessmannH (2006) Evolutionary fate of retroposed gene copies in the human genome. Proc Natl Acad Sci U S A 103 : 3220–3225.
34. WoodAJ, SchulzR, WoodfineK, KoltowskaK, BeecheyCV, et al. (2008) Regulation of alternative polyadenylation by genomic imprinting. Genes Dev 22 : 1141–1146.
35. CowleyM, WoodAJ, BohmS, SchulzR, OakeyRJ (2012) Epigenetic control of alternative mRNA processing at the imprinted Herc3/Nap1l5 locus. Nucleic Acids Res 40 : 8917–8926.
36. ShearwinKE, CallenBP, EganJB (2005) Transcriptional interference–a crash course. Trends Genet 21 : 339–345.
37. LiJ, AkagiK, HuY, TrivettAL, HlynialukCJ, et al. (2012) Mouse endogenous retroviruses can trigger premature transcriptional termination at a distance. Genome Res 22 : 870–884.
38. DrukerR, BruxnerTJ, LehrbachNJ, WhitelawE (2004) Complex patterns of transcription at the insertion site of a retrotransposon in the mouse. Nucleic Acids Res 32 : 5800–5808.
39. MorganHD, SutherlandHG, MartinDI, WhitelawE (1999) Epigenetic inheritance at the agouti locus in the mouse. Nat Genet 23 : 314–318.
40. PeastonAE, EvsikovAV, GraberJH, de VriesWN, HolbrookAE, et al. (2004) Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev Cell 7 : 597–606.
41. WolffEM, ByunHM, HanHF, SharmaS, NicholsPW, et al. (2010) Hypomethylation of a LINE-1 promoter activates an alternate transcript of the MET oncogene in bladders with cancer. PLoS Genet 6: e1000917 doi:10.1371/journal.pgen.1000917.
42. MuotriAR, ChuVT, MarchettoMC, DengW, MoranJV, et al. (2005) Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature 435 : 903–910.
43. ThomasCA, PaquolaAC, MuotriAR (2012) LINE-1 Retrotransposition in the Nervous System. Annu Rev Cell Dev Biol 28 : 555–573.
44. CoufalNG, Garcia-PerezJL, PengGE, YeoGW, MuY, et al. (2009) L1 retrotransposition in human neural progenitor cells. Nature 460 : 1127–1131.
45. EvronyGD, CaiX, LeeE, HillsLB, ElhosaryPC, et al. (2012) Single-neuron sequencing analysis of l1 retrotransposition and somatic mutation in the human brain. Cell 151 : 483–496.
46. Bourc'hisD, BestorTH (2004) Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431 : 96–99.
47. SuzukiS, OnoR, NaritaT, PaskAJ, ShawG, et al. (2007) Retrotransposon silencing by DNA methylation can drive mammalian genomic imprinting. PLoS Genet 3: e55 doi:10.1371/journal.pgen.0030055.
48. BarlowDP (1993) Methylation and imprinting: from host defense to gene regulation? Science 260 : 309–310.
49. HaigD (2012) Retroviruses and the placenta. Curr Biol 22: R609–613.
50. RougetC, PapinC, BoureuxA, MeunierAC, FrancoB, et al. (2010) Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo. Nature 467 : 1128–1132.
51. WatanabeT, TomizawaS, MitsuyaK, TotokiY, YamamotoY, et al. (2011) Role for piRNAs and noncoding RNA in de novo DNA methylation of the imprinted mouse Rasgrf1 locus. Science 332 : 848–852.
52. DrakeNM, ParkYJ, ShiraliAS, ClelandTA, SolowayPD (2009) Imprint switch mutations at Rasgrf1 support conflict hypothesis of imprinting and define a growth control mechanism upstream of IGF1. Mamm Genome 20 : 654–663.
53. YanY, ZhangY, YangK, SunZ, FuY, et al. (2011) Small RNAs from MITE-derived stem-loop precursors regulate abscisic acid signaling and abiotic stress responses in rice. Plant J 65 : 820–828.
54. WongS, WolfeKH (2005) Birth of a metabolic gene cluster in yeast by adaptive gene relocation. Nat Genet 37 : 777–782.
55. LeeJM, SonnhammerEL (2003) Genomic gene clustering analysis of pathways in eukaryotes. Genome Res 13 : 875–882.
56. CohenBA, MitraRD, HughesJD, ChurchGM (2000) A computational analysis of whole-genome expression data reveals chromosomal domains of gene expression. Nat Genet 26 : 183–186.
57. KunarsoG, ChiaNY, JeyakaniJ, HwangC, LuX, et al. (2010) Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat Genet 42 : 631–634.
58. JordanIK, RogozinIB, GlazkoGV, KooninEV (2003) Origin of a substantial fraction of human regulatory sequences from transposable elements. Trends Genet 19 : 68–72.
59. PolavarapuN, Marino-RamirezL, LandsmanD, McDonaldJF, JordanIK (2008) Evolutionary rates and patterns for human transcription factor binding sites derived from repetitive DNA. BMC Genomics 9 : 226.
60. MacfarlanTS, GiffordWD, DriscollS, LettieriK, RoweHM, et al. (2012) Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487 : 57–63.
61. MacfarlanTS, GiffordWD, AgarwalS, DriscollS, LettieriK, et al. (2011) Endogenous retroviruses and neighboring genes are coordinately repressed by LSD1/KDM1A. Genes Dev 25 : 594–607.
62. RoweHM, JakobssonJ, MesnardD, RougemontJ, ReynardS, et al. (2010) KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463 : 237–240.
63. SchultzDC, AyyanathanK, NegorevD, MaulGG, RauscherFJ3rd (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16 : 919–932.
64. GellersenB, BrosensIA, BrosensJJ (2007) Decidualization of the human endometrium: mechanisms, functions, and clinical perspectives. Semin Reprod Med 25 : 445–453.
65. LynchVJ, LeclercRD, MayG, WagnerGP (2011) Transposon-mediated rewiring of gene regulatory networks contributed to the evolution of pregnancy in mammals. Nat Genet 43 : 1154–1159.
66. GerloS, DavisJR, MagerDL, KooijmanR (2006) Prolactin in man: a tale of two promoters. Bioessays 28 : 1051–1055.
67. EmeraD, CasolaC, LynchVJ, WildmanDE, AgnewD, et al. (2012) Convergent evolution of endometrial prolactin expression in primates, mice, and elephants through the independent recruitment of transposable elements. Mol Biol Evol 29 : 239–247.
68. DupressoirA, VernochetC, HarperF, GueganJ, DessenP, et al. (2011) A pair of co-opted retroviral envelope syncytin genes is required for formation of the two-layered murine placental syncytiotrophoblast. Proc Natl Acad Sci U S A 108: E1164–1173.
69. SamsteinRM, JosefowiczSZ, ArveyA, TreutingPM, RudenskyAY (2012) Extrathymic generation of regulatory T cells in placental mammals mitigates maternal-fetal conflict. Cell 150 : 29–38.
70. NaitoK, ZhangF, TsukiyamaT, SaitoH, HancockCN, et al. (2009) Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461 : 1130–1134.
71. LeeE, IskowR, YangL, GokcumenO, HaseleyP, et al. (2012) Landscape of somatic retrotransposition in human cancers. Science 337 : 967–971.
72. Fiston-LavierAS, CarriganM, PetrovDA, GonzalezJ (2011) T-lex: a program for fast and accurate assessment of transposable element presence using next-generation sequencing data. Nucleic Acids Res 39: e36.
73. StewartC, KuralD, StrombergMP, WalkerJA, KonkelMK, et al. (2011) A comprehensive map of mobile element insertion polymorphisms in humans. PLoS Genet 7: e1002236 doi:10.1371/journal.pgen.1002236.
74. PoppC, DeanW, FengS, CokusSJ, AndrewsS, et al. (2010) Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463 : 1101–1105.
75. LaneN, DeanW, ErhardtS, HajkovaP, SuraniA, et al. (2003) Resistance of IAPs to methylation reprogramming may provide a mechanism for epigenetic inheritance in the mouse. Genesis 35 : 88–93.
76. RakyanVK, ChongS, ChampME, CuthbertPC, MorganHD, et al. (2003) Transgenerational inheritance of epigenetic states at the murine Axin(Fu) allele occurs after maternal and paternal transmission. Proc Natl Acad Sci U S A 100 : 2538–2543.
77. StoyeJP (2012) Studies of endogenous retroviruses reveal a continuing evolutionary saga. Nature reviews Microbiology 10 : 395–406.
78. CopelandNG, HutchisonKW, JenkinsNA (1983) Excision of the DBA ecotropic provirus in dilute coat-color revertants of mice occurs by homologous recombination involving the viral LTRs. Cell 33 : 379–387.
79. CordauxR, HedgesDJ, HerkeSW, BatzerMA (2006) Estimating the retrotransposition rate of human Alu elements. Gene 373 : 134–137.
80. XingJ, ZhangY, HanK, SalemAH, SenSK, et al. (2009) Mobile elements create structural variation: analysis of a complete human genome. Genome Res 19 : 1516–1526.
81. MoyesD, GriffithsDJ, VenablesPJ (2007) Insertional polymorphisms: a new lease of life for endogenous retroviruses in human disease. Trends Genet 23 : 326–333.
82. BurnsKH, BoekeJD (2012) Human transposon tectonics. Cell 149 : 740–752.
83. FinneganDJ (2012) Retrotransposons. Curr Biol 22: R432–437.
Štítky
Genetika Reprodukční medicína
Článek Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across PathogensČlánek TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association StudiesČlánek Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization inČlánek Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA ExpressionČlánek The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of GenesČlánek The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 1- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
-
Všechny články tohoto čísla
- A Model of High Sugar Diet-Induced Cardiomyopathy
- Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across Pathogens
- Emerging Function of Fat Mass and Obesity-Associated Protein (Fto)
- Positional Cloning Reveals Strain-Dependent Expression of to Alter Susceptibility to Bleomycin-Induced Pulmonary Fibrosis in Mice
- Genetics of Ribosomal Proteins: “Curiouser and Curiouser”
- Transposable Elements Re-Wire and Fine-Tune the Transcriptome
- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- MAML1 Enhances the Transcriptional Activity of Runx2 and Plays a Role in Bone Development
- Predicting Mendelian Disease-Causing Non-Synonymous Single Nucleotide Variants in Exome Sequencing Studies
- A Systematic Mapping Approach of 16q12.2/ and BMI in More Than 20,000 African Americans Narrows in on the Underlying Functional Variation: Results from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- Transcription of the Major microRNA–Like Small RNAs Relies on RNA Polymerase III
- Histone H3K56 Acetylation, Rad52, and Non-DNA Repair Factors Control Double-Strand Break Repair Choice with the Sister Chromatid
- Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 12q23.1 for Lung Squamous Cell Carcinoma in Han Chinese
- Genetic Disruption of the Copulatory Plug in Mice Leads to Severely Reduced Fertility
- The [] Prion Exists as a Dynamic Cloud of Variants
- Adult Onset Global Loss of the Gene Alters Body Composition and Metabolism in the Mouse
- Fis Protein Insulates the Gene from Uncontrolled Transcription
- The Meiotic Nuclear Lamina Regulates Chromosome Dynamics and Promotes Efficient Homologous Recombination in the Mouse
- Genome-Wide Haplotype Analysis of Expression Quantitative Trait Loci in Monocytes
- TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association Studies
- Structural Basis of a Histone H3 Lysine 4 Demethylase Required for Stem Elongation in Rice
- The Ecm11-Gmc2 Complex Promotes Synaptonemal Complex Formation through Assembly of Transverse Filaments in Budding Yeast
- MCM8 Is Required for a Pathway of Meiotic Double-Strand Break Repair Independent of DMC1 in
- Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications
- Integration of Nodal and BMP Signals in the Heart Requires FoxH1 to Create Left–Right Differences in Cell Migration Rates That Direct Cardiac Asymmetry
- Pharmacodynamics, Population Dynamics, and the Evolution of Persistence in
- A Hybrid Likelihood Model for Sequence-Based Disease Association Studies
- Aberration in DNA Methylation in B-Cell Lymphomas Has a Complex Origin and Increases with Disease Severity
- Multiple Opposing Constraints Govern Chromosome Interactions during Meiosis
- Transcriptional Dynamics Elicited by a Short Pulse of Notch Activation Involves Feed-Forward Regulation by Genes
- Dynamic Large-Scale Chromosomal Rearrangements Fuel Rapid Adaptation in Yeast Populations
- Heterologous Gln/Asn-Rich Proteins Impede the Propagation of Yeast Prions by Altering Chaperone Availability
- Gene Copy-Number Polymorphism Caused by Retrotransposition in Humans
- An Incompatibility between a Mitochondrial tRNA and Its Nuclear-Encoded tRNA Synthetase Compromises Development and Fitness in
- Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization in
- Single-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
- Tbx2 Controls Lung Growth by Direct Repression of the Cell Cycle Inhibitor Genes and
- Suv4-20h Histone Methyltransferases Promote Neuroectodermal Differentiation by Silencing the Pluripotency-Associated Oct-25 Gene
- A Conserved Helicase Processivity Factor Is Needed for Conjugation and Replication of an Integrative and Conjugative Element
- Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
- Genome-Wide Analysis Reveals Selection for Important Traits in Domestic Horse Breeds
- Coordinated Degradation of Replisome Components Ensures Genome Stability upon Replication Stress in the Absence of the Replication Fork Protection Complex
- Nkx6.1 Controls a Gene Regulatory Network Required for Establishing and Maintaining Pancreatic Beta Cell Identity
- HIF- and Non-HIF-Regulated Hypoxic Responses Require the Estrogen-Related Receptor in
- Delineating a Conserved Genetic Cassette Promoting Outgrowth of Body Appendages
- The Telomere Capping Complex CST Has an Unusual Stoichiometry, Makes Multipartite Interaction with G-Tails, and Unfolds Higher-Order G-Tail Structures
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Loci Associated with -Glycosylation of Human Immunoglobulin G Show Pleiotropy with Autoimmune Diseases and Haematological Cancers
- Switchgrass Genomic Diversity, Ploidy, and Evolution: Novel Insights from a Network-Based SNP Discovery Protocol
- Centromere-Like Regions in the Budding Yeast Genome
- Sequencing of Loci from the Elephant Shark Reveals a Family of Genes in Vertebrate Genomes, Forged by Ancient Duplications and Divergences
- Mendelian and Non-Mendelian Regulation of Gene Expression in Maize
- Mutational Spectrum Drives the Rise of Mutator Bacteria
- Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA Expression
- The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of Genes
- Sex-Specific Signaling in the Blood–Brain Barrier Is Required for Male Courtship in
- A Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
- Is Required for Leptin-Mediated Depolarization of POMC Neurons in the Hypothalamic Arcuate Nucleus in Mice
- Unlocking the Bottleneck in Forward Genetics Using Whole-Genome Sequencing and Identity by Descent to Isolate Causative Mutations
- The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
- MTERF3 Regulates Mitochondrial Ribosome Biogenesis in Invertebrates and Mammals
- Downregulation and Altered Splicing by in a Mouse Model of Facioscapulohumeral Muscular Dystrophy (FSHD)
- NBR1-Mediated Selective Autophagy Targets Insoluble Ubiquitinated Protein Aggregates in Plant Stress Responses
- Retroactive Maintains Cuticle Integrity by Promoting the Trafficking of Knickkopf into the Procuticle of
- Phenome-Wide Association Study (PheWAS) for Detection of Pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network
- Genetic and Functional Modularity of Activities in the Specification of Limb-Innervating Motor Neurons
- A Population Genetic Model for the Maintenance of R2 Retrotransposons in rRNA Gene Loci
- A Quartet of PIF bHLH Factors Provides a Transcriptionally Centered Signaling Hub That Regulates Seedling Morphogenesis through Differential Expression-Patterning of Shared Target Genes in
- A Genome-Wide Integrative Genomic Study Localizes Genetic Factors Influencing Antibodies against Epstein-Barr Virus Nuclear Antigen 1 (EBNA-1)
- Mutation of the Diamond-Blackfan Anemia Gene in Mouse Results in Morphological and Neuroanatomical Phenotypes
- Life, the Universe, and Everything: An Interview with David Haussler
- Alternative Oxidase Expression in the Mouse Enables Bypassing Cytochrome Oxidase Blockade and Limits Mitochondrial ROS Overproduction
- An Evolutionarily Conserved Synthetic Lethal Interaction Network Identifies FEN1 as a Broad-Spectrum Target for Anticancer Therapeutic Development
- The Flowering Repressor Underlies a Novel QTL Interacting with the Genetic Background
- Telomerase Is Required for Zebrafish Lifespan
- and Diversified Expression of the Gene Family Bolster the Floral Stem Cell Network
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Increased Maternal Genome Dosage Bypasses the Requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis Seed Development
- WNK1/HSN2 Mutation in Human Peripheral Neuropathy Deregulates Expression and Posterior Lateral Line Development in Zebrafish ()
- Synergistic Interaction of Rnf8 and p53 in the Protection against Genomic Instability and Tumorigenesis
- Dot1-Dependent Histone H3K79 Methylation Promotes Activation of the Mek1 Meiotic Checkpoint Effector Kinase by Regulating the Hop1 Adaptor
- A Heterogeneous Mixture of F-Series Prostaglandins Promotes Sperm Guidance in the Reproductive Tract
- Starvation, Together with the SOS Response, Mediates High Biofilm-Specific Tolerance to the Fluoroquinolone Ofloxacin
- Directed Evolution of a Model Primordial Enzyme Provides Insights into the Development of the Genetic Code
- Genome-Wide Screens for Tinman Binding Sites Identify Cardiac Enhancers with Diverse Functional Architectures
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



