-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaAnalysis of nuclear and organellar genomes of in humans reveals ancient population structure and recent recombination among host-specific subpopulations
Plasmodium knowlesi, a common malaria parasite of long-tailed and pig-tailed macaques, is now recognized as a significant cause of human malaria, accounting for up to 70% of malaria cases in certain areas in Southeast Asia including Malaysian Borneo. Rapid human population growth, deforestation and encroachment on wild macaque habitats potentially increase contact with humans and drive up the prevalence of human Plasmodium knowlesi infections. Appropriate molecular tools and sampling are needed to assist surveillance by malaria control programmes, and to understand the genetics underpinning Plasmodium knowlesi transmission and switching of hosts from macaques to humans. We report a comprehensive analysis of the largest assembled set of Plasmodium knowlesi genome sequences from Malaysia. It reveals genetic regions that have been recently exchanged between long-tailed and pig-tailed macaques, which contain genes with signals indicative of rapid contemporary ecological change, including deforestation. Additional analyses partition Plasmodium knowlesi infections in Borneo into 3 deeply branched lineages of ancient origin, which founded the two divergent populations associated with long-tailed and pig-tailed macaques and a third, highly diverse population, on the Peninsular mainland. Overall, the complex Plasmodium parasite evolution observed and likelihood of further host transitions are potential challenges to malaria control in Malaysia.
Published in the journal: . PLoS Genet 13(9): e32767. doi:10.1371/journal.pgen.1007008
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1007008Summary
Plasmodium knowlesi, a common malaria parasite of long-tailed and pig-tailed macaques, is now recognized as a significant cause of human malaria, accounting for up to 70% of malaria cases in certain areas in Southeast Asia including Malaysian Borneo. Rapid human population growth, deforestation and encroachment on wild macaque habitats potentially increase contact with humans and drive up the prevalence of human Plasmodium knowlesi infections. Appropriate molecular tools and sampling are needed to assist surveillance by malaria control programmes, and to understand the genetics underpinning Plasmodium knowlesi transmission and switching of hosts from macaques to humans. We report a comprehensive analysis of the largest assembled set of Plasmodium knowlesi genome sequences from Malaysia. It reveals genetic regions that have been recently exchanged between long-tailed and pig-tailed macaques, which contain genes with signals indicative of rapid contemporary ecological change, including deforestation. Additional analyses partition Plasmodium knowlesi infections in Borneo into 3 deeply branched lineages of ancient origin, which founded the two divergent populations associated with long-tailed and pig-tailed macaques and a third, highly diverse population, on the Peninsular mainland. Overall, the complex Plasmodium parasite evolution observed and likelihood of further host transitions are potential challenges to malaria control in Malaysia.
Introduction
Plasmodium knowlesi, a common malaria parasite of long-tailed Macaca fascicularis (Mf) and pig-tailed M. nemestrina (Mn) macaques in Southeast Asia, is now recognized as a significant cause of human malaria. A cluster of human P. knowlesi cases were reported from Malaysian Borneo in 2004 [1], but now human infections are known to be widespread in Southeast Asia [2,3], and have been reported in travellers from outside the region [2,4]. Clinical symptoms range from asymptomatic carriage to high parasitaemia with severe complications including death [5,6]. As rapid human population growth, deforestation and encroachment on remaining wild macaque habitats potentially increases contact with humans [7], in Southeast Asian countries P. knowlesi is now coming to the attention of national malaria control and elimination programmes that have hitherto focused on P. vivax and P. falciparum [2].
P. knowlesi commonly displays multi-clonality in humans and macaques, and analysis of microsatellite markers, csp, 18S rRNA, and mtDNA sequences indicates no systematic differences between human and macaque isolates from Malaysian Borneo [8]. Whole genome-level genetic diversity among P. knowlesi from human infections in Sarikei in Sarawak demonstrates substantial dimorphism extending over at least 50% of the genome [9]. This finding is supported by analysis of microsatellite diversity in parasites from Mf, Mn and human infections across Peninsular and Borneo Malaysia [10]. It also provides evidence that the two distinct genome dimorphs reflect adaptation to either of the two host macaque species, although no evidence of a complete barrier in primate host susceptibility was found [10]. A third genome cluster has been described from geographically distinct Peninsular Malaysia [11, 12, 13, 14].
Studies of mtDNA have revealed that ancestral P. knowlesi predates the settlement of Homo sapiens in Southeast Asia, the evolutionary emergence of P. falciparum and P. vivax, and underwent population expansion 30–40 thousand years ago [8]. Diversity at the genomic level is thus likely to reflect host - and geography-related partitioning during this expansion, as well as additional recent complexity due to contemporary changes in host and vector distributions during ongoing ecological transition in the region [15]. Several Anopheles species, all from the Leuchosphyrus group, are capable of transmitting P. knowlesi malaria, including A. latens and A. balbacensis in Malaysian Borneo [16, 17, 18], A. hackeri and A. cracens in Peninsular Malaysia [19] and A. dirus in southern Vietnam [20]. It is thus likely that patterns of genome diversity in natural populations of P. knowlesi reflect partitioning among both Dipteran and primate hosts occurring on varying time-scales through the evolutionary history of the species. Such partitioning can plausibly prevent or reduce panmictic genetic exchange.
Genomic studies of P. knowlesi to date have considered nuclear gene diversity and dimorphism among naturally-infected human hosts, and macaque-derived laboratory-maintained isolates from the 1960s [10, 12]. However, these studies did not consider non-nuclear organellar genomes in the mitochondrion and apicoplast of malaria parasites, which are non-recombinant and uniparentally inherited, and can provide evidence of genome evolution on a longer timescale [21]. Recombination barriers among insect and primate hosts may have less impact on sequence diversity in the organellar genomes of P. knowlesi. Utilising a new P. knowlesi reference genome generated using long-read technology [22] we performed a new analysis of all available nuclear and non-nuclear genome sequences. Patterns of polymorphisms were analysed to identify evolutionary signals of both recent and ancient events associated with the partitioning of the di - or tri-morphic genomes previously reported.
Results
Sequence data reveals multiplicity of infection
Raw short-read sequence data from all available P. knowlesi isolates (S1 Fig) were mapped to a new reference genome [22] from the human-adapted P. knowlesi line A1-H.1 genome [23], yielding an average coverage of ~120-fold across 99% of the reference genome (S1 Table), and 1,632,024 high quality SNPs. The high density of point mutations (1 every 15bp) in P. knowlesi compared to P. vivax and P. falciparum has been previously noted [10]. Seven macaque-derived isolates were found to have high multiplicity of infection (S2 Fig), and were excluded, leaving an analysis set of 60 isolates.
Population structure analysis reveals new natural genetic exchange
SNP-based neighbour-joining tree analysis revealed three subpopulation groups that coincide with isolates presenting the Mf-associated P. knowlesi genotype (Mf-Pk, Borneo Malaysia, Cluster 1), the Mn-associated P. knowlesi genotype (Mn-Pk, Borneo Malaysia, Cluster 2) [10, 11, 12, 14], and older Peninsular Malaysia strains (Cluster 3) (Fig 1A). Within Cluster 1 we observed two geographic sub-groups that coincide with Kapit and Betong regions in Malaysian Borneo. The samples from Sarikei region (DIM prefix), geographically located equidistant between Kapit and Betong, fall into either cluster (S3 Fig). Overall, the regional clusters from Kapit and Betong were more genetically similar to each other (mean fixation index FST 0.03, S4 Fig) than were the host-associated clusters (Cluster 1 vs. 2, mean FST 0.21). However, a significant chromosomal anomaly was identified that differentiated the Kapit and Betong Mf-Pk subgroups; this occurred in a multi-gene region on chromosome 8 (~500 SNPs with FST values >0.4; Fig 1B; S4 Fig).
Fig. 1. Whole genome population structure and evidence of genetic exchange in chromosome 8. 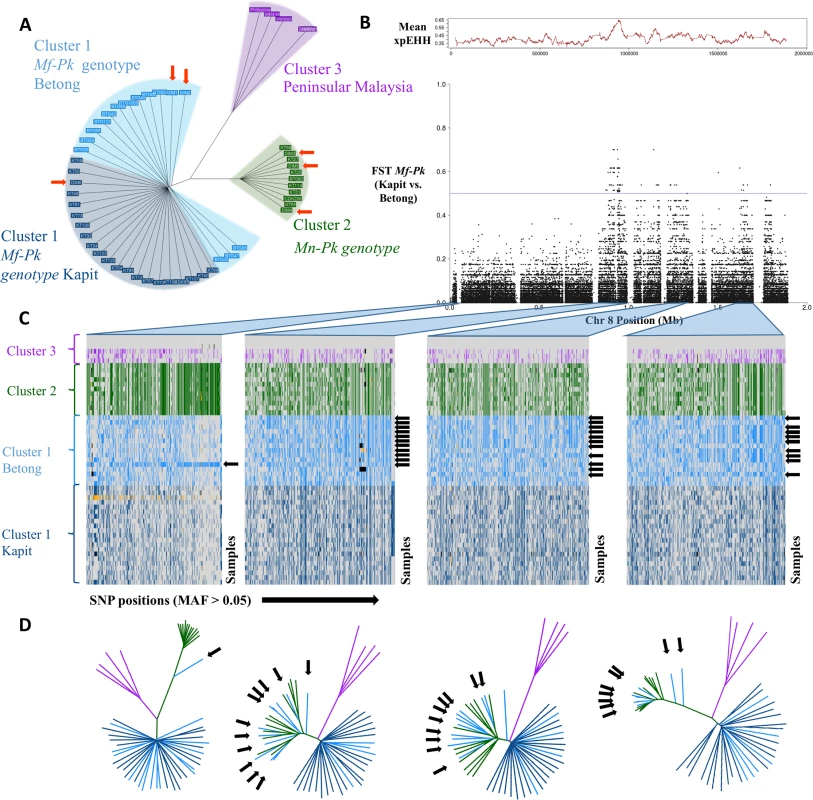
A) Neighbour joining tree constructed using 1,632,024 genome-wide SNPs across the 60 P. knowlesi (Pk) samples. The tree shows two levels of resolution involved in the clustering of genotypes. The first level differentiates Peninsular Malaysia samples (Cluster 3, purple) from the Malaysian-Borneo host-related Pk genotypes (Cluster 1, M. fascicularis macaques (Mf-Pk), blue; Cluster 2, M. nemestrina macaques (Mn-Pk), green). The second level differentiates within Cluster 1, where Mf-Pk genotypes fall in subgroups from Betong (light blue) and from Kapit (dark blue). Samples from Sarikei have been highlighted using orange arrows. B) Allele frequency differences between Betong and Kapit regional subgroups of the Mf-Pk genotype in chromosome 8 SNPs using the population differentiation measure FST. There is high differentiation (FST > 0.4) in several regions across chromosome 8 (0.85-1Mb, 1.2Mb-1.35Mb, and 1.6–1.7Mb), and these signals overlap with strong evidence of recent positive selection, measured by the average XP-EHH calculated in 1kbp windows (red trace above). C) Haplotype plots for all samples (y-axis) at common SNP positions (MAF >5%, x-axis) highlighting the regions with abnormally high FST values (0.85-1Mb, 1.2Mb-1.35Mb, and 1.6–1.7Mb), as well as the low Fst region spanning from 0.1 to 0.2Mb for comparison. The black arrows indicate samples with the Mf-Pk genotype from Betong present with a Mn-Pk Cluster 2-like haplotype. These patterns are indicative of genetic exchange between the Mf-Pk and Mn-Pk genotype clusters, which is supported by the neighbour joining trees included in D). Missing calls are coloured in black and mixed calls are coloured in yellow. D) Neighbour joining trees constructed using SNPs in each of the regions in C). The trees show clear clustering of Mf-Pk Betong samples with the Mn-Pk genotype cluster in the genetic regions of abnormal FST (2nd, 3rd and 4th trees) compared to the 1st tree where only sample DIM2 presents introgression. Signatures of introgression events in chromosome 8
To explore the anomaly in chromosome 8, individual haplotypes and neighbour-joining trees were constructed across several loci (Fig 1C and Fig 1D) revealing two very distinct patterns. The first pattern was observed in the chromosomal sections with low genetic diversity between the two Mf-Pk regional clusters (FST < 0.2, Fig 1B). The tree structure for these genomic regions (Fig 1D, 1st tree) mimics that of the genome-wide tree in Fig 1A. Strong haplotype differentiation between the host-associated Clusters 1 (Mf-Pk) and 2 (Mn-Pk) was confirmed in the SNP-based profiles (Fig 1C, 1st column).
A second pattern was observed in regions of chromosome 8 with distinct genetic differentiation between Kapit and Betong subgroups (FST > 0.4). Many Mf-Pk Betong subgroup isolates presented segments almost identical to chromosome 8 sequences of the Mn-Pk genotype from Cluster 2 (Fig 1D, 2nd, 3rd and 4th trees). This exchange is supported by the SNP-based haplotype patterns, where a distinct haplotype in the Betong samples is Cluster 2-like (Fig 1C, 2nd, 3rd and 4th columns, black arrows), suggesting the introgression of large chromosomal regions (up to 200Kb) between Mf-Pk (Cluster 1) and Mn-Pk (Cluster 2). This is consistent with a very recent event of natural genetic exchange between these subgroups of P. knowlesi recently isolated from human infections. The high frequency of the new haplotype (73%) in the Betong subgroup suggests that it is under (recent) strong selection pressure in this region. The presence of differences in extended haplotype homozygosity between the recombinant and non-recombinant regional Mf-Pk subpopulations provides additional evidence of recent positive selection (XP-EHH peak, P<0.0001) in a region of increased population differentiation (FST > 0.4, Fig 1B).
The functional nature of genes in chromosome 8 involved in these putative introgression events was investigated (FST > 0.4, Table 1), and found to include loci that are important in the vector component of the Plasmodium life cycle. For example, cap380 (PKNH_0820800, 101 SNPs with FST > 0.4) encodes a protein expressed in the external capsule of the oocyst. This gene is essential in the maturation from ookinete into oocyst in P. berghei, and is assumed to assist in evasion of mosquito immune mechanisms [24]. Another gene, PKNH_0826900 (19 SNPs) encodes for the circumsporozoite - and TRAP-related protein (CTRP), which has an established role in ookinete motility in P. berghei and is essential for binding to and invading the mosquito midgut [25]. Further, homologues of PKNH_0826400 (21 SNPs) display increased transcription levels in ookinete and gametocyte V sexual stages in both P. falciparum [26] and P. berghei [27] compared to the asexual ring stage (fold change of at least 2). The transcriptomic profiles of these strongly selected genes are shown in S5 Fig.
Tab. 1. Genes located within the chromosome 8 regions of genetic exchange and transcriptional changes. 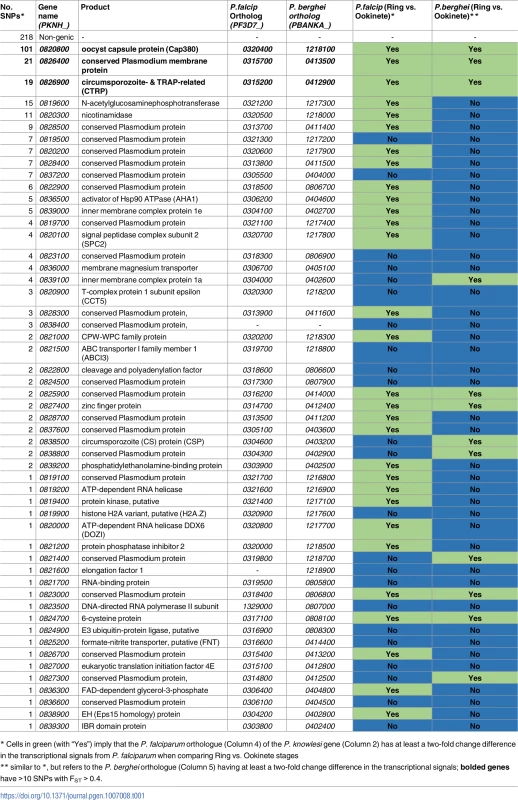
* Cells in green (with “Yes”) imply that the P. falciparum orthologue (Column 4) of the P. knowlesi gene (Column 2) has at least a two-fold change difference in the transcriptional signals from P. falciparum when comparing Ring vs. Ookinete stages Genome-wide evidence of genetic exchange events in P. knowlesi
By applying a combination of neighbour joining trees and SNP diversity analysis across 50 Kbp windows, we identified that 33/60 isolates show clear evidence of genetic exchange between Clusters 1 and 2 (S2 Table). Regions involved in exchange (recombination) (137/494 regions, 86% contained an ookinete related gene) showed evidence of enrichment for ookinete-expressed genes compared to other (non-recombinant) chromosome regions (357/494 regions, 77% contained an ookinete related gene) (Chi Square P = 0.03). One such region in chromosome 12 included the Pf47-like (PKH_120710) gene, where the orthologue in P. falciparum is a known mediator of the evasion of the mosquito immune system [28]. Furthermore, it has been shown that a change in haplotype in this gene in a P. falciparum isolate is sufficient to make it compatible to a different mosquito species [28]. Nearly half (45%) of isolates from Betong presented with a recombinant profile in PKH_120710.
In general, the genetic exchanges generated differing levels of mosaicism in each population and among individual isolates across all chromosomes (S6 Fig). One isolate from Sarikei with the Mf genome dimorph type (DIM2) appeared to harbour Mn-type introgressed sequences in 8% of the genome, occurring across 6 chromosomes (6, 7, 8, 9, 11 and 12), including an almost complete Mn-type chromosome 8. Of the 33 samples with evidence of exchanges, 13 were from the Betong region, 14 from Kapit and 6 from Sarikei, which indicates that the events are not geographically restricted. Although, the majority of genetic exchange events involve the integration of Mn-type motifs into Mf-type genomes, introgression in the opposite direction was also observed, but on a smaller scale and at lower frequency.
Organellar genomes also reflect genetic exchange events
The mitochondrial and apicoplast genomes of each P. knowlesi isolate was interrogated for signals of evolutionary history over longer time-scales, as in previous studies [21, 29, 30]. Combining the mitochondrial sequence data from the 60 P. knowlesi isolates from this study together with 54 previously published mitochondrial sequences including human and both Mn and Mf samples [9], we generated a phylogenetic tree (Fig 2). This tree shows four clades (shown in purple, red, blue and green). To interpret these clades, they were cross-referenced to the previously defined 3 nuclear genotypes (Clusters 1 to 3) and the host contributing the sample (human, macaque-type). The red and purple clades possess similar mitochondrial haplotypes as highlighted by their inter-cluster average FST (red vs. purple: average FST = 0.16), which is lower than comparisons including the other two clusters (red or purple vs. blue or green: average FST > 0.18). The purple clade consists of cultured isolates from Peninsular Malaysia, and is associated with the Peninsular nuclear genotype (Cluster 3). The red and green clades each contain a mixture of Borneo Malaysia samples from both humans and macaques with nuclear genotypes from Clusters 1 and 2. The green clade also includes the only sequence sourced from a M. nemestrina host. The blue clade contains samples from humans and macaques, all with Cluster 1 nuclear genotypes. The divergence of these mitochondrial clades from their common ancestor was estimated to be 72k years ago, and younger than the previous the estimate of 257k but within error [8]. Furthermore, the presence of monkey-derived sequences spread across the tree seems to indicate that none of the mitochondrial genotypic groups found is human-specific as all have also been observed in macaques, also consistent with previous findings [9].
Fig. 2. Phylogenetic tree constructed from P. knowlesi mitochondrial sequences for the 60 whole genome sequenced samples and 54 published others [6] sourced from human, M. nemestrina (Mn) and M. fascicularis (Mf) samples. ![Phylogenetic tree constructed from <i>P</i>. <i>knowlesi</i> mitochondrial sequences for the 60 whole genome sequenced samples and 54 published others [<em class="ref">6</em>] sourced from human, <i>M</i>. <i>nemestrina (Mn)</i> and <i>M</i>. <i>fascicularis (Mf)</i> samples.](https://pl-master.mdcdn.cz/media/cache/media_object_image_small/media/image/8f2a8d7177f7347a0b682db31fe66280.png)
The mitochondrial genotype groups defined here are cross-referenced to the nuclear genotypes in Fig 1A (pentagons in the outer ring, missing pentagons relate to the 54 samples with only mitochondrial sequence data [6]). Samples sourced from the different macaques are highlighted in the tree branches. The tree shows three main subpopulations: (i) two clades including Peninsular Malaysia (Peninsular nuclear genotype, Cluster 3, purple) and Borneo Malaysia (mix of Mf-Pk and Mn-Pk nuclear genotypes, Cluster 1 and 2, red) presenting a very similar mitochondrial haplotype; (ii) the majority of the samples with a Mn-Pk nuclear genotype together with the only sequence obtained from a Mn sample (Cluster 2, green); (iii) samples with a Mf-Pk nuclear genotype (Cluster 1, blue). These clusters are consistent with microsatellite-based trees [12]. The presence of monkey samples spread throughout the tree indicates that none of the mitochondrial genotypes groups are human-specific, consistent with microsatellite-based analysis [9]. Black arrows indicate the presence of samples with mismatched nuclear and mitochondrial subtypes. Using the common SNPs (280/425 with MAF > 5%: apicoplast 252, mitochondria 28 SNPs) in the 60 isolates with the sequence data we confirmed that the organellar genomes are co-inherited (mean pairwise organellar linkage disequilibrium D’ = 0.99). SNP-based haplotype profile analysis (S7A Fig) revealed clustering that is consistent with the three main clusters seen in Fig 2. Similarly, a phylogenetic tree constructed using only apicoplast SNPs (S7B Fig) is congruent with the mitochondrial based tree (Fig 2). The presence of mismatched nuclear and organellar type genomes in two of the three clusters (black arrows in Fig 2) and the presence of such mismatched samples with little or no evidence of nuclear genome recombination suggests ancient genetic exchange events between distinct lineages. The nuclear footprints of such exchanges are likely to have been broken down by recombination over time. We observed a significant incongruence between the robust phylogenetic tree topologies based on organellar and nuclear genome SNPs (Shimodaira-Hasegawa test P = 0.001; Templeton test P = 0.003) (Fig 2). These results from organellar and nuclear genomes, in a small but geographically diverse set of P. knowlesi, indicate that there have been several genetic exchanges between the host-associated clusters in Malaysian Borneo.
Discussion
P. knowlesi is now the major cause of malaria in Malaysian Borneo, but the biology of the parasite [15, 22, 23], host and vector interactions, and disease distribution and epidemiology [19, 31, 32] are not well understood. The availability of a new high-quality reference sequence and a more robust approach to MOI were used to re-evaluate the previously described peninsular and macaque-associated subpopulations of P. knowlesi parasites. We report two major new findings. First, clear evidence of natural genetic exchanges between the divergent Mf- and Mn-associated subpopulations of P. knowlesi, including a major segment of introgression on chromosome 8, is presented. Second, the presence of haplotype sub-divisions in the organellar genomes that do not map onto the subpopulations implied by nuclear genome analysis indicate that exchange events have previously occurred in non-recent history. A similar multi-tiered pattern of evolution among nuclear and organellar genomes has been found in Trypanosoma cruzi, an unrelated protozoan parasite with a mammalian host-insect vector life cycle [29, 30].
Unexpectedly, observed mosaicism and population differentiation signals were not encountered equally across the P. knowlesi nuclear genome, but were particularly prominent on chromosome 8, with genes expressed in mosquito stages over-represented. For example, the majority (73%) of Mf-associated isolates from Betong harboured the Mn-associated allele of the oocyst-expressed cap380 gene, which differs at 101 positions from the allele found in the Mf-associated cluster. This is essential for ookinete to oocyst maturation and therefore for the transmission of the parasite during the vector stage [24, 25]; here, we identify signals of recent selective pressure on this locus (Fig 1B). Other vector-related genes were identified within the introgressed segment, and point towards strong evolutionary selection pressure on the parasites driven by the transmitting Anopheles vector species. Such effects have been found in P. falciparum [28] and P. vivax genomes [33], and highlight the importance of understanding the distribution of the different Anopheles vector species, their host feeding preferences, and their interactions with the parasite in highly dynamic and complex environments such as the ecological niche of P. knowlesi.
Nearly 80% of Malaysian Borneo has undergone deforestation or agricultural expansion, which have driven habitat modification affecting both macaque and Anopheles host species, and the proximity of humans to both [8, 31]. Furthermore, studies have predicted that Mn predominantly inhabits forested areas while Mf reside in more cosmopolitan areas, which include croplands, vegetation mosaics, rubber plantations and forested areas [8, 34]. The main genomic exchange event on chromosome 8 involves essential vector-related genes and is pin-pointed geographically to the Betong area. This region has undergone significant forest degradation due to expansion of industrial plantations in the recent years [35]. These types of environmental changes have been previously related to alterations in the vector species distribution in Malaysia, leading to malaria epidemics [36]. Environmental changes also affect macaque habitats, and increase the opportunities for human-macaque interaction [31], but selection events highlighted in this study seem to primarily reflect adaptation of the parasite to changes in mosquito distribution or to recent changes in the vectorial capacity of the existing vectors. The depth, breadth and spread of the genetic exchanges observed in three different areas (Betong, Kapit and Sarikei) in Sarawak highlight the potential importance of these events for parasite adaptation in both vertebrate and invertebrate species.
Although, the level of genetic diversity between Mf- and Mn-associated P. knowlesi has some similarity to that observed between P. ovale curtisi and P. o. wallikeri, now considered separate species [37], the evidence of recombination and genetic exchanges observed in this study precludes species designation, as reproductive isolation is not complete. Nevertheless, better understanding of P. knowlesi population structure could aid future studies across the regions where human populations have been identified at risk of infection including both symptomatic and asymptomatic cases [4, 38, 39]. This would assist with characterising and tracking subpopulations and genetic exchanges, and provide a flexible framework for better understanding P. knowlesi diversity across the region.
Our work has provided insight into Plasmodium parasite evolution. It has been suggested that malaria parasites have survived using either adaptive radiation where host switching plays a key role [40], or alternatively adaptation to complex historical and geographical environments leading to speciation [41]. Plasmodium species in non-human natural conditions in the absence of drug selection pressure have a wide range of possible hosts [41, 42]. The P. knowlesi data has shown that geographical or ecological isolation of the different hosts over an extended time can generate subgroups of parasites with substantial genetic differentiation, but capable of recombining when in contact [12, 30, 31]. This pattern has a major impact on the parasite genome, as illustrated by the profound chromosome mosaicism observed among our study isolates. Our data suggest that the broad host specificity of some of the Plasmodium species are important drivers of parasite genomic diversity. In P. knowlesi this means that genetic divergence is enabled not only by long-term geographic isolation, as is the case between Peninsular and Bornean isolates, but also via the isolation afforded by extended transmission cycles within different primate hosts. The genetic trimorphism suggests that the separate macaque hosts provides sufficient genetic isolation to allow for host specific adaptations to occur, even within relatively small geographic areas. Furthermore, the possibility of recombination between partially differentiated parasite genomes increases opportunities for new adaptation, including further host transitions, and can only make malaria control more difficult. Genome-level studies on P. knowlesi isolates from Mf and Mn across the parasite’s geographic range are now needed to test the generalizability of this remarkable conclusion.
Materials and methods
P. knowlesi sequence data
Raw sequence data were downloaded for 48 isolates from Kapit and Betong in Malaysian Borneo [11], 6 isolates from Sairikei in Malaysian Borneo (S1 Fig) [9] and 6 long-time isolated lines, maintained in rhesus monkeys sourced originally from Peninsular Malaysia and Philippines [11]. The sequence data accession numbers can be found in S1 Table. The samples were aligned against the new reference for the human-adapted line A1-H.1 (pathogenseq.lshtm.ac.uk/knowlesi_1, accession number ERZ389239, [22]) using bwa-mem [43] and SNPs were called using the Samtools suite [44], and filtered for high quality SNPs using previously described methods [45, 46]. In particular, the SNP calling pipeline generated a total of 2,020,452 SNP positions, which were reduced to 1,632,024 high quality SNPs after removing those in non-unique regions, and in low quality and coverage positions. Samples were individually assessed for detecting multiplicity of infection (MOI) using: (i) estMOI [47] software, and (ii) quantifying the number of positions with mixed genotypes (if more than one allele at a specific position have been found in at least 20% of the reads [46]). The measures led to correlated results (r2 = 0.8), which highlighted the robustness of these two methods. Samples were classified into three subcategories: (i) single infections (> = 98% genome showing no evidence of MOI and < = 1/10,000 SNP positions with mixed genotypes), (ii) low MOI (>85% genome showing no evidence of MOI and < = 4/10,000 SNPs positions with mixed genotypes); (iii) high MOI (<85% genome showing no evidence of MOI, and > 4/10,000 SNPs positions with mixed genotypes). Samples with high MOI were removed from subsequent analyses.
Population genetics analysis
For comparisons between populations, we first applied the principal component analysis (PCA) and neighbourhood joining tree clustering based on a matrix of pairwise identity by state values calculated from the SNPs. We used the ranked FST statistics to identify the informative polymorphism driving the clustering observed in the PCA [48]. Finally, we created haplotype plots using only SNP positions with MAF > 0.05 over all the populations, and displayed each sample as a row to allow closer inspection of the chromosome regions where interesting recombination events are observed. The XP-EHH metric [49] implemented within the rehh R package was used to assess evidence of recent relative positive selection between regional clusters from Kapit and Betong. The results were smoothed by calculating means in 1 Kbp windows, where windows overlapped by 250bp. The raXML software (v.8.0.3, 1000 bootstrap samples) was used to construct robust phylogenetic trees (90% bootstrap values > 95) for nuclear and organellar SNPs. Estimates of divergence times for subpopulations was based on a Bayesian Markov Chain Monte Carlo (MCMC) (BEAST, v.1.8.1) approach applied to mitochondrial sequences, with identical parameters settings to those described elsewhere [8]. The Shimodaira-Hasegawa [50] and the Templeton [51] tests were used to detect incongruence between the tree topologies.
Identification of introgressed regions in the different chromosomes
In order to identify regions that have undergone introgression we calculated the pairwise SNP diversity (π) of each sample against all the Borneo samples using a 50 Kbp sliding window. This window size was sufficient to include the required number of SNPs for the robust identification of introgression events. The average π in the M. nemestrina associated (Mn-Pk) and M. fascicularis associated (Mf-Pk) clusters was calculated, leading to two diversity values for each sample (Mfπ and Mnπ) and thereby a measure of genetic distance to the average of the two clusters. For Mf samples, an increase in the Mfπ and a decrease in Mnπ would mean the sample is more similar to the Mn-Pk cluster than the average; vice versa for the Mf samples. In order to avoid the identification of spurious events, we applied a threshold of a 0.001 increase in the deviation from the original cluster.
Characterization of genes under strong selection after recombination
For P. knowlesi genes of interest, orthologues in P. falciparum and P. berghei genomes were identified using PlasmoDB (plasmodb.org). Gene expression data (including from the RNAseq platform) for these genes across different stages of the life cycle of the parasite were considered [26, 27]. In particular, we compared the average of the asexual blood stages and the sexual ookinete stage, highlighting the genes upregulated with a two-fold change (P<0.000001), for P. falciparum [26] and P. berghei [27].
Supporting Information
Zdroje
1. Singh B, Kim Sung L, Matusop A, Radhakrishnan A, Shamsul SS, Cox-Singh J, et al. A large focus of naturally acquired Plasmodium knowlesi infections in human beings. Lancet 2004; 363, 1017–1024. doi: 10.1016/S0140-6736(04)15836-4 15051281
2. Kantele A, Jokiranta TS. Review of cases with the emerging fifth human malaria parasite, Plasmodium knowlesi. Clin Infect Dis 2011; 52, 1356–1362. doi: 10.1093/cid/cir180 21596677
3. Putaporntip C, Hongsrimuang T, Seethamchai S, Kobasa T, Limkittikul K, Cui L, et al. Differential Prevalence of Plasmodium Infections and Cryptic Plasmodium Knowlesi Malaria in Humans in Thailand. J Infect Dis 2009; 199 : 1143–50. doi: 10.1086/597414 19284284
4. Muller M, Schlagenhauf P. Plasmodium knowlesi in travellers, update 2014. Int J infect Dis 2014; 22, 55–64.
5. Singh B, Daneshvar C. Human infections and detection of Plasmodium knowlesi. Clinical microbiology reviews 2013; 26, 165–184. doi: 10.1128/CMR.00079-12 23554413
6. Lubis IND, Wijaya H, Lubis M, Lubis CP, Divis PCS, Beshir KB, et. al. Contribution of Plasmodium knowlesi to Multispecies Human Malaria Infections in North Sumatera, Indonesia. J Infect Dis 2017; 215(7), 1148–1155. doi: 10.1093/infdis/jix091 28201638
7. Imai N, White MT, Ghani AC, Drakeley CJ. Transmission and Control of Plasmodium Knowlesi: A Mathematical Modelling Study. PLOS Negl Trop Dis 2014; 8 e2978. doi: 10.1371/journal.pntd.0002978 25058400
8. Lee KS, Divis PC, Zakaria SK, Matusop A, Julin RA, Conway DJ, et al. Plasmodium knowlesi: reservoir hosts and tracking the emergence in humans and macaques. PLoS Pathog 2011; 7, e1002015. doi: 10.1371/journal.ppat.1002015 21490952
9. Pinheiro MM, Ahmed MA, Millar SB, Sanderson T, Otto TD, Lu WC, et al. Plasmodium knowlesi Genome Sequences from Clinical Isolates Reveal Extensive Genomic Dimorphism. PLoS ONE 2015; 10(4), e0121303. doi: 10.1371/journal.pone.0121303 25830531
10. Divis PC, Singh B, Anderios F, Hisam S, Matusop A, Kocken CH, et al. Admixture in Humans of Two Divergent Plasmodium knowlesi Populations Associated with Different Macaque Host Species. PLoS Pathog 2015; 11(5), e1004888. doi: 10.1371/journal.ppat.1004888 26020959
11. Assefa S, Lim C, Preston MD, Duffy CW, Nair MB, Adroub SA, et al. Population genomic structure and adaptation in the zoonotic malaria parasite Plasmodium knowlesi. Proc National Academy Sci U.S.A 2015; 112(42), 13027–13032.
12. Ahmed MA, Fong MY, Lau YL., Yusof R. Clustering and genetic differentiation of the normocyte binding protein (nbpxa) of Plasmodium knowlesi clinical isolates from Peninsular Malaysia and Malaysia Borneo. Malaria J 2016; 15, 241.
13. Divis PC, Lin LC, Rovie-Ryan JJ, Kadir KA, Anderios F, Hisam S, et al. Three Divergent Subpopulations of the Malaria Parasite Plasmodium knowlesi. Emerging Infectious Diseases 2017; 23(4), 616–624. doi: 10.3201/eid2304.161738 28322705
14. Fornace KM, Abidin TR, Alexander N, Brock P, Grigg MJ, Murphy A, et al. Association between landscape factors and spatial patterns of Plasmodium knowlesi Infections in Sabah, Malaysia. Emerg Infect Dis. 2016; 22, 201–208. doi: 10.3201/eid2202.150656 26812373
15. Yusof R, Ahmed MA, Jelip J, Ngian HU, Mustakim S, Hussin HM, et al. Phylogeographic Evidence for 2 Genetically Distinct Zoonotic Plasmodium knowlesi Parasites, Malaysia. Emerging Infectious Diseases 2016; 22(8), 1371–1380. doi: 10.3201/eid2208.151885 27433965
16. Vythilingam I, Tan CH, Asmad M, Chan ST, Lee KS, Singh B. Natural transmission of Plasmodium knowlesi to humans by Anopheles latens in Sarawak, Malaysia. Trans Roy Soc Tropl Med Hyg 2006; 100(11), 1087–1088.
17. Tan CH, Vythilingam I, Matusop A, Chan ST, Singh B. Bionomics of Anopheles latens in Kapit, Sarawak, Malaysian Borneo in relation to the transmission of zoonotic simian malaria parasite Plasmodium knowlesi. Malaria J. 2008; 7(1), 52.
18. Brant HL, Ewers RM, Vythilingam I, Drakeley C, Benedick S, Mumford JD. D. Vertical stratification of adult mosquitoes (Diptera: Culicidae) within a tropical rainforest in Sabah, Malaysia. Malaria J. 2016; 15(1), 370.
19. Vythilingam I, Noorazian YM, Huat TC, Jiram AI, Yusri YM, Azahari AH, et al. Plasmodium knowlesi in humans, macaques and mosquitoes in peninsular Malaysia. Parasit Vectors 2008; 1(1):26. doi: 10.1186/1756-3305-1-26 18710577
20. Moyes CL, Henry AJ, Golding N, Huang Z, Singh B, Baird JK, et al. Defining the Geographical Range of the Plasmodium knowlesi Reservoir. PLoS Negl Trop Dis 2014; 8: e2780. doi: 10.1371/journal.pntd.0002780 24676231
21. Preston MD, Campino S, Assefa SA, Echeverry DF, Ocholla H, Amambua-Ngwa A, et al. A barcode of organellar genome polymorphisms identifies the geographic origin of Plasmodium falciparum strains. Nature Comm2014; 5, 4052. 24923250
22. Benavente ED, de Sessions PF, Moon RW, Grainger M, Holder AA, Blackman MJ, et al. A reference genome and methylome for the Plasmodium knowlesi malaria A1-H.1 line. Int J Parasit. In press. doi: 10.1645/12-11.1
23. Moon RW, Sharaf H, Hastings CH, Ho YS, Nair MB, Rchiad Z, et al. A. Normocyte-binding protein required for human erythrocyte invasion by the zoonotic malaria parasite Plasmodium knowlesi. Proc Natl Acad Sci U S A 2016; 113 : 7231–6. doi: 10.1073/pnas.1522469113 27303038
24. Srinivasan P, Fujioka H, Jacobs-Lorena M. PbCap380, a novel oocyst capsule protein, is essential for malaria parasite survival in the mosquito. Cellular Microbiology 2008; 10(6), 1304–1312. doi: 10.1111/j.1462-5822.2008.01127.x 18248630
25. Dessens JT, Beetsma AL, Dimopoulos G, Wengelnik K, Crisanti A, Kafatos FC, et al. CTRP is essential for mosquito infection by malaria ookinetes. The EMBO Journal 1999; 18(22), 6221–6227. doi: 10.1093/emboj/18.22.6221 10562534
26. López-Barragán MJ, Lemieux J, Quiñones M, Williamson KC, Molina-Cruz A, Cui K et al. Directional gene expression and antisense transcripts in sexual and asexual stages of Plasmodium falciparum. BMC Genomics 2011; 12(1), 587.
27. López-Barragán MJ, Lemieux J, Quiñones M, Williamson KC, Molina-Cruz A, Cui K, et al. A comprehensive evaluation of rodent malaria parasite genomes and gene expression. BMC Biology 2014; 12(1), 86.
28. Molina-Cruz A, Garver LS, Alabaster A, Bangiolo L, Haile A, Winikor J, et al. The human malaria parasite Pfs47 gene mediates evasion of the mosquito immune system. Science 2013; 340(6135):984–7. doi: 10.1126/science.1235264 23661646
29. Messenger LA, Llewellyn MS, Bhattacharyya T, Franzén O, Lewis MD, Ramírez JD, et al. Multiple mitochondrial introgression events and heteroplasmy in Trypanosoma cruzi revealed by Maxicircle MLST and Next Generation Sequencing. PLoS Negl Trop Dis 2012; 6(4), e1584. doi: 10.1371/journal.pntd.0001584 22506081
30. Messenger LA, Miles MA. Evidence and importance of genetic exchange among field populations of Trypanosoma cruzi. Acta Tropica 2015; 151, 150–155. doi: 10.1016/j.actatropica.2015.05.007 26188331
31. Brock PM, Fornace KM, Parmiter M, Cox J, Drakeley CJ, Ferguson HM, et al. Plasmodium knowlesi transmission: integrating quantitative approaches from epidemiology and ecology to understand malaria as a zoonosis. Parasitology 2016; 143(4), 389–400. doi: 10.1017/S0031182015001821 26817785
32. Vythilingam I, Wong ML, Wan-Yussof WS. Current status of Plasmodium knowlesi vectors: a public health concern? Parasitology 2016; 1–9.
33. Diez Benavente E, Ward Z, Chan W, Mohareb FR, Sutherland CJ, Roper C, et al. Genomic variation in Plasmodium vivax malaria reveals regions under selective pressure. PLOS ONE 2017; 12(5), e0177134. doi: 10.1371/journal.pone.0177134 28493919
34. Moyes CL, Shearer FM, Huang Z, Wiebe A, Gibson HS, Nijman V, et al. Predicting the geographical distributions of the macaque hosts and mosquito vectors of Plasmodium knowlesi malaria in forested and non-forested areas. Parasites & Vectors 2016; 9, 242.
35. Miettinen J, Shi C, Liew SC. Land cover distribution in the peatlands of Peninsular Malaysia, Sumatra and Borneo in 2015 with changes since 1990. Global Ecology and Conservation 2016; 6, 67–78.
36. Yasuoka J, Levins R. Impact of deforestation and agricultural development on anopheline ecology and malaria epidemiology. Am J Trop Med Hyg 2007; 76(3), 450–460. 17360867
37. Ansari HR, et al. Genome-scale comparison of expanded gene families in Plasmodium ovale wallikeri and Plasmodium ovale curtisi with Plasmodium malariae and with other Plasmodium species. Int J Parasitol 2016; 46(11):685–96. doi: 10.1016/j.ijpara.2016.05.009 27392654
38. Ansari HR, Templeton TJ, Subudhi AK, Ramaprasad A, Tang J, Lu F, et al. Estimating Geographical Variation in the Risk of Zoonotic Plasmodium knowlesi Infection in Countries Eliminating Malaria. PLOS Negl Trop Dis 2016; 10(8), e0004915. doi: 10.1371/journal.pntd.0004915 27494405
39. Fornace KM, Nuin NA, Betson M, Grigg MJ, William T, Anstey NM, et al. Asymptomatic and Submicroscopic Carriage of Plasmodium knowlesi Malaria in household and community members of clinical cases in Sabah, Malaysia. J Infect Dis 2016; 213(5), 784–787. doi: 10.1093/infdis/jiv475 26433222
40. Hayakawa T, Culleton R, Otani H, Horii T, Tanabe K. Big bang in the evolution of extant malaria parasites. Mol Biol Evol 2008; 25(10), 2233–2239. doi: 10.1093/molbev/msn171 18687771
41. Muehlenbein MP, Pacheco MA, Taylor JE, Prall SP, Ambu L, Nathan S, et al. Accelerated diversification of nonhuman primate malarias in Southeast Asia: adaptive radiation or geographic speciation? Molecular Biology and Evolution 2015; 32(2), 422–439. doi: 10.1093/molbev/msu310 25389206
42. Sutherland CJ. Persistent Parasitism: The Adaptive Biology of Malariae and Ovale Malaria. Trends in Parasitology 2016; 32(10), 808–819. doi: 10.1016/j.pt.2016.07.001 27480365
43. Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25(14), 1754–1760. doi: 10.1093/bioinformatics/btp324 19451168
44. Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011; 27(21), 2987–2993. doi: 10.1093/bioinformatics/btr509 21903627
45. Campino S, Benavente ED, Assefa S, Thompson E, Drought LG, Taylor CJ, et al. Genomic variation in two gametocyte non-producing Plasmodium falciparum clonal lines. Malaria J 2016; 15(1), 229. 27098483
46. Samad H, Coll F, Preston MD, Ocholla H, Fairhurst RM, Clark TG. Imputation-based population genetics analysis of Plasmodium falciparum malaria parasites. PLoS Genet. 2015; 11(4):e1005131. doi: 10.1371/journal.pgen.1005131 25928499
47. Assefa SA, Preston MD, Campino S, Ocholla H, Sutherland CJ, Clark TG. estMOI: estimating multiplicity of infection using parasite deep sequencing data. Bioinformatics 2014; 30(9), 1292–1294. doi: 10.1093/bioinformatics/btu005 24443379
48. Holsinger KE, Weir BS. Genetics in geographically structured populations: defining, estimating and interpreting FST. Nat Rev Genet 2009; 10(9), 639–650. doi: 10.1038/nrg2611 19687804
49. Sabeti PC, Varilly P, Fry B, Lohmueller J, Hostetter E, Cotsapas C, et al. Genome-wide detection and characterization of positive selection in human populations. Nature 2007; 449(7164), 913–918. doi: 10.1038/nature06250 17943131
50. Shimodaira H, Hasegawa H. Multiple comparisons of loglikelihoods with applications to phylogenetic inference. Mol Biol Evol 1999; 16, 1114.
51. Templeton AR. Phylogenetic inference from restriction endonuclease cleavage site maps with particular reference to the evolution of humans and the apes. Evolution 1983; 37, 221–244. doi: 10.1111/j.1558-5646.1983.tb05533.x 28568373
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2017 Číslo 9- Kazuistika – Perspektivy využití precizované medicíny v rámci personalizované specifické terapie onkologických pacientů
- Nobelova cena za chemii pro genetické nůžky: Objev, který změní naši budoucnost?
- Technologie na bázi RNA v klinické praxi: od přebarvených petúnií k terapii vzácných a dosud jen obtížně léčitelných chorob u lidí
- „Nepředstavovali jsme si, že náš výzkum povede přímo ke vzniku nových léků, dokonce ještě za našeho života“
- Bezplatné služby pro diagnostiku ATTRv amyloidózy pro kardiology
Nejčtenější v tomto čísle- "T-bet"-ing on autoimmunity variants
- Analysis of nuclear and organellar genomes of in humans reveals ancient population structure and recent recombination among host-specific subpopulations
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Autoři: prof. MUDr. Pavel Horák, CSc., doc. MUDr. Ludmila Brunerová, Ph.D., doc. MUDr. Václav Vyskočil, Ph.D., prim. MUDr. Richard Pikner, Ph.D., MUDr. Olga Růžičková, MUDr. Jan Rosa, prof. MUDr. Vladimír Palička, CSc., Dr.h.c.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



