-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Incorporation of tissue-based genomic biomarkers into localized prostate cancer clinics
Localized prostate cancer (PCa) is a clinically heterogeneous disease, which presents with variability in patient outcomes within the same risk stratification (low, intermediate or high) and even within the same Gleason scores. Genomic tools have been developed with the purpose of stratifying patients affected by this disease to help physicians personalize therapies and follow-up schemes. This review focuses on these tissue-based tools. At present, four genomic tools are commercially available: Decipher™, Oncotype DX®, Prolaris® and ProMark®. Decipher™ is a tool based on 22 genes and evaluates the risk of adverse outcomes (metastasis) after radical prostatectomy (RP). Oncotype DX® is based on 17 genes and focuses on the ability to predict outcomes (adverse pathology) in very low-low and low-intermediate PCa patients, while Prolaris® is built on a panel of 46 genes and is validated to evaluate outcomes for patients at low risk as well as patients who are affected by high risk PCa and post-RP. Finally, ProMark® is based on a multiplexed proteomics assay and predicts PCa aggressiveness in patients found with similar features to Oncotype DX®. These biomarkers can be helpful for post-biopsy decision-making in low risk patients and post-radical prostatectomy in selected risk groups. Further studies are needed to investigate the clinical benefit of these new technologies, the financial ramifications and how they should be utilized in clinics.
Keywords:
Prostate cancer, Radical prostatectomy, Genetic tools, Decipher, Oncotype DX, Prolaris
Authors: Marco Moschini 1; Martin Spahn 2; Agostino Mattei 3; John Cheville 4; R. Jeffrey Karnes 1*
Authors place of work: Department of Urology, Mayo Clinic, Rochester, MN, USA. 1; Department of Urology, University Hospital of Bern, Inselspital, Bern, Switzerland. 2; Klinik für Urologie, Luzerner Kantonsspital, Luzern, Switzerland. 3; Department of Laboratory Medicine and Pathology, Mayo Clinic , Rochester, MN, USA. 4
Published in the journal: BMC Medicine 2016, 14:67
Category: Review
doi: https://doi.org/10.1186/s12916-016-0613-7© 2016 Moschini et al. Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
The electronic version of this article is the complete one and can be found online at: https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-016-0613-7Summary
Localized prostate cancer (PCa) is a clinically heterogeneous disease, which presents with variability in patient outcomes within the same risk stratification (low, intermediate or high) and even within the same Gleason scores. Genomic tools have been developed with the purpose of stratifying patients affected by this disease to help physicians personalize therapies and follow-up schemes. This review focuses on these tissue-based tools. At present, four genomic tools are commercially available: Decipher™, Oncotype DX®, Prolaris® and ProMark®. Decipher™ is a tool based on 22 genes and evaluates the risk of adverse outcomes (metastasis) after radical prostatectomy (RP). Oncotype DX® is based on 17 genes and focuses on the ability to predict outcomes (adverse pathology) in very low-low and low-intermediate PCa patients, while Prolaris® is built on a panel of 46 genes and is validated to evaluate outcomes for patients at low risk as well as patients who are affected by high risk PCa and post-RP. Finally, ProMark® is based on a multiplexed proteomics assay and predicts PCa aggressiveness in patients found with similar features to Oncotype DX®. These biomarkers can be helpful for post-biopsy decision-making in low risk patients and post-radical prostatectomy in selected risk groups. Further studies are needed to investigate the clinical benefit of these new technologies, the financial ramifications and how they should be utilized in clinics.
Keywords:
Prostate cancer, Radical prostatectomy, Genetic tools, Decipher, Oncotype DX, ProlarisBackground
Prostate cancer (PCa) represents the most commonly diagnosed internal malignancy among men in the Western world. In 2015, the number of men diagnosed with PCa was estimated to be over 220,000 in the United States alone [1]. The introduction of prostate-specific antigen (PSA) in the late 80s and the associated ‘stage migration’ have led to an increase of PCa diagnoses and specifically lower risk disease [2]. Although a number of these newly diagnosed patients are potentially eligible for active surveillance (AS), a large percentage still undergo active treatment given some uncertainty as to whether they harbor a ‘higher risk’ disease. Additionally, the number of patients diagnosed with high risk disease with an elevated risk of metastasis and death is still considerable, and an increasing trend has been described [3, 4]. In this setting, it is pivotal to increase our ability over-and-above routine clinicopathologic variables (T stage, PSA, Gleason scores and other cancer characteristics from biopsies and/or pathology specimens) to identify patients that could benefit from an active treatment after prostate biopsy, as well as patients that are at added risk for recurrence(s) after radical prostatectomy (RP). This is fundamental considering the potential morbidity associated with any active treatment, such as radiation and/or RP for PCa. None of the above represents novel concepts; however, there now exists biomarkers that may allow for better risk stratification to help decide on treatment versus no treatment. Patients can often be categorized into subgroups using the routine clinicopathologic parameters. However, better understanding of PCa biology using tissue-based biomarkers might help clinicians provide more ‘personalized’ treatments.
In this context, not only is the clinical behavior of PCa heterogeneous but so are the genomic alteration patterns within the biologic pathways, and several different mutations are needed during tumorigenesis to develop an aggressive PCa. For this reason, studying and cataloging these albeit quite complex alterations [5, 6, 7] has contributed to the need for development of complex biomarkers or genomic panels that are combined to improve prognostication of localized PCa, and more so to tailor both primary or secondary treatment and follow-up. Clinicians, most frequently urologists, are often challenged how to best advise newly diagnosed male patients with low-to-intermediate (3 + 4) risk prostate cancer, to undergo active surveillance versus active treatment, and patients are often left wanting more information to make that decision. Even after a PCa has been treated by a RP, patients with adverse pathology (non-organ-confined disease) or those that develop biochemical recurrence (BCR) have a disparate risk of developing metastasis, and secondary treatment, often radiation, is advocated. As alluded earlier, there are inherent limitations in the ability to prognosticate a potentially lethal cancer using pathology reports and PSA values. Genomic sequencing of advanced PCa highlights the complexity of this disease and its different states [8, 9]. In this context, biomarker panels or genomic tools have been developed for more localized disease that perhaps help to ‘simplify’ a response to a complex question, i.e. is there a lethal component of this hormone-sensitive PCa in the treatment-naïve state or post-RP? The information provided by these tools are presented as a risk percentage; for example, the percentage risk of upgrading from biopsy to a higher Gleason score, such as 4 + 3 at RP, or percentage chance of BCR or metastasis after RP. These tools do not provide a binary answer to treat or not to treat for the clinician or patient. They use archived formalin-fixed, paraffin-embedded (FFPE) tissue of either the highest volume tumor foci or Gleason score from either a biopsy or RP specimen. Each assay is unique. To date, there are no head-to-head studies evaluating one of these panels over the other when a comparison is relevant. Prior to these biomarker or genomic panels, there were no routine tissue-based biomarkers that were used in practice. As mentioned, the goal is early detection of a potentially ‘lethal’ PCa to treat and separation of this from a potentially indolent course. The development of metastasis is usually lethal despite survival gains by different systemic therapies [10, 11, 12, 13, 14]. Considering the vast research and data in this field, several previous reviews assessed the importance of tissue-based biomarkers [15, 16, 17, 18]. In this context, we present an up-to-date overview of available tissue markers correlated with clinical utility in the field of PCa.
Commercially available tissue-based prognostic biomarker panels
Decipher™
Decipher™ is a genomic test, co-developed by GenomeDx Biosciences (Vancouver, BC, Canada) and Mayo Clinic, to assess the probability of developing metastases after RP (Table 1). This tool was developed by analyzing 1.4 million genomic markers, including coding and non-coding RNA. The signature is based on 22 expressed RNA biomarkers involved in various biologic pathways (cell differentiation, proliferation, structure, adhesion and motility, immune modulation, cell-cycle progression, androgen signaling) [19]. The outcome of the test is represented by a derived continuous risk score—a genomic classifier (GC) ranging from 0 to 1. Each score has an associated probability of metastasis at 5 years postoperatively, with 1 being the highest risk. Specifically, Decipher™ has been studied and approved in the United States for PCa patients treated with RP and adverse pathology (pT3 and/or positive margins or BCR) to evaluate the risk of developing BCR or clinical progression (metastasis) during follow-up. However, although these recommendations have been reported in the National Comprehensive Cancer Network (NCCN) guidelines [20], clinical use and long-term data are needed to judge the real added value [16].
Tab. 1. Characteristics of studies evaluating Decipher™ 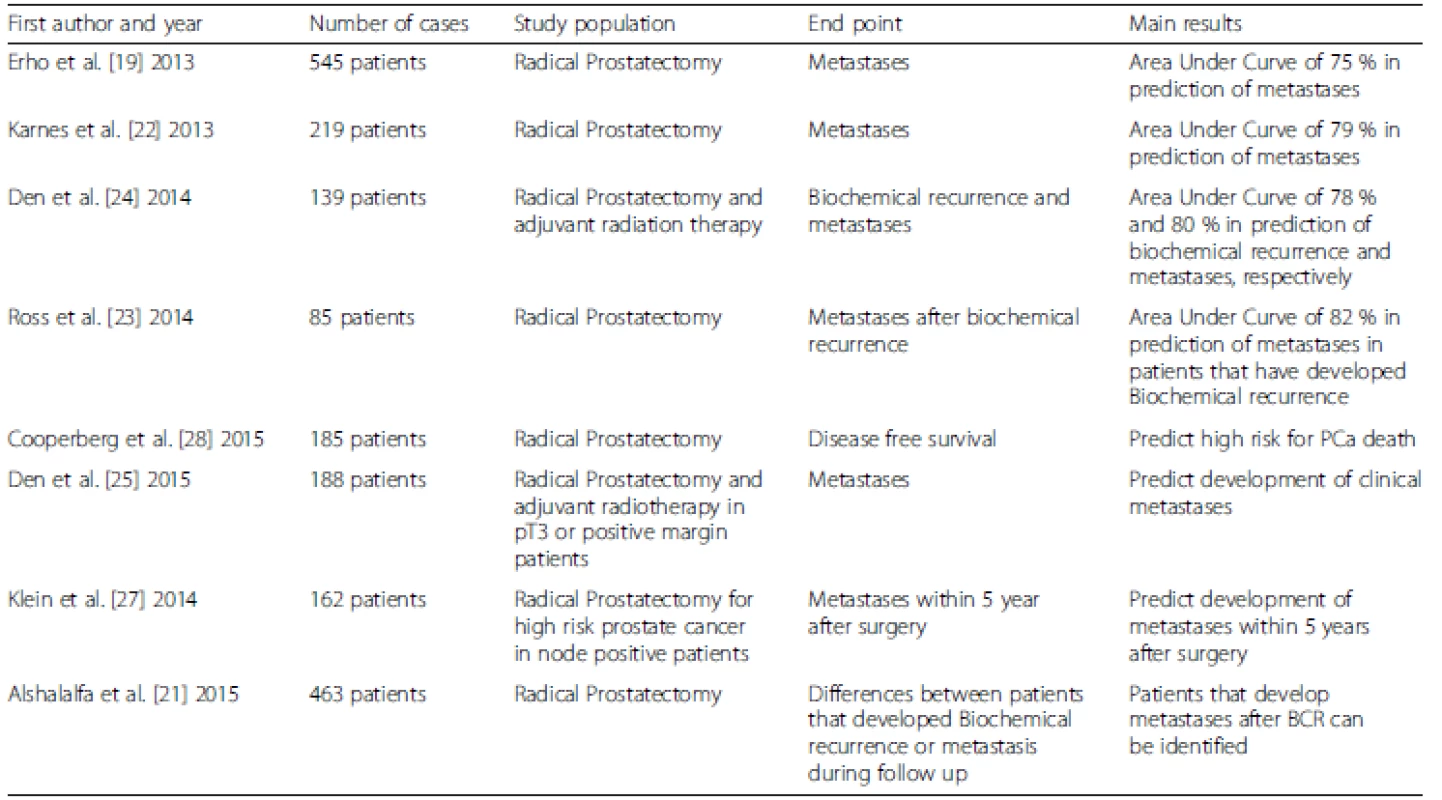
Natural history after BCR in RP PCa patients
Alshalalfa et al. [21] analyzed the differences in patients that developed BCR, clinical metastases or were free from any adverse outcomes during follow-up, and found that Decipher™ can represent a useful tool to better prognosticate patients who develop metastasis after BCR. In this study, patients without evidence of disease and those with BCR-only were found with similar transcriptional profiles, in contrast to patients that developed metastases and showed a distinct transcriptional profile that can be detected in the primary tumor/RP specimen. These findings were validated in 219 high risk PCa patients treated at Mayo Clinic with RP. On multivariable analyses, higher GC scores were most prognostic for the development of metastases [22]. Similar results were observed by Ross et al. [23] with 85 high risk patients who developed BCR after RP. Moreover, with this natural history study, i.e. without secondary treatments potentially confounding the results, the GC score was able to differentiate between patients that eventually progress after BCR and those that do not, indicating perhaps the patients to earlier intervene on versus those that might continue to be followed.
Immediate versus delayed adjuvant radiotherapy after RP
Den et al. [24] studied patients treated by RP and adjuvant radiotherapy (aRT) for pT3 or positive surgical margin PCa, and found the area under the curve (AUC) predicted BCR and clinical metastases of 78 % and 80 %, respectively. Patients with a lower GC score on their RP specimen were those who may benefit from delayed RT at further BCR, in comparison to patients with higher GC who may benefit from aRT. In a follow-up study [25], 188 patients from both Thomas Jefferson University and Mayo Clinic treated with RP and adjuvant or salvage RT were evaluated with a median follow-up of 10 years. The GC score was able to best prognosticate the occurrence of metastases, and provided further support that patients with low GC scores can probably be treated with observation and salvage RT as needed, while patients with high GC scores could benefit the most from aRT. This is a step towards balancing the risks and benefits of delivering aRT, since all these male patients would have been advised to consider aRT based on their RP pathology, and evidence that the GC score is not only prognostic but potentially predictive in regards to the timing of postoperative RT. Furthermore, a study by Lobo et al. [26] showed a benefit in quality-adjusted life-years (QALYs) by using the Decipher™ test in the postoperative setting.
Combination of validated clinical and genomic risk stratification for prediction of survival after RP
An improvement in the prognostication of clinical outcomes after RP has been described by Klein [27] and Cooperberg [28] by including validated clinical parameters and Decipher™ scores in the same model. The AUC in prognostication of metastases was 77 %, 75 % and 72 % for Decipher™, and the commonly used clinical scores from the Stephenson model and Cancer of the Prostate Risk Assessment Score (CAPRA-S), respectively. However, when Decipher™ was integrated into the Stephenson nomogram, the AUC rose to 79 %. Similar results were presented for cancer-specific mortality (CSM) after RP; in fact, the AUCs were 75 % and 78 % for CAPRA-S and Decipher™, respectively. Patients with both high GC and CAPRA-S risk scores were at major risk for CSM. Moreover, Decipher™ was able to reclassify many patients stratified to high risk based on CAPRA-S to an actual lower risk based on a low GC, thus potentially sparing patients postoperative therapy, and at least providing less worry to the patients.
Oncotype DX®
Oncotype DX® is a test that has been developed by Genomic Health (Redwood City, CA, USA) for prostate cancer, and is well known for its prognostic and predictive capability in breast oncology. This tool is a quantitative RT-PCR assay performed on FFPE tissue from needle biopsies, incorporating 12 cancer-related genes representing four biological pathways (stromal response, androgen signaling, proliferation, cellular organization) and five reference genes, which are algorithmically combined to calculate the Genomic Prostate Score (GPS) [29]. The derived GPS ranges from 0 to 100, with the higher number correlating with a higher probability of harboring adverse pathology at RP in men diagnosed with low or low-intermediate risk prostate cancer at prostate biopsy (Table 2). Ostensibly, this test is utilized to counsel men regarding the best course of therapy (active surveillance vs. treatment) in newly diagnosed low or low-intermediate PCa. Specifically, NCCN guidelines recommend its utilization in post-biopsy decision-making for NCCN very low and low risk PCa at diagnosis with 10–20 years’ life expectancy [20].
Tab. 2. Characteristics of studies evaluating OncotypeDX<sup>®</sup> 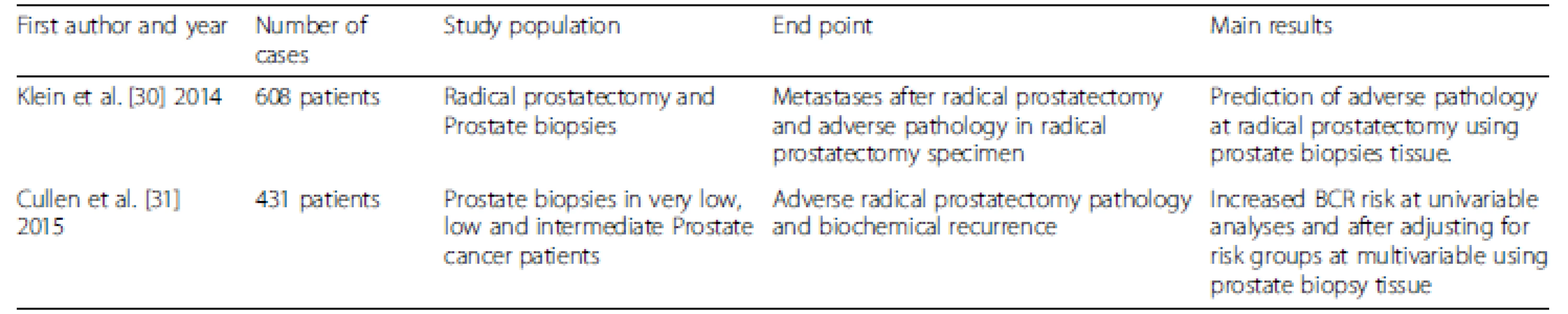
Prediction of survival outcomes in low or intermediate risk PCa
Klein et al. [30] designed an initial study including 395 men who had low to intermediate risk prostate cancer who underwent RP. The biopsy-based 17-gene GPS improved on the prediction of the presence or absence of adverse pathology, defined as primary Gleason score pattern of 4 or any pattern of 5, or non-organ-confined disease. Cullen et al. [31] described 431 patients with NCCN very low, low and intermediate risk-stratified PCa at biopsy, and demonstrated the ability of Oncotype DX® on a biopsy specimen (largest tumor foci) to independently better predict adverse pathology at RP and consequently BCR than the risk group variables.
Prolaris®
Prolaris® is also a prognostic test developed by Myriad Genetics (Salt Lake City, UT, USA). This test is based on the expression of 31 cell-cycle progression (CCP) and 15 housekeeping genes (Table 3), and is an extension of their breast cancer test. The result is represented by a proliferative index, expressed as a CCP score [32]. The result has shown improved prognostic ability over clinicopathologic variables in various settings and tissues (biopsy and RP), and for various outcomes. This tool is used on biopsies from low and very low risk men to help better determine if immediate or deferred (conservatively managed) treatment is the better course. Again, like others, it is a prognostic assay and does not provide an answer to the treatment conundrums, but a probability of an event in the future, such as BCR and prostate cancer-specific mortality. In context, NCCN recommends its use in the post-biopsy period for NCCN very low and low risk PCa at diagnosis for patients with at least 10 years’ life expectancy [20].
Tab. 3. Characteristics of studies evaluating Prolaris<sup>®</sup> 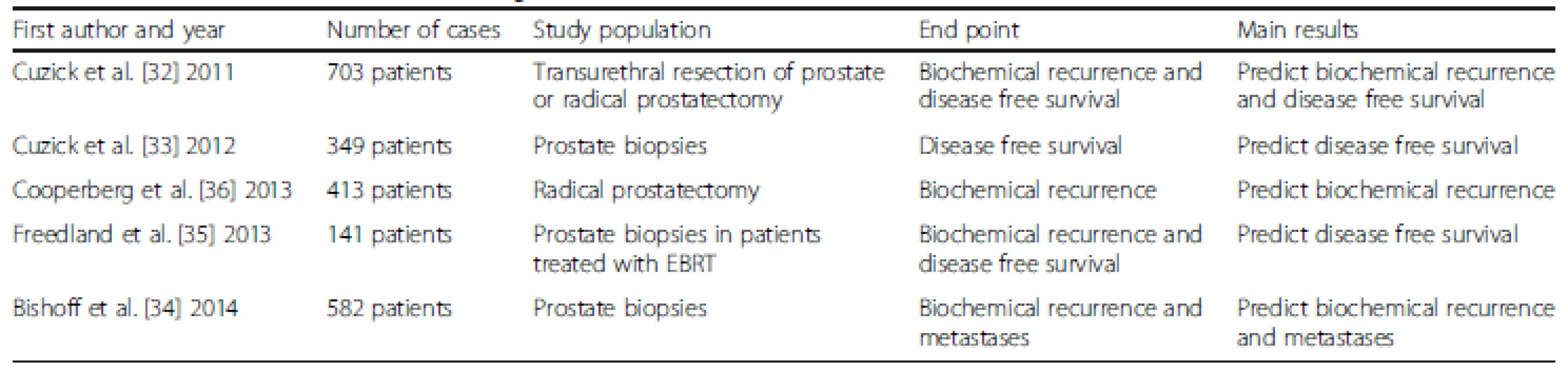
Prediction of survival outcomes in low or very low risk patients
Cuzick et al. [33] investigated 349 prostate biopsy samples evaluating the impact of CCP score—Prolaris®—on disease-free survival. At multivariable analysis, the CCP score showed a hazard ratio of 1.65, which was the strongest factor. Bishoff et al. [34] analyzed 582 patients from the Martini Clinic (n = 283), Durham Veterans Affairs Medical Center (n = 176) and Intermountain Healthcare (n = 123), and showed an association of the biopsy CCP score with adverse outcomes after surgery. In a larger study, Cuzick et al. [32] evaluated the CCP score and its ability of prognosticating BCR in a US cohort of RP patients and mortality in a UK cohort of patients treated with transurethral resection of the prostate (TURP). Additionally, the CCP was validated by Freedland et al. [35] in primary external radiation-treated patients. Cooperberg et al. [36] analyzed 413 RP patients with a CCP score in prognosticating BCR, and the best AUC came from a combined CCP score and CAPRA-S for the overall cohort and the low risk subset.
ProMark®
ProMark® is the newest commercially available biomarker panel and is a protein-based test developed by Metamark (Cambridge, MA, USA) as a quantitative multiplexed proteomics assay. Originally, there were 12 protein markers that demonstrated the best prognostication from a biopsy specimen [37]. Subsequently, the signature was refined and includes eight protein markers [38], with a score (0–1) reflecting the probability of adverse pathology at RP. To our knowledge, there is a lack of validation of this panel, most likely related to the more recent development.
Prediction of aggressiveness of prostate biopsy samples
Blume-Jensen et al. [38] found a favorable biomarker risk score (defined as ≤ 0.33) and a non-favorable risk score (defined as > 0.80) in 10 % and 5 % patients, respectively. The predictive value for non-favorable pathology was 76.9 % considering a biomarker risk score of > 0.80. At a biomarker risk score of ≤ 0.33 evaluating the prediction of favorable pathology disease, the predictive values were 95 %, 81.5 % and 87.2 % in NCCN very low and low risk, and low risk D’Amico groups, respectively [38].
ConfirmMDx
ConfirmMDx is a methylation marker genetic test developed by MDxHealth (Irvine, CA, USA). The test records a 90 % negative predictive value within 30 months of the initial biopsy, and has been reported as the most significant predictor of biopsy results [39]. The impact of this test on re-biopsy was recently assessed in 138 patients; among those with a negative ConfirmMDx assay, only 6 patients (4 %) underwent repeat biopsies [40]. ConfirmMDx may therefore be of value in the decision of repeat biopsies after a negative initial prostate biopsy.
Other available tissue-based markers
Ki-67
Ki-67 is a nuclear protein associated with ribosomal RNA synthesis and has been typically measured by semi-quantitative immunohistochemistry (IHC) to assay cell proliferation in cancer. Although not tied to a commercial entity nor licensed, the IHC is typically performed using a MIB-1 antibody and reported as a percent of cells staining positively, which reflects their proliferation and correlates with various cancer outcomes, including PCa. Although several reports have assessed the impact of Ki-67 evaluation at the time of biopsy or after RP, at the present time no clinical use has been recommended by NCCN, which recently excluded the use of this test from the recommendation [20].
Prostate biopsies
The prognostic value of K1-67 in prostate needle biopsies are documented by several authors [41, 42, 43]. Tollefson et al. [44] most recently reported data of 451 consecutive patients with biopsy-proven PCa treated with RP at Mayo Clinic. The results confirmed that a combination of Gleason score, perineural invasion and Ki-67 expression is the most effective predictor of cancer outcome at the time of prostate needle biopsy. It is not known how this single marker of cell proliferation compares to the CCP. Ki-67 staining index on biopsies has been tested on 573 patients treated with PCa radiotherapy with a median follow-up of 96 months [45]. The Ki67 staining index was the most significant determinant of distant metastases and cancer-specific mortality during follow-up.
Outcomes after radical prostatectomy for localized PCa
Ki-67 staining index has also been reported to be a powerful predictor of survival after RP [42, 46, 47]. Furthermore, Mathieu et al. [48] analyzed data of 3,123 patients who underwent RP and confirmed that Ki-67 is a strong prognostic variable for BCR, and might be informative in clinical decision-making regarding adjuvant therapy and optimizing follow-up schedules.
PTEN
Considering mutations that have a well-established role in PCa, dysregulation of PTEN seems to be involved more frequently than others in advanced, localized or metastatic PCa, and has shown a prognostic impact [49, 50, 51, 52]. PTEN is located on chromosome 10 and its mutation is considered a part of the PI3K/AKT signaling pathway and functions as a tumor suppressor gene. PTEN appears to be the most frequently mutated tumor suppressor gene and deletion of PTEN has been recorded in 10–70 % of RP specimens [53, 54, 55, 56, 57], with the variability likely related to the risk group, with the higher the risk the higher likelihood of a PTEN deletion. A recent publication by Murphy et al. [58] demonstrated that PTEN loss is infrequent in clinically insignificant PCa (Gleason score 6 and low volume) and is associated with higher-grade tumors. This finding is also supported by others whether on biopsy [59, 60] or RP specimens [49, 61] and in castration-resistant tumors [62]. Again, it is unknown how a single gene may perform in relation to an aforementioned panel and what the prognostic differences would be. The costs of a single gene assay should be cheaper than a panel, but given the heterogeneity of PCa in terms of foci and different biologic pathways there is an obvious potential to not completely capture what is needed with a single marker. As per Ki-67 marker, its use has been excluded from the clinical recommendation by NCCN [20].
Conclusions
Available genomic assays have improved the prognostic ability over clinicopathologic parameters of localized PCa. Ideally, these assays should be prospectively applied, or even retrospectively applied to prospective studies, to further validate their clinical utility in prognostication and even prediction in terms of what treatment should be applied either at a new diagnosis or post-RP. In addition to their clinical value, more work is needed in regards to their financial impact on the cost of localized PCa care.
Competing interests
JC and RJK are employees of Mayo Clinic. As there is a partnership with GenomeDx and Mayo Clinic that co-developed Decipher, Mayo Clinic has a financial interest in Decipher. MM, MS and AM declare that they have no competing interests.
Authors’ contributions
MM drafted the manuscript, and undertook analysis and interpretation of data. MS conceived, designed and supervised the study, and critically revised and drafted the manuscript. AM supervised the study and critically revised the manuscript. JC critically revised the manuscript. RJK conceived, designed and supervised the study, analyzed and interpreted the data, and drafted and critically revised the manuscript. All authors read and approved the final manuscript.
Authors’ information
MM MD is a PhD student and research fellow at Mayo Clinic. MS MD is Associate Professor and Consultant at the Department of Urology, University Hospital of Bern. AM MD is Chairman and Professor at the Department of Urology, Luzerner Kantonsspital. JC MD is Professor of Laboratory Medicine and Pathology at Mayo Clinic. RJK MD is Vice-chair of Urology at the Mayo Clinic. JC and RJK are co-directors of the Prostate Biomarker Discovery Program, Center of Individualized Medicine at Mayo Clinic.
Received: 12 September 2015
Accepted: 30 March 2016
Published: 4 April 2016* Correspondence:
R. Jeffrey Karnes
Department of Urology
Mayo Clinic, Rochester
MN, USAkarnes.r@mayo.edu
Zdroje
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65 : 5–29. doi:10.3322/caac.21254.
2. Shao YH, Demissie K, Shih W, Mehta AR, Stein MN, Roberts CB, et al. Contemporary risk profile of prostate cancer in the United States. J Natl Cancer Inst. 2009;101 : 1280–3. doi:10.1093/jnci/djp262.
3. Budäus L, Spethmann J, Isbarn H, Schmitges J, Beesch L, Haese A, et al. Inverse stage migration in patients undergoing radical prostatectomy: results of 8916 European patients treated within the last decade. BJU Int. 2011;108 : 1256–61. doi:10.1111/j.1464-410X.2010.09982.x.
4. Silberstein JL, Vickers AJ, Power NE, Fine SW, Scardino PT, Eastham JA, et al. Reverse stage shift at a tertiary care center: escalating risk in men undergoing radical prostatectomy. Cancer. 2011;117 : 4855–60. doi:10.1002/cncr.26132.
5. Barbieri CE, Bangma CH, Bjartell A, Catto JWF, Culig Z, Grönberg H, et al. The mutational landscape of prostate cancer. Eur Urol. 2013;64 : 567–76. doi:10.1016/j.eururo.2013.05.029.
6. Haffner MC, Mosbruger T, Esopi DM, Fedor H, Heaphy CM, Walker DA, et al. Tracking the clonal origin of lethal prostate cancer. J Clin Invest. 2013;123 : 4918–22. doi:10.1172/JCI70354.
7. Kumar A, White TA, MacKenzie AP, Clegg N, Lee C, Dumpit RF, et al. Exome sequencing identifies a spectrum of mutation frequencies in advanced and lethal prostate cancers. Proc Natl Acad Sci U S A. 2011;108 : 17087–92. doi:10.1073/pnas.1108745108.
8. Grasso CS, Wu YM, Robinson DR, Cao X, Dhanasekaran SM, Khan AP, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature. 2012;487 : 239–43. doi:10.1038/nature11125.
9. Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18 : 11–22. doi:10.1016/j.ccr.2010.05.026.
10. Fizazi K, Scher HI, Molina A, Logothetis CJ, Chi KN, Jones RJ, et al. Abiraterone acetate for treatment of metastatic castration-resistant prostate cancer: final overall survival analysis of the COU-AA-301 randomised, double-blind, placebocontrolled phase 3 study. Lancet Oncol. 2012;13 : 983–92. doi:10.1016/S1470-2045(12)70379-0.
11. Scher HI, Fizazi K, Saad F, Taplin ME, Sternberg CN, Miller K, et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N Engl J Med. 2012;367 : 1187–97. doi:10.1056/NEJMoa1207506.
12. Beer TM, Tombal B. Enzalutamide in metastatic prostate cancer before chemotherapy. N Engl J Med. 2014;371 : 1755–6. doi:10.1056/NEJMc1410239.
13. Tannock IF, de Wit R, Berry WR, Horti J, Pluzanska A, Chi KN, et al. Docetaxel plus prednisone or mitoxantrone plus prednisone for advanced prostate cancer. N Engl J Med. 2004;351 : 1502–12. doi:10.1056/NEJMoa040720.
14. de Bono JS, Oudard S, Ozguroglu M, Hansen S, MacHiels JP, Kocak I, et al. Prednisone plus cabazitaxel or mitoxantrone for metastatic castrationresistant prostate cancer progressing after docetaxel treatment: a randomised open-label trial. Lancet. 2010;376 : 1147–54. doi:10.1016/S0140-6736(10)61389-X.
15. Sternberg IA, Vela I, Scardino PT. Molecular profiles of prostate cancer: to treat or not to treat. Annu Rev Med. 2015;67 : 119–35. doi:10.1146/annurevmed-060413-112226.
16. Boström PJ, Bjartell AS, Catto JWF, Eggener SE, Lilja H, Loeb S, et al. Genomic predictors of outcome in prostate cancer. Eur Urol. 2015;68 : 1033–44. doi:10.1016/j.eururo.2015.04.008.
17. Sartori DA, Chan DW. Biomarkers in prostate cancer: what’s new? Curr Opin Oncol. 2014;26 : 259–64. doi:10.1097/CCO.0000000000000065.
18. Spahn M, Boxler S, Joniau S, Moschini M, Tombal B, Karnes RJ. What is the need for prostatic biomarkers in prostate cancer management? Curr Urol Rep. 2015;16 : 70. doi:10.1007/s11934-015-0545-3.
19. Erho N, Crisan A, Vergara IA, Mitra AP, Ghadessi M, Buerki C, et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS One. 2013;8, e66855. doi:10.1371/journal.pone.0066855.
20. Mohler JL, Armstrong AJ, Bahnson RR, D’Amico AV, Davis BJ, Eastham JA, et al. Prostate Cancer, Version 1.2016. J Natl Compr Canc Netw. 2016;14 : 19–30.
21. Alshalalfa M, Crisan A, Vergara IA, Ghadessi M, Buerki C, Erho N, et al. Clinical and genomic analysis of metastatic prostate cancer progression with a background of postoperative biochemical recurrence. BJU Int. 2015;116 : 556–67. doi:10.1111/bju.13013.
22. Karnes RJ, Bergstralh EJ, Davicioni E, Ghadessi M, Buerki C, Mitra AP, et al. Validation of a genomic classifier that predicts metastasis following radical prostatectomy in an at risk patient population. J Urol. 2013;190 : 2047–53. doi:10.1016/j.juro.2013.06.017.
23. Ross AE, Feng FY, Ghadessi M, Erho N, Crisan A, Buerki C, et al. A genomic classifier predicting metastatic disease progression in men with biochemical recurrence after prostatectomy. Prostate Cancer Prostatic Dis. 2014;17 : 64–9. doi:10.1038/pcan.2013.49.
24. Den RB, Feng FY, Showalter TN, Mishra MV, Trabulsi EJ, Lallas CD, et al. Genomic prostate cancer classifier predicts biochemical failure and metastases in patients after postoperative radiation therapy. Int J Radiat Oncol. 2014;89 : 1038–46. doi:10.1016/j.ijrobp.2014.04.052.
25. Den RB, Yousefi K, Trabulsi EJ, Abdollah F, Choeurng V, Feng FY, et al. Genomic classifier identifies men with adverse pathology after radical prostatectomy who benefit from adjuvant radiation therapy. J Clin Oncol. 2015;33 : 944–51. doi:10.1200/JCO.2014.59.0026.
26. Lobo JM, Dicker AP, Buerki C, Daviconi E, Karnes RJ, Jenkins RB, et al. Evaluating the clinical impact of a genomic classifier in prostate cancer using individualized decision analysis. PLoS One. 2015;10, e0116866. doi:10.1371/journal.pone.0116866.
27. Klein EA, Yousefi K, Haddad Z, Choeurng V, Buerki C, Stephenson AJ, et al. A genomic classifier improves prediction of metastatic disease within 5 years after surgery in node-negative high-risk prostate cancer patients managed by radical prostatectomy without adjuvant therapy. Eur Urol. 2014;67 : 778–86. doi:10.1016/j.eururo.2014.10.036.
28. Cooperberg MR, Davicioni E, Crisan A, Jenkins RB, Ghadessi M, Karnes RJ. Combined value of validated clinical and genomic risk stratification tools for predicting prostate cancer mortality in a high-risk prostatectomy cohort. Eur Urol. 2015;67 : 326–33. doi:10.1016/j.eururo.2014.05.039.
29. Knezevic D, Goddard AD, Natraj N, Cherbavaz DB, Clark-Langone KM, Snable J, et al. Analytical validation of the Oncotype DX prostate cancer assay – a clinical RT-PCR assay optimized for prostate needle biopsies. BMC Genomics. 2013;14 : 690. doi:10.1186/1471-2164-14-690.
30. Klein EA, Cooperberg MR, Magi-Galluzzi C, Simko JP, Falzarano SM, Maddala T, et al. A 17-gene assay to predict prostate cancer aggressiveness in the context of gleason grade heterogeneity, tumor multifocality, and biopsy undersampling. Eur Urol. 2014;66 : 550–60. doi:10.1016/j.eururo.2014.05.004.
31. Cullen J, Rosner IL, Brand TC, Zhang N, Tsiatis AC, Moncur J, et al. A biopsybased 17-gene genomic prostate score predicts recurrence after radical prostatectomy and adverse surgical pathology in a racially diverse population of men with clinically low - and intermediate-risk prostate cancer. Eur Urol. 2015;68 : 123–31. doi:10.1016/j.eururo.2014.11.030.
32. Cuzick J, Swanson GP, Fisher G, Brothman AR, Berney DM, Reid JE, et al. Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: a retrospective study. Lancet Oncol. 2011;12 : 245–55. doi:10.1016/S1470-2045(10)70295-3.
33. Cuzick J, Berney DM, Fisher G, Mesher D, Møller H, Reid JE, et al. Prognostic value of a cell cycle progression signature for prostate cancer death in a conservatively managed needle biopsy cohort. Br J Cancer. 2012;106 : 1095–9. doi:10.1038/bjc.2012.39.
34. Bishoff JT, Freedland SJ, Gerber L, Tennstedt P, Reid J, Welbourn W, et al. Prognostic utility of the cell cycle progression score generated from biopsy in men treated with prostatectomy. J Urol. 2014;192 : 409–14. doi:10.1016/j.juro.2014.02.003.
35. Freedland SJ, Gerber L, Reid J, Welbourn W, Tikishvili E, Park J, et al. Prognostic utility of cell cycle progression score in men with prostate cancer after primary external beam radiation therapy. Int J Radiat Oncol Biol Phys. 2013;86 : 848–53. doi:10.1016/j.ijrobp.2013.04.043.
36. Cooperberg MR, Simko JP, Cowan JE, Reid JE, Djalilvand A, Bhatnagar S, et al. Validation of a cell-cycle progression gene panel to improve risk stratification in a contemporary prostatectomy cohort. J Clin Oncol. 2013;31 : 1428–34. doi:10.1200/JCO.2012.46.4396.
37. Shipitsin M, Small C, Choudhury S, Giladi E, Friedlander S, Nardone J, et al. Identification of proteomic biomarkers predicting prostate cancer aggressiveness and lethality despite biopsy-sampling error. Br J Cancer. 2014;111 : 1201–12. doi:10.1038/bjc.2014.396.
38. Blume-Jensen P, Berman DM, Rimm DL, Shipitsin M, Putzi M, Nifong TP, et al. Development and clinical validation of an in situ biopsy-based multimarker assay for risk stratification in prostate cancer. Clin Cancer Res. 2015;21 : 2591–600. doi:10.1158/1078-0432.CCR-14-2603.
39. Partin AW, Van Neste L, Klein EA, Marks LS, Gee JR, Troyer DA, et al. Clinical validation of an epigenetic assay to predict negative histopathological results in repeat prostate biopsies. J Urol. 2014;192 : 1081–7. doi:10.1016/j.juro.2014.04.013.
40. Wojno KJ, Costa FJ, Cornell RJ, Small JD, Pasin E, Van Criekinge W, et al. Reduced rate of repeated prostate biopsies observed in ConfirmMDx clinical utility field study. Am Heal Drug Benefits. 2014;7 : 129–34.
41. Fisher G, Yang ZH, Kudahetti S, Møller H, Scardino P, Cuzick J, et al. Prognostic value of Ki-67 for prostate cancer death in a conservatively managed cohort. Br J Cancer. 2013;108 : 271–7. doi:10.1038/bjc.2012.598.
42. Rubio J, Ramos D, López-Guerrero JA, Iborra I, Collado A, Solsona E, et al. Immunohistochemical expression of Ki-67 antigen, cox-2 and Bax/Bcl-2 in prostate cancer; prognostic value in biopsies and radical prostatectomy specimens. Eur Urol. 2005;48 : 745–51. doi:10.1016/j.eururo.2005.06.014.
43. Jhavar S, Bartlett J, Kovacs G, Corbishley C, Dearnaley D, Eeles R, et al. Biopsy tissue microarray study of Ki-67 expression in untreated, localized prostate cancer managed by active surveillance. Prostate Cancer Prostatic Dis. 2009;12 : 143–7. doi:10.1038/pcan.2008.47.
44. Tollefson MK, Karnes RJ, Kwon ED, Lohse CM, Rangel LJ, Mynderse LA, et al. Prostate cancer Ki-67 (MIB-1) expression, perineural invasion, and gleason score as biopsy-based predictors of prostate cancer mortality: the Mayo model. Mayo Clin Proc. 2014;89 : 308–18. doi:10.1016/j.mayocp.2013.12.001.
45. Pollack A, DeSilvio M, Khor LY, Li R, Al-Saleem TI, Hammond ME, et al. Ki-67 staining is a strong predictor of distant metastasis and mortality for men with prostate cancer treated with radiotherapy plus androgen deprivation: Radiation Therapy Oncology Group Trial 92-02. J Clin Oncol. 2004;22 : 2133–40. doi:10.1200/JCO.2004.09.150.
46. Khatami A, Hugosson J, Wang W, Damber JE. Ki-67 in screen-detected, low-grade, low-stage prostate cancer, relation to prostate-specific antigen doubling time, Gleason score and prostate-specific antigen relapse after radical prostatectomy. Scand J Urol Nephrol. 2009;43 : 12–8. doi:10.1080/00365590802469543.
47. Aaltomaa S, Kärjä V, Lipponen P, Isotalo T, Kankkunen JP, Talja M, et al. Expression of Ki-67, cyclin D1 and apoptosis markers correlated with survival in prostate cancer patients treated by radical prostatectomy. Anticancer Res. 2006;26 : 4873–8.
48. Mathieu R, Shariat SF, Seitz C, Karakiewicz PI, Fajkovic H, Sun M, et al. Multiinstitutional validation of the prognostic value of Ki-67 labeling index in patients treated with radical prostatectomy. World J Urol. 2015;33 : 1165–71. doi:10.1007/s00345-014-1421-3.
49. Leinonen KA, Saramäki OR, Furusato B, Kimura T, Takahashi H, Egawa S, et al. Loss of PTEN is associated with aggressive behavior in ERG-positive prostate cancer. Cancer Epidemiol Biomarkers Prev. 2013;22 : 2333–44. doi:10.1158/1055-9965.EPI-13-0333-T.
50. Yoshimoto M, Joshua AM, Cunha IW, Coudry RA, Fonseca FP, Ludkovski O, et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod Pathol. 2008;21 : 1451–60. doi:10.1038/modpathol.2008.96.
51. Krohn A, Diedler T, Burkhardt L, Mayer PS, De Silva C, Meyer-Kornblum M, et al. Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am J Pathol. 2012;181 : 401–12. doi:10.1016/j.ajpath.2012.04.026.
52. McCall P, Witton CJ, Grimsley S, Nielsen KV, Edwards J. Is PTEN loss associated with clinical outcome measures in human prostate cancer? Br J Cancer. 2008;99 : 1296–301. doi:10.1038/sj.bjc.6604680.
53. Cairns P, Okami K, Halachmi S, Halachmi N, Esteller M, Herman JG, et al. Frequent inactivation of PTEN/MMAC1 in primary prostate cancer. Cancer Res. 1997;57 : 4997–5000.
54. Feilotter HE, Nagai MA, Boag AH, Eng C, Mulligan LM. Analysis of PTEN and the 10q23 region in primary prostate carcinomas. Oncogene. 1998;16 : 1743–8. doi:10.1038/sj.onc.1200205.
55. Pesche S, Latil A, Muzeau F, Cussenot O, Fournier G, Longy M, et al. PTEN/MMAC1/TEP1 involvement in primary prostate cancers. Oncogene. 1998;16 : 2879–83. doi:10.1038/sj.onc.1202081.
56. Wang SI, Parsons R, Ittmann M. Homozygous deletion of the PTEN tumor suppressor gene in a subset of prostate adenocarcinomas. Clin Cancer Res. 1998;4 : 811–5.
57. Whang YE, Wu X, Suzuki H, Reiter RE, Tran C, Vessella RL, et al. Inactivation of the tumor suppressor PTEN/MMAC1 in advanced human prostate cancer through loss of expression. Proc Natl Acad Sci U S A. 1998;95 : 5246–50.
58. Murphy SJ, Karnes RJ, Kosari F, Castellar BE, Kipp BR, Johnson SH, et al. Integrated analysis of the genomic instability of PTEN in clinically insignificant and significant prostate cancer. Mod Pathol. 2016;29 : 143–56. doi:10.1038/modpathol.2015.136.
59. Mithal P, Allott E, Gerber L, Reid J, Welbourn W, Tikishvili E, et al. PTEN loss in biopsy tissue predicts poor clinical outcomes in prostate cancer. Int J Urol. 2014;21 : 1209–14. doi:10.1111/iju.12571.
60. Lotan TL, Carvalho FLF, Peskoe SB, Hicks JL, Good J, Fedor HL, et al. PTEN loss is associated with upgrading of prostate cancer from biopsy to radical prostatectomy. Mod Pathol. 2015;28 : 128–37. doi:10.1038/modpathol.2014.85.
61. Lotan TL, Gurel B, Sutcliffe S, Esopi D, Liu W, Xu J, et al. PTEN protein loss by immunostaining: analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin Cancer Res. 2011;17 : 6563–73. doi:10.1158/1078-0432.CCR-11-1244.
62. Ferraldeschi R, Nava Rodrigues D, Riisnaes R, Miranda S, Figueiredo I, Rescigno P, et al. PTEN protein loss and clinical outcome from castrationresistant prostate cancer treated with abiraterone acetate. Eur Urol. 2015;67 : 795–802. doi:10.1016/j.eururo.2014.10.027.
Článek vyšel v časopiseBMC Medicine
Nejčtenější tento týden
2016 Číslo 67- Bezpečnost dlouhodobé terapie osteoporózy – aktuální data
- Máj pod bílým pláštěm aneb když mezi směnami vykvete láska
- Dlouhodobá recidiva a komplikace spojené s elektivní operací břišní kýly
- Mluvit bez hlasivek? Pomoci by měla nová technologie s podporou AI
- Na inkontinenční pomůcky nově dosáhne více pacientů
Nejčtenější v tomto čísle
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: MUDr. Petr Výborný, CSc., FEBO
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



