-
Medical journals
- Career
Detection of the etiological agents of hospital-acquired pneumonia – validity and comparison of different types of biological sample collection: a prospective, observational study in intensive care patients
Authors: L. Doubravská 1; R. Uvízl 1; T. Herkeľ 1; M. Kolář 2; T. Gabrhelík 3; M. Röderová 2; M. Htoutou Sedláková 2; K. Langová 4; V. Kolek 5; P. Jakubec 5; M. Adamus 1
Authors‘ workplace: Department of Anesthesiology and Intensive Care Medicine, University Hospital Olomouc and Faculty of Medicine and Dentistry, Palacký University Olomouc 1; Department of Microbiology, University Hospital Olomouc and Faculty of Medicine and Dentistry, Palacký University Olomouc 2; Department of Anesthesiology, Tomas Bata Hospital, Zlín 3; Department of Medical Biophysics, Faculty of Medicine and Dentistry, Palacký University Olomouc 4; Department of Respiratory Medicine, University Hospital Olomouc and Faculty of Medicine and Dentistry, Palacký University Olomouc 5
Published in: Epidemiol. Mikrobiol. Imunol. 66, 2017, č. 4, s. 155-162
Category: Original Papers
Overview
Background:
There is still a lack of evidence as to which method of biological sample collection is optimal for identifying bacterial pathogens causing hospital-acquired pneumonia (HAP). Much effort has been made to find an easy and valid approach to be used in clinical practice.Methods:
The primary endpoint of this prospective, observational study was to determine the predictive value of oropharyngeal swab (OS) and gastric aspiration (GA) as simple and non-invasive methods for diagnosing HAP. Their efficacy was compared to endotracheal aspiration (ETA) and protected specimen brushing (PSB), the standard methods approved for HAP diagnosis.Results:
Initially, 56 patients were enrolled. Significant amounts of bacterial pathogens were detected in 48 patients (79 isolates) in Round A and in 39 patients (45 isolates) in Round B (after 72 hours). The sensitivity rates were: ETA 98%, PSB 31%, OS 64% and GA 67% in Round A and ETA 87%, PSB 32%, OS 74% and GA 42% in Round B. Strains of 12 bacterial species were identified in the samples. The three most common etiological agents (both rounds together) were Klebsiella pneumoniae (23.7%), Burkholderia multivorans (21.1%) and Pseudomonas aeruginosa (15.8%).Conclusions:
Blind ETA is an optimum method for obtaining biological samples for identification of etiological agents causing HAP in intubated patients. Microbial etiological agents were more frequently detected in ETA samples than in those collected by PSB. If ETA/PSB results are negative, samples may be collected by OS and/or GA as these techniques followed ETA in terms of the frequency of pathogen detection.Keywords:
hospital-acquired pneumonia – bacterial pathogens – tracheal aspiration – protected specimen brushingINTRODUCTION
Hospital-acquired pneumonia (HAP) accounts for 10–50% of nosocomial infections in the intensive care unit (ICU) [1–3] and for more than 50% of the antibiotics prescribed [4]. Mechanical ventilation and intubation increase the risk of HAP 3 - to 21-fold [5]. HAP prolongs hospital stay by 7–9 days [6], increases treatment costs and is associated with a mortality of 20-60% [7, 8], especially if accompanied by severe sepsis [9, 10]. As many as half of all deaths related to nosocomial infections are due to nosocomial pneumonia [11]. The 2005 American Thoracic Society and Infectious Diseases Society of America (ATS/IDSA) guidelines define HAP as pneumonia occurring 48 hours or more after hospital admission, which was not incubating at the time of admission [7]. According to some authors, HAP may be further classified into early-onset pneumonia, arising within 48-96 hours after hospital admission and late-onset pneumonia occurring after day 5 of admission, with the latter being associated with a higher risk of infection with multidrug-resistant (MDR) pathogens [12].
The clinical diagnosis of HAP is based on the presence of a new lung infiltrate plus clinical evidence that the infiltrate is of infectious origin, which includes the new onset of fever, purulent sputum, leukocytosis, and decline in oxygenation [12, 13].
Pneumonia in ICU patients is mostly due to aspiration of microorganisms from the nasal, oropharyngeal or gastric flora [3]. The lower respiratory tract may be contaminated prior to ICU admission due to impaired barrier function of the upper respiratory tract, usually as a consequence of altered consciousness, trauma, surgery, or invasive airway management.
Gram-negative bacilli account for more than 50% of commonly isolated pathogens in HAP. Notable among these are Klebsiella pneumoniae, Pseudomonas aeruginosa, Enterobacter spp. and Acinetobacter spp. [13–15]. Recent trends show an increase in the prevalence of HAP caused by MDR bacteria [2, 16, 17]. Inadequate initial antibiotic therapy is associated with a significant increase in mortality in ICU patients suffering from HAP [10, 18]. Targeted antibiotic therapy may only be started after the etiological agent is recognized and its susceptibility to antibiotics is determined. In clinical practice, however, it may be difficult to detect and correctly identify the etiological agent of HAP in a biological sample. The diagnosis and treatment of pneumonia are fully described in two sets of guidelines currently in force, American (IDSA/ATS 2016) and European (ERS/ESCMID/ESICM 2009) [12, 19]. The most direct and specific method of biological sample collection for microbiological examination is bronchoscopy-assisted protected specimen brushing (PSB) [20]. The technically most feasible sampling method is endotracheal aspiration (ETA) in intubated patients. The use of other bronchoscopy and puncture methods for obtaining samples from the lower respiratory tract is also beneficial [21]. However, the results may be ambiguous and these techniques may also pose a potential risk of injury for the patient. In an effort to find the optimal approach to biological sample collection that is technically and economically feasible as well as suf-ficiently sensitive, it is useful to compare different types of sampling methods. Study of their validity will aid in determining whether ETA, gastric aspirate or an oropharyngeal swab are comparable to the technically more difficult PSB. According to a current weak recommendation with low quality of evidence from the IDSA/ATS on the management of adult with HAP and ventilator-assisted pneumonia, treatment should be initiated according to the results of non-invasive sampling and semi-quantitative cultures [12].
MATERIALS AND METHODS
Study design
The study was designed as prospective and observa-tional. It was approved by the Ethics Committee of the University Hospital Olomouc and Faculty of Medicine and Dentistry, Palacký University in Olomouc on 25 June 2012 (reference number 98/12) and registered in the Clinicaltrials.gov database (ID: NCT03039998). Informed consent was not possible to obtain from participants prior their enrollment as the study was conducted on unconscious patients. Informed consent was obtained later from the participants that where in sufficient level of consciousness.
The primary endpoint was to assess the sensitivity and specificity of diagnostic sampling methods with respect to identification of the etiological agent of HAP and to determine the method with the highest sensitivity. The aim was to test the hypothesis that in addition to PSB, considered a gold standard, there is a technically simpler method of obtaining biological samples for identification of bacteria causing HAP.
Given the two-round system of sample collection, the possibility of detecting pathogens in the same group of patients on two occasions at a 72-hour interval was verified. The secondary endpoint was to determine how frequently identical etiological agents causing HAP were detected by various sampling methods.
Patients
The study group comprised patients hospitalized in the Department of Anesthesiology and Intensive Care Medicine, University Hospital Olomouc and Faculty of Medicine and Dentistry, Palacký University Olomouc between 1 March 2013 and 31 May 2015 who developed clinical signs of HAP. The diagnostic criteria for HAP included the presence of newly developed or progressive infiltrates on chest radiographs in patients hospitalized for 48 hours or more plus at least two other signs of respiratory tract infection such as temperature over 38 °C, purulent sputum, leukocytosis (WBC > 12 x 103/mm3) or leukopenia (WBC < 4 x 103/mm3), signs of inflammation on auscultation, cough and/or respiratory insufficiency (PaO2/FiO2 ≤ 300 mmHg). Another inclusion criterion was the need for tracheal intubation. The following basic patient data were recorded: gender (male/female) and age at enrolment (years). The detection of specific bacterial strains in four types of biological samples as well as the quantity of culture and identity of isolated bacterial strains were studied.
All the patients with basic internal as well as surgical disea-ses regardless of whether the disease was acute or chronic were enrolled into the study. The only decisive criterion was the length of stay in the hospital and fulfillment of the HAP criteria. The duration of hospitalization in the study was always longer than 48 hours to meet the criteria for at least early HAP. At the time of inclusion, all patients were mechanically ventilated. The length of mechanical ventilation was not monitored because the file was not divided into HAP/VAP (Ventilator-associated pneumonia).
In all patients, antibiotic treatment was initiated after diagnosis of HAP. The therapy was guided empirically and respected the guidelines’ principles for the treatment of nosocomial pneumonia [22], till the determing etiological agents and its sensitivity to antibiotics. After determination of the etiological agent of pneumonia the treatment was subjected to modification, escalation or deescalation.
Collection of samples for microbiological culture
Four types of biological material samples were collected at once in each patient at the time of enrolment, that is, within 24 hours from the appearance of clinical signs of HAP.
- Oropharyngeal swab (OS) – Samples were collected from the back wall of the oropharynx using a commercially available sample collection kit with a transport medium (Copan Diagnostics).
- Gastric aspiration (GA) – Ten milliliters of gastric juices were aspirated with an irrigation syringe from a nasogastric tube into a sterile plastic container at the end of the feeding interval, just prior to administering another dose of enteral nutrition.
- Endotracheal aspiration (ETA) – Samples were collected by aspiration of secretions from an orotracheal tube using a sterile closed collecting system, with subsequent rinsing of the suction catheter with 10 mL of sterile saline and closing of the test tube with a sterile stopper.
- Protected specimen brushing (PSB) – A flexible bronchoscope was introduced near the orifice of the segmental bronchus with the most marked opacities detected by high-resolution computed tomography. If secretion was visible in the subsegmental bronchus, the PSB catheter was advanced into this secretion, the protected brush was opened and the secretion was sampled from this area. If secretion was not bronchoscopically visible, the entire PSB catheter was advanced 2–3 cm from the bronchoscope tip and then the brush was advanced by another 2–4 cm into the relevant subsegmental bronchus. Then the brush was moved back and forth and rotated several times, retracted into the PSB catheter and this was removed from the bronchoscope. The distal portion of the PSB catheter was washed in 70% alcohol and cut.
Biological sample collection was performed in two rounds, immediately after enrollment of the patient into the study (Round A) and 72 hours later (Round B). In patients who were extubated during the first 72 hours after their enrolment, Round B samples were not obtained due to the impossibility of performing PSB and ETA sampling.
The two-round design of microbiological examinations was selected in the study design to increase the number of identified HAP bacterial agents and to evaluate the effect of antibiotic therapy (these data are not part of this work).
Sample processing
The time between collection of samples and their proces-sing in a microbiology lab did not exceed 30 minutes. The transport temperature range was 18–26 °C. Each sample was processed by a semi-quantitative method based on the four-quadrant streak technique using a calibrated loop, with the quantity of isolated microorganisms ranging between 102 and 1011 colony forming units (CFUs) per mL. The samples were processed by standard microbiological methods. The microorganisms were identified by MALDI--TOF mass spectrometry (Bruker Daltonics). If the same bacterial species were detected in more than one sample, their relationship and/or identity were determined.
Definition of positive findings
An etiological agent was considered relevant if the quantity exceeded 105 CFU/mL and 103 CFU/mL in ETA secretion and PSB samples, respectively [23]. Since there are no such thresholds for OS and GA samples, positive detection in these samples was not considered a confirmed etiological agent.
Detection of identical pathogens
To see whether bacterial pathogens detected by different biological sampling techniques were unique or not, DNA profiles of the tree most frequently cultured pathogens (Klebsiella pneumoniae, Burkholderia multivorans and Pseudomonas aeruginosa) were compared. In Klebsiella pneumoniae and Pseudomonas aeruginosa strains, the relationship and/or identity were determined by pulsed-field gel electrophoresis (PFGE) as described by Husičková et al. [24]. The similarity of isolates was determined in accordance with interpretation criteria by Tenover et al. [25]. Isolates of Burkholderia multivorans were typed using random amplification of polymorphic DNA (RAPD) as described by Mahenthiralingam et al. [26].
Statistical analysis
Excluded from the study were patients with incomplete data. Statistical analyses were performed using visual inspection of data and normality tests; redundancy of predictors was assessed by analyzing their associations (correlation for continuous variables and contingency table analysis for categorical variables). Standard descriptive statistics were used to summarize primary data; continuous variables were used to determine the confidence interval, median and range; and categorical variables were used to determine absolute and relative frequencies. The selection of variables for a multivariate model was based on univariate P < 0.1 and redundancy analysis of preselected predictors. All analyses were carried out at a P ≤ 0.05 level of statistical significance. The software used was SPSS 21 (IBM Corporation, 2012).
RESULTS
Patients
In Round A, comprising 56 patients, a total of 79 bacterial isolates were detected. Eight patients had no positive culture findings in any of their biological samples. In 15 patients, no bacterial pathogens meeting the above quantity criteria were isolated from ETA or PSB samples. Thus, a total of 33 patients were assessed, from whom 45 positive culture samples were obtained. Twenty-two patients had one etiological agent, ten patients had two agents and one patient had three agents (Fig. 1). Between Round A and Round B, two patients died and 15 patients were extubated. In Round B, 39 patients were included and 45 bacterial isolates were obtained. Five patients had negative lower respiratory tract findings. In 10 patients, the quantity of their agents did not reach the threshold. A total of 24 patients had significant findings in ETA or PSB samples comprising 31 agents. Eighteen patients had one etiological agent, five patients had two agents and one patient had three agents (Fig. 2).
Descriptive data
The mean age of patients was 67.2±14.8 years; the median age was 70 years. The group comprised 40 males (71.4%) and 16 females (28.6%). The mean BMI was 29.6 ± 6.2 kg/m2, range 18.4–44.1 kg/m2 and median 28.5 kg/m2.
Main results
In Rounds A and B, ETA sampling was able to detect 99.7% and 87.1% of positive findings of etiological agents causing HAP, respectively (93.4% for both rounds).
PSB sampling detected 35.6% and 35.4% of positive findings in Rounds A and B, respectively (35.5% for both rounds).
In the subgroup of patients with positive findings obtained by ETA or PSB sampling, identical agents were also detected by OS sampling in 62.2% and 54.8% in Rounds A and B, respectively (59.2% for both rounds). GA sampling detected the same agents in 66.6% and 41.9% in Rounds A and B, respectively (56.6% for both rounds).
Bacterial strains
A total of 12 different bacterial species were detected in both rounds of sample collection (Table 1).
1. Frequency of bacterial species – etiological agents causing HAP (A – Round A, B – Round B) 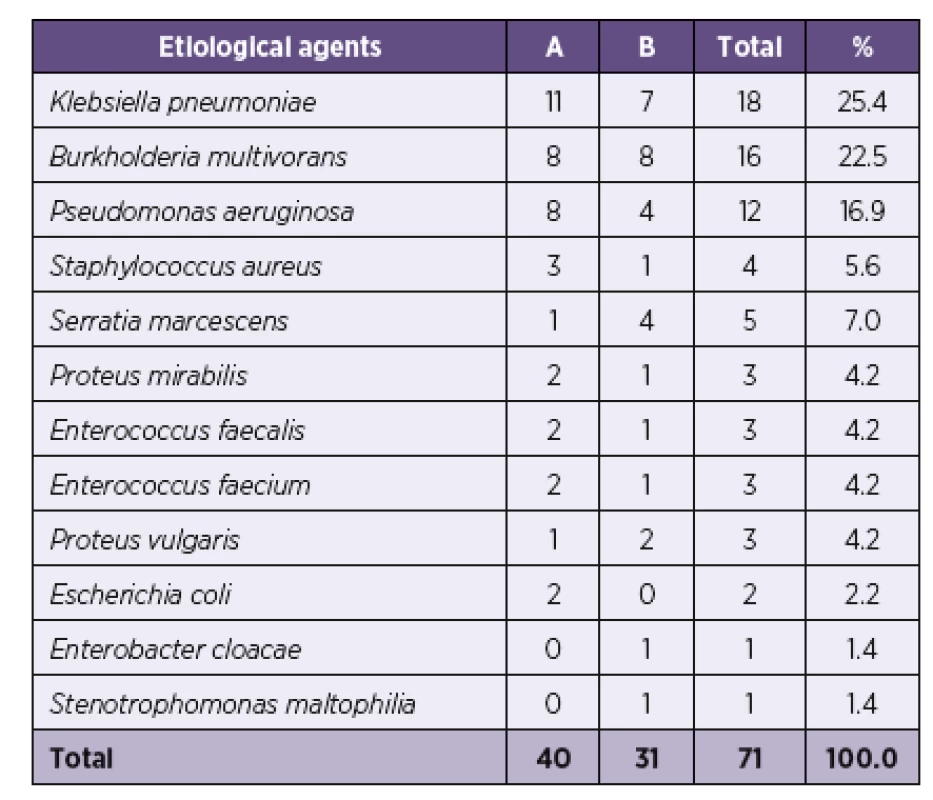
In biological samples obtained from the lower respiratory tract, the most frequently identified strains were those of Klebsiella pneumoniae (25.4%), Burkholderia multivorans (22.5%) and Pseudomonas aeruginosa (16.9%). The three species accounted for 56.2% of agents detected in ETA samples, 56.0% of agents in PSB samples, 64.6% in OS samples and 70.7% in GA samples (Table 2).
2. Positive culture findings obtained by ETA, PSB, OS and GA sampling in Rounds A and B 
ETA – endotracheal aspiration, PSB – protected specimen brushing, OS – orotracheal swab, GA – gastric aspiration, A – Round A, B – Round B In the subgroup of patients in whom Klebsiella pneumoniaewas found to be the etiological agent causing HAP, identical strains were found by ETA+GA in 12 patients (66.6%), by ETA+OS in 10 patients (55.5%), by ETA+PSB in 4 patients (22.2%), by PSB+GA in 5 patients (27.7%) and by PSB+OS in 4 patients (22.2%).
In case of Pseudomonas aeruginosa, identical strains were detected by ETA+GA in 8 patients (66.6%), by ETA+OS in 8 patients (66.6%), by ETA+PSB in 4 patients (33.3%), by PSB+GA in 2 patients (16.7%) and by PSB+OS in 2 patients (16.7%).
In case of Burkholderia multivorans, identical strains were found by ETA+GA in 6 patients (37.5%), by ETA+OS in 11 patients (68.7%), by ETA+PSB in 3 patients (18.7%), by PSB+GA in no patient (0%) and by PSB+OS in 2 patients (12.5%), see Table 3.
3. Simultaneous detection of identical etiological agents causing HAP by Round A and B sample collection (three most frequent agents) 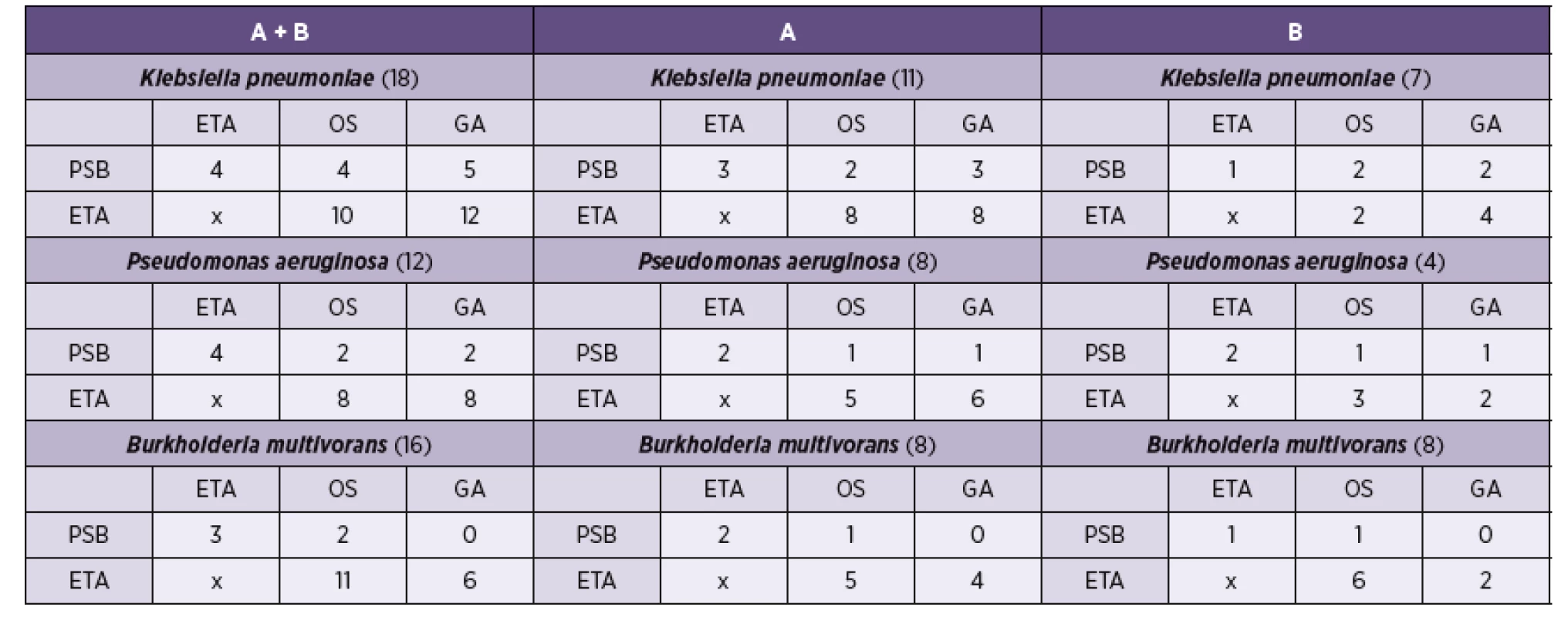
The number of identical isolates in the individual pairs of examined clinical specimens is reported. Among the 24 patients with significant bacterial pathogens detected in Round B (72-hour interval), 16 patients (66.7%) were found to have bacterial pathogens isolated from the lower respiratory tract (ETA or PSB) identical to those detected in Round A.
The sensitivity and specificity of the sample collection methods and their combinations are shown in Tables 4 and 5.
4. Sampling method sensitivity 
5. Sampling method specificity 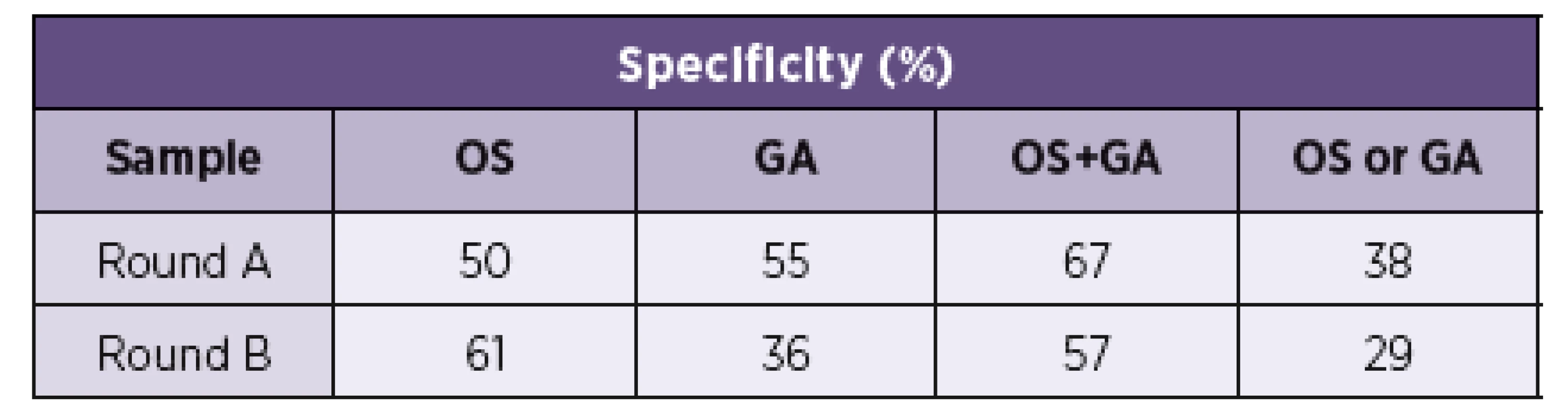
DISCUSSION
The present study provides data comparing the validity, sensitivity and specificity of commonly available and technically straightforward biological sample collection methods used to screen for HAP with PSB sampling, a costly, time-consuming reference method requiring more sophisticated equipment that is considered the gold standard. ETA, OS and GA are generally believed to have low validity, being unable to detect the etiological agents of pneumonia with sufficient sensitivity [20]. By contrast, PSB has been reported to have sensitivity and specificity rates of 33-100% and 50–100%, respectively [20, 27, 28]. This is in contrast with the present study’s results showing that the frequency of pathogen detection was considerably higher when samples were obtained by ETA (93.4%) as compared with OS (59.2%), GA (56.6%) and, in particular, PSB (35.5%). Because of the targeted nature of PSB, contamination of samples with upper respiratory tract microflora is almost impossible. PSB is also considered a standard method for identification of lower respiratory tract pathogens [20]. Yet it is seldom used in routine practice due to technical, time and financial reasons.
In addition, it should be noted that the recent literature on ETA sets a threshold of 106 CFU/ml, which is considered to be sufficiently sensitive and with the corresponding specificity [29]. However, the present study was conducted in 2013-2015 and worked with current data that determined the threshold at 105 CFU/ml.
In this context, however, it is appropriate to add that the present study was working with a group of patients with clinically unambiguously proved HAP, and in some pa-tients a bacterial agent level of 105 CFU/ml was detected, with an identical isolate present in PSB. This demonstrates the need for a rational assessment of all clinical and microbiological data and the clinical significance of the indicated quantity of 105 CFU/ml. In this context, the authors consider that this work of Czech authors can contribute to a modified view of the evaluation of the quantity of bacterial pathogens in HAP. It has to be taken into consideration whether the low frequency of pneumonia etiological agent detected by the PSB method was not only due to the fact that the other methods used identified the colonizing agent in supra-treshhold value. However, if the patient fulfils criteria of HAP according to guidelines, a positive bacterial cultivation of 106 CFU/ml is found in ETA, and identical bacteria are present e.g. in gastric lavage, but PSB is negative, of course, it can be assumed that the bacteria in ETA and GA is a colonizer. Nevertheless, the authors believe that the more likely explanation is simply the presence of false negativity of PSB. And this is the main message of the work presented.
In case of the three most commonly isolated bacteria, Klebsiella pneumoniae, Burkholderia multivorans and Pseudomonas aeruginosa, identical pathogens were most frequently detected by ETA+GA and ETA+OS. This is consistent with the notion that HAP is often caused by translocation of the patient’s primary or secondary microflora into the lower respiratory tract. Thus, microbiological examination using OS and GA may be a valuable complement to ETA and aid in identifying the bacterial pathogens. In the present study, the sample collection methods used for screening were also selected based on the assumption that pathogens causing HAP originate from the patient’s bacterial (in particular secondary) microflora [2]. However, there is no consistency as to whether the transmission is via the upper respiratory tract or from the gastrointestinal tract [30]. The present results suggest that for clinical practice, ETA is an ideal screening method for detecting the etiological agents causing HAP. Assuming that PSB is a standard approach in the diagnosis of HAP and ETA is a screening test, the sensitivity and specificity of ETA were 92% and 41%, respectively. The positive and negative predictive values were 29% and 95%, respectively. When comparing the ETA and PSB results with clinical signs (all patients clearly suffered from bacterial lung disease), the sensitivity rates were 66% for ETA and only 17% for PSB.
The methods for biological sample collection may be classified into non-targeted, or blind, and targeted, or bronchoscopy-guided. Both targeted and non-targeted sampling may be carried out in a protected manner to reduce the risk of contamination. In clinical practice, bacterial pathogens causing HAP are routinely isolated and identified using non-targeted sample collection, with sensitivity and specificity rates of 38–87% and 31–92%, respectively, for ETA and 58–96% and 71–100%, respectively, for blind PSB [21]. In non-ventilated patients, the most common technique is sputum sampling. However, its validity is severely compromised by a drawback common to all non-targeted types of sampling, that is, potential contamination of samples by bacteria that primarily or secondarily colonize the upper respiratory tract, resulting in potential false-positive findings. Although clinical practice guidelines recommend a blood culture in febrile patients with signs of pneumonia, the sensitivity of this test is low given the etiological agents causing HAP [31]. Apart from ETA, other non-targeted techniques are used to obtain samples from the lower respiratory tract, such as blind bronchial suction, blind bronchoalveolar lavage (BAL), blind PSB, blind protected BAL (pBAL), miniBAL or a blind plugged telescopic catheter (PTC). However, these sample collection techniques are rarely routinely used in intubated patients, the only exception being ETA. According to some authors, the yields of non-bronchoscopic and bronchoscopic methods are comparable [32, 33], even though bronchoscopic methods are associated with higher quantities of microbiology examinations. The concordance between non-bronchoscopic and bronchoscopic methods is approximately 80%. A certain proportion of non-bronchoscopic examinations yields false negative results, especially in left lung involvement [34]. However, the present study failed to confirm this as well as conclusions from a study by Chastre et al., stating that bronchoscopic methods are able to identify 80% of all bacteria in the lungs with considerable correlation with lung tissue biopsy [34]. Consistently with the present study, other studies questioned the usefulness of targeted sample collection since results of PSB sampling from the site of infiltration as shown by radiography and simultaneous blind PSB sampling from the contralateral lung were identical in 53% of cases [35]. Another study showed that results of repeated BAL from the same lung region were identical in 75% of cases [36]. Similarly, targeted diagnostic sample collection other than PSB, namely bronchoscopy-guided BAL, pBAL and PTC, are not routinely used in clinical practice. Moreover, bronchoscopy-guided methods place more stress on pa-tients and may not even provide clear results [37]. A 2005 meta-analysis failed to show the effect of bronchoscopic methods on reduced mortality [21].
The main limitation of the present study is a relatively small group of patients. Despite their best efforts, the investigators failed to include all consecutive patients meeting the above criteria during the study period. This was due to the fact that bronchoscopy was less accessible outside normal working hours and on the weekend. On the other hand, the fact that PSB sample collection was exclusively performed by experienced bronchoscopists routinely using the method considerably reduced the chance for erroneous sampling and thus false negative results. The group comprised a relatively high proportion of patients with no bacterial pathogen detected in sufficient quantity in any of their samples. The explanation may be that either the tests were performed at the time of early infection with a low number of bacteria or the pathogen had been eradicated by early adequate initial antibiotic therapy administered prior to randomization of the patient. Other reasons may be ETA sample collection from the unaffected lung region, irregular distribution of the pathogen in secretions from the lower respiratory tract or low amounts of material obtained by PSB.
A considerable proportion of GA and OS samples were culture-positive even in patients with negative ETA or PSB findings. Although the positive findings suggested a potential agent causing pneumonia in patients showing clinical signs of lung inflammation, these isolates could not be classified as HAP pathogens. The results add to the discussion about whether it is useful and sensible to consider possible positive culture findings in OS and GA samples in the diagnosis of HAP pathogens if the patient shows clinical signs of lung infection while the pathogen was not detected in PSB or ETA samples or these samples could not be collected in the patient without airway management. As seen from the study results, such considerations are supported by the high proportion of identical ETA/PSB and OS/GA findings.
Another limitation of the study is that the previous antibiotic therapy was not evaluated.
CONCLUSION
The study shows that blind endotracheal aspiration appears to be an optimum method for obtaining biological samples for identification of etiological agents causing HAP in intubated patients. The approach is not only technically feasible and easy to use in clinical practice but also sensitive enough. Microbial etiological agents causing HAP were more frequently detected in ETA samples than in those collected by PSB. If the results are ambiguous or negative, samples may be collected by oropharyngeal swabs and/or gastric aspiration as these techniques followed ETA in terms of the frequency of pathogen detection. Of the four types of biological sample collection compared in the study, PSB was the least successful in detecting HAP pathogens.
Do redakce došlo dne 13. 5. 2017.
Adresa pro korespondenci:
MUDr. Radovan Uvízl, Ph.D.
KARIM FN Olomouc
I.P. Pavlova 6
779 00 Olomouc
e-mail: radovan.uvizl@seznam.cz
Sources
1. Vincent JL, Bihari DJ, Suter PM, et al. The prevalence of nosocomial infection in intensive care units in Europe. Results of the European Prevalence of Infection in Intensive Care (EPIC) Study. EPIC International Advisory Committee. JAMA, 1995;274(8):639–644. doi:10.1001/jama.1995.03530080055041.
2. Uvizl R, Hanulik V, Husickova V, et al. Hospital-acquired pneumonia in ICU patients. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub, 2011;155(4):373–378. doi: 10.5507/bp.2011.067.
3. File TM Jr, Bartlett JG, Thorner AR. UpToDate Epidemiology, pathogenesis, microbiology, and diagnosis of hospital-acquired, ventilator-associated pneumonia in adults. [cit. 2017-03-20] available from: https://www.uptodate.com/contents/epidemiology-pathogenesis-microbiology-and-diagnosis-of-hospital-acquired-and-ventilator-associated-pneumonia-in-adults.
4. Richards MJ, Edwards JR, Culver DH, et al. Nosocomial infections in medical intensive care in the United States. Nosocomial Infections Surveillance System. Crit Care Med, 1991;27 : 887–892.
5. Fabregas N, Ewig S, Torres A, et al. Clinical diagnosis of ventilator associated pneumonia revisited: comparative validation using immediate post-mortem lung biopsies. Thorax, 1999;54(10):867–873.
6. Chastre J, Fagon JY. Ventilator-associated pneumonia. Am J Respir Crit Care. 2002;165(7):867–903. doi: 10.1164/ajrccm.165.7.2105078.
7. American Thoracic Society, Infectious Diseases Society of America. Guidelines for the management of adults with hospital-acquired, ventilator-associated, and healthcare-associated pneumonia. Am J Respir Crit Care Med, 2005;171 : 388–416. doi: 10.1164/rccm.200405-644ST.
8. Craven DE, Palladino R, McQuillen DP. Healthcare-associated pneumonia in adults: management principles to improve outcomes. Infect Dis Clin North Am, 2004;18(4):939. doi: 10.1016/j.idc.2004.08.001
9. Uvizl R, Adamus M, Cerny V, et al. Patient survival, predictive factors and disease course of severe sepsis in Czech intensive care units: a multicentre, retrospective, observational study. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub, 2016;160(2):287–297. doi: 10.5507/bp.2015.052.
10. Herkel T, Uvizl R, Adamus M, et al. Epidemiology of hospital-acquired pneumonia: results of a Central European multicenter, prospective, observational study compared with data from the European region. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub, 2016;160(3):448–455. doi: 10.5507/bp.2016.014.
11. Schulgen G, Kropec A, Kappstein I, et al. Estimation of extra hospital stay attributable to nosocomial infections: heterogenity and timing ef events. J Clin Epidemiol, 2000;53(4):409–417. doi: 10.1016/S0895-4356(99)00182-1.
12. Kalil AC, Metersky ML, Klompas M, et al. Management of Adults With Hospital-acquired and Ventilator-associated Pneumonia: 2016 Clinical Practice Guidelines by the Infectious Diseases Society of America and the American Thoracic Society. Clin Infect Dis, 2016;63(5): 575–582. doi: 10.1093/cid/ciw504e61-e111.
13. Kasper DL, TR Harrison. Hospital-Acquired (Nosocomial) Pneumonia. In: Harrison´s Principles of Internal Medicine. 16th edn. New York: McGraw-Hill; Medical Pub. Division; 2005 pp 1538–1541.
14. Jones RN. Microbial etiologies of hospital-acquired bacterial pneumonia and ventilator-associated bacterial pneumonia. Clin Infect Dis, 2010;Suppl 1:S81–87. doi: 10.1086/653053.
15. Sievert DM, Ricks P, Edwards JR. Antimicrobial-resistant pathogens associated with healthcare-associated infections: summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009-2010. Infect Control Hosp Epidemiol, 2013;34 : 1–14.
16. Jones RF. Microbial etiologies of hospital-acquired pneumonia and ventilator-associated bacterial pneumonia. Clin Index Dis, 2010;51(1 Suppl):81–87. doi: 10.1086/653053.
17. Hanulík V, Uvízl R, Husičková V, et al. Pneumonia-causing bacterial pathogens in intensive care patients. Klin Mikrobiol Inf Lek, 2011;17(4):135–140.
18. Irequi M, Ward S, Sherman G, et al. Clinical importance of delays in the initiation of appropriate antibiotic treatment for ventilator-associated pneumonia. Chest, 2002; 122(1):262–268. doi: 10.1378/chest.122.1.262.
19. Torres A, Ewig S, Lode H, et al. European HAP working group. Defining, treating and preventing hospital acquired pneumonia: European perspective. Intensive Care Med, 2009;35 : 9–29. doi: 10.1007/s00134-008-1336-9.
20. Baughmann RP. Protected-specimen brush technique in the diagnosis of ventilator-associated pneumonia. Chest, 2000;117(4/Suppl 2):203S–206S. doi:10.1378/chest.117.4_suppl_2.203S.
21. Campbell GD Jr. Blinded invasive diagnostic procedures in ventilator-associated pneumonia. Chest, 2000;117(4/Suppl 2):207S–222S. doi: 10.1378/chest.117.4_suppl_2.207S.
22. Herkeľ T, Uvizl R, Kolar M, Htoutou Sedlakova M, Adamus M, Doubravska L, Gabrhelik T, Pudova V, Langová K, Zazula R, Rezac T, Moravec M, Cermak P, Sevcik P, Stasek J, Sevcikova A, Hanslianova M, Turek Z, Cerny V, Paterova P. Health care associated pneumonia in intensive care patients – optimal settings for the initial empirical antimicrobial therapy: Reults of multicenter obsevational study Anest. intenziv. Med. 2017;28(3):154–162.
23. Joseph HM, Sistla S, Dutta TK, et al. Ventilator-associated pneumonia: A review. Eur J Intern Med, 2010;21 : 360–368. doi: 10.1016/j.ejim.2010.07.006.
24. Husickova, V., Cekanova, L., Chroma, M., et al. Carriage of ESBL - and AmpC-positive Enterobacteriaceae in the gastrointestinal tract of community subjects and hospitalized patients in the Czech Republic. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech Repub, 2012;156(4):348–353. doi: 10.5507/bp.2012.039.
25. Tenover, F.C., Arbeit, R.D., Goering, R.V., et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J. Clin. Microbiol, 1995;33(9):2233–2239.
26. Mahenthiralingam E, Campbell ME, Henry DA, et al. Epidemiology of Burkholderia cepacia infection in Patients with Cystic Fibrosis: Analysis by Random Amplified Polymorphic DNA Fingerprinting. Journal of Clinical Microbiology, 1996;34 : 2914–2920.
27. Shorr AF, Sherner JH, Jackson WL, et al. Invasive approaches to the diagnosis of ventilator-associated pneumonia: a meta-analysis. Crit Care Med, 2005;33 : 46–53. doi: 10.1097/01.CCM.0000149852.32599.31.
28. Rea-Neto A. Youssef NCM, Tuche F, et al. Diagnosis of ventilator-associated pneumonia: a systematic review of the literature. Critical Care, 2008;12:R56. doi: 10.1186/cc6877.
29. Kollef MH. Ventilator-Associated Pneumonia: the role of emerging therapies and diagnostics. Chest, 2015;147 : 1448–1450. doi: 10.1378/chest.14-2745.
30. Koleff MH, Morrow LE, Niederman MS. Clinical Characteristics and Treatment Patterns Among Patients With Ventilator-Associated Pneumonia. Chest, 2006;129 : 1210–1218. doi: 10.1378/chest.129.5.1210.
31. Fujitami S, Sun HY, Yu VL, et al. Pneumonia due to Pseudomonas aeruginosa: part I: epidemiology, clinical diagnosis, and source. Chest, 2011;39 : 909–919. doi: 10.1378/chest.10-0166.
32. Kowalzcyk W, Rybicky Z, Tomaszewsky D, et al. The comparison of different bronchial aspirate culturing methods in patients with ventilator-associated pneumonia. Anaesthesiology Intensive Therapy, 2001;43(2):64–68.
33. Clec’h C, Jauréguy F, Hamza L, et al. Agreement Between Quantitative Cultures of Postintubation tracheal Aspiration and Plugged Telescoping Catheter, Protected Specimen Brush, or BAL for the Diagnosis of Nosocomial Pneumonia. Chest, 2006;130 : 956–961. doi: 10.1378/chest.130.4.956.
34. Chastre J, Combes A, Luyt CE. The Invasive (Quantitative) Diagnosis of Ventilator-Associated Pneumonia. Respir Care, 2005;50(6):797–807.
35. Butler KL, Best IM, Oster RA, et al. Is bilateral protecter specimen brush sampling necessary for the accurate diagnosis of ventilator-associated pneumonia. J Trauma, 2004;57 : 316–322. doi: 10.1097/01.TA.0000088858.22080.CB.
36. Gerbeaux P, Ledorav V, Boussuges A, et al. Diagnosis of nosocomial pneumonia in mechanically ventilated patients: repeatability of the bronchoalveolar lavage. Am J Respir Crit Care Med, 1998;157 : 76–80.
37. Porzecanski I., Bowton DL. Diagnosis and Treatment of Ventilator-Associated Pneumonia. Chest, 2006;130 : 597–604. doi: 10.1378/chest.130.2.597.
Labels
Hygiene and epidemiology Medical virology Clinical microbiology
Article was published inEpidemiology, Microbiology, Immunology

2017 Issue 4-
All articles in this issue
- The prevalence, incidence, persistence and transmission ways of human papillomavirus infection (HPV)
- Flow cytometry in microbiology
- Crohn’s disease and ulcerative colitis – current view on genetic determination, immunopathogenesis and biologic therapy
- Mycological diagnosis of pulmonary Aspergillus infections with a focus on serological methods
- Human alveolar echinococcosis and an overview of the occurrence of Echinococcus multilocularis in animals in the Czech Republic
- Detection of the etiological agents of hospital-acquired pneumonia – validity and comparison of different types of biological sample collection: a prospective, observational study in intensive care patients
- Detection of antigen-specific T cells in patients with neuroborreliosis
- Epidemiology, Microbiology, Immunology
- Journal archive
- Current issue
- Online only
- About the journal
Most read in this issue- Human alveolar echinococcosis and an overview of the occurrence of Echinococcus multilocularis in animals in the Czech Republic
- Crohn’s disease and ulcerative colitis – current view on genetic determination, immunopathogenesis and biologic therapy
- Flow cytometry in microbiology
- Mycological diagnosis of pulmonary Aspergillus infections with a focus on serological methods
Login#ADS_BOTTOM_SCRIPTS#Forgotten passwordEnter the email address that you registered with. We will send you instructions on how to set a new password.
- Career



