-
Medical journals
- Career
Predicted strain coverage of a new protein-based meningococcal vaccine in the Czech Republic
Authors: P. Křížová 1; M. Musílek 1; Z. Vacková 1; J. Kozáková 1; H. Claus 2; U. Vogel 2; D. Medini 3
Authors‘ workplace: Centre for Epidemiology and Microbiology, National Institute of Public Health, Prague, Czech Republic 1; University of Würzburg, Institute for Hygiene and Microbiology, Würzburg, Germany 2; Novartis Vaccines and Diagnostics, Siena, Italy 3
Published in: Epidemiol. Mikrobiol. Imunol. 63, 2014, č. 2, s. 103-106
Category: Review articles, original papers, case report
Overview
Background:
Recent data indicate that Neisseria meningitidis B strains cause about 70% of invasive meningococcal disease (IMD) cases in Europe and the availability of a vaccine effective against N. meningitidis B is desirable. A new protein-based MenB vaccine was licensed for use in Europe in January 2013. Meningococcal antigen typing system (MATS) was developed to predict strain coverage of this vaccine. Reports have recently been published for a European consortium, including aggregated data for the Czech Republic. The aim of this paper is to provide a detailed breakdown of MATS results for the Czech N. meningitidis B isolates.Materials and methods:
One hundred and eight N. meningitidis B isolates from IMD collected in the Czech Republic during 2007–2010 were selected. MATS analysis was done according to the method previously published.Results:
Based on MATS analysis, the overall estimate of strain coverage of the new MenB vaccine for a panel of 108 Czech N. meningitidis B strains is 74% (95% CI: 59–87%). Thirty-nine strains (36%) are predicted to be covered by a single antigen and 41 strains (38%) by more than one antigen. For 28 strains (26%), no antigen coverage was found.Conclusions:
MATS analysis showed that the new protein-based MenB vaccine could protect against a substantial proportion of IMD caused by N. meningitidis B in the Czech Republic. Continued detailed surveillance of IMD will be essential if the MenB vaccine is introduced to the country.Keywords:
Neisseria meningitidis B – MenB vaccine – vaccine coverage – MATS – Meningococcal Antigen Typing SystemINTRODUCTION
Invasive meningococcal disease (IMD) is one of the most devastating infectious diseases causing a high case fatality rate at times of advanced medical care. Thanks to the development and implementation of conjugate vaccines against meningococcal serogroups A, C, Y, and W135, the incidence of IMD cases caused by serogroup C has decreased in Europe and recently, serogroup B has become the leading cause of IMD. Neisseria meningitidis B strains have been reported to cause about 70% of IMD cases in Europe [1] and about 40–50% of IMD cases in the USA [2–4]. For this reason, the availability of a vaccine effective against N. meningitidis B (MenB vaccine) is desirable. However, the most significant challenges are the diversity of N. meningitidis B population and poor immunogenicity of serogroup B capsule. Various attempts have been made to develop a MenB vaccine effective against the whole N. meningitidis B population and reverse vaccinology represents a new approach [5–7]. This method started by whole genome sequencing of N. meningitidis and more than 600 antigen candidates were identified. The final version of the new four-component MenB vaccine (4CMenB) combines three primary recombinant components: fHbp (factor H binding protein) fusion protein, NadA (Neisserial adhesin A) and NHBA (Neisserial Heparin Binding Antigen) fusion protein, and the outer membrane protein PorA P1.4 of the strain causing the New Zealand outbreak. This new MenB vaccine (BEXSERO) was licensed for use in Europe in January 2013.
Meningococcal antigen typing system (MATS) ELISA was developed to predict meningococcal strain coverage of the protein-based meningococcal vaccine 4CMenB – Figure 1 [8]. MATS is considered as correlated with bactericidal activity and allows typing of large panels of strains.
Fig 1. Schematic of the MATS ELISA method (A) MenB bacteria are grown overnight on chocolate agar. (B) A suspension of bakteria taken from the plate is prepared to a specifield OD600. (C) detergent is added to the suspension to extract the capsule and expose the antigens. (D) Serial dilutions of extract are tested in the MATS ELISA. A specific capture antibody (yellow) binds one of the antigens (example: fHbp, blue) from the extract, which is then detected with a specific biotin-labeled antibody (yellow and purple) and a streptavidin-enzyme conjugate (green and gold). (E) Plates are read at 490 nm in an ELISA reader. (F) Results are calculated by comparing the curve of OD490 vs. dilution obtained with the serially diluted unknown strain to a serially diluted reference strain tested in the same ELISA plate. Obr. 1. Schematické znázornění metody MATS ELISA [8] (A) Bakterie meningokoka B se kultivují přes noc na čokoládovém agaru. (B) Připraví se bakteriální suspenze odpovídající zadané optické denzitě OD600. (C) Do suspenze se přidá detergent, jehož působením se extrahuje pouzdro a dojde k uvolnění antigenů. (D) Sériová ředění extraktu se testují pomocí typizačního systému MATS ELISA. Specifická vychytávací protilátka (žlutá) na sebe naváže jeden z antigenů (například fHbp, modrá) z extraktu, který je následně detekován specifickou protilátkou značenou biotinem (žlutá a fialová), na kterou se naváže konjugát streptavidinu s enzymem (zelená a zlatá). (E) Destičky se odečtou při 490 nm pomocí čtečky ELISA. (F) Provede se výpočet výsledků na základě porovnání křivky pro OD490 proti ředění sériově naředěného neznámého kmene se sériově naředěným referenčním kmenem testovaným na stejné destičce. ![Fig 1. Schematic of the MATS ELISA method (A) MenB bacteria are grown overnight on chocolate agar. (B) A suspension of bakteria taken from the plate is prepared to a specifield OD600. (C) detergent is added to the suspension to extract the capsule and expose the antigens. (D) Serial dilutions of extract are tested in the MATS ELISA. A specific capture antibody (yellow) binds one of the antigens (example: fHbp, blue) from the extract, which is then detected with a specific biotin-labeled antibody (yellow and purple) and a streptavidin-enzyme conjugate (green and gold). (E) Plates are read at 490 nm in an ELISA reader. (F) Results are calculated by comparing the curve of OD490 vs. dilution obtained with the serially diluted unknown strain to a serially diluted reference strain tested in the same ELISA plate.
Obr. 1. Schematické znázornění metody MATS ELISA [8] (A) Bakterie meningokoka B se kultivují přes noc na čokoládovém agaru. (B) Připraví se bakteriální suspenze odpovídající zadané optické denzitě OD600. (C) Do suspenze se přidá detergent, jehož působením se extrahuje pouzdro a dojde k uvolnění antigenů. (D) Sériová ředění extraktu se testují pomocí typizačního systému MATS ELISA. Specifická vychytávací protilátka (žlutá) na sebe naváže jeden z antigenů (například fHbp, modrá) z extraktu, který je následně detekován specifickou protilátkou značenou biotinem (žlutá a fialová), na kterou se naváže konjugát streptavidinu s enzymem (zelená a zlatá). (E) Destičky se odečtou při 490 nm pomocí čtečky ELISA. (F) Provede se výpočet výsledků na základě porovnání křivky pro OD490 proti ředění sériově naředěného neznámého kmene se sériově naředěným referenčním kmenem testovaným na stejné destičce.](https://pl-master.mdcdn.cz/media/image/969cee79081c79310f6bd07b995c5ff0.jpg?version=1537795405)
MATS was used to test a collection of N. meningitidis B isolates, mostly from the epidemiological year 2007/2008, from seven European countries: Germany, France, England & Wales, Italy, Norway, Spain, and the Czech Republic. The predicted coverage was 78% (95% CI 63–90), ranging from 73 to 87% in individual countries [9]. The aim of this paper is to provide a detailed breakdown of the MATS results for the Czech N. meningitidis B isolates.
MATERIALS AND METHODS
Neisseria meningitidis strain selection
One hundred and eight N. meningitidis B strains collected in the Czech Republic during 2007–2010 were selected. The original collection included 120 N. meningitidis B isolates but 12 strains that failed to grow in the laboratory were excluded from this study. The original collection comprised all serogroup B isolates from IMD cases obtained by the National Reference Laboratory for Meningococcal Infections of the National Institute of Public Health, Prague, Czech Republic within the above mentioned period, i.e. 69 CSF isolates, 46 blood culture isolates, four isolates from other sites, and one isolate from a nasopharyngeal swab.
Neisseria meningitidis strain characterisation
Serogrouping was performed by slide agglutination, using commercial antisera. PorA variable region typing was done by sequencing, according to the Neisseria Sequence Typing Home Page [10]. Multilocus sequence typing (MLST) was done in accordance with the pubMLST neisseria sequence typing database guidelines [10]. Serogroup, PorA variable regions, and MLST were determined by the National Reference Laboratory for Meningococcal Infections of the Czech Republic.
Meningococcal antigen typing system (MATS) ELISA
MATS analysis was done according to the method published previously [8]. MATS relative potencies for fHbp, NHBA, and nadA were calculated by comparison with the N. meningitidis B reference strains. The predicted strain coverage was defined as the proportion of strains with MATS relative potencies greater than the positive bactericidal thresholds (PBTs) [8, 9]. Novartis supplied with the MATS methodology the University of Wurzburg, Germany, which performed MATS testing of the Czech N. meningitidis B isolates. Quality-assured data from the laboratory of the University of Wurzburg were collected by Novartis, subjected to further quality control, and strains with complete (4 antigens) and valid data (according to MATS acceptance criteria, consistency of documentation and QA of the originating laboratory) were selected for analysis.
Statistical analysis
95% Confidence Intervals for the positive bactericidal thresholds were estimated on the basis of MATS precision (reproducibility) determined during inter-laboratory standardization [11]. CIs do not take into account the number of strains typed.
RESULTS
Based on MATS analysis, the overall estimate of strain coverage for all 108 N. meningitidis B Czech strains is 74% (95% CI: 59–87%). When considering strains predicted to be covered (i.e. those that express levels of any 4CMenB antigen above the PBT or harbour PorA VR2 4), a single antigen coverage is predicted for 39 strains (36%) and 41 strains (38% of the total Czech collection spanning 2007–2010) are predicted to be covered by more than one antigen – Figure 2. Thus, the redundancy ensured by the strategy of targeting bacteria with antibodies to multiple antigens may provide an additional measure of protection against antigen mutation or loss.
Fig 2. Coverage of N. meningitidis B strains by number of 4CMenB vaccine antigens, Czech Republic, 2007–2010 Obr. 2. Pokrytí kmenů N. meningitidis B vakcínou 4CMenB podle počtu antigenů, Česká republika, 2007–20010 
The distribution of PorA VR2 subtypes showed that P1.15 and P1.14 VR2 subtypes predominated in 2007-2010 (both 19%). A single strain expressed PorA P1.4 subtype, the immunodominant protein antigen contained in the OMV, and is considered covered by 4CMenB for the purpose of coverage estimation.
The coverage of strains in the panel based on the expression of fHbp, NadA, or NHBA above the MATS PBT or PorA P1.4 was then analyzed. Next, the nature of the combinations of multiple antigens present in strains that may act as targets for bactericidal antibodies elicited by 4CMenB vaccination was analysed in greater detail. The proportions of the total (n = 108) predicted to be covered based on the presence of the listed antigen(s) are shown in Table 1.
1. Coverage of N. meningitidis B strains by antigen combinationa, Czech Republic, 2007–2010 Tabulka 1. Pokrytí kmenů N. meningitidis B vakcínou 4CMenB podle kombinací antigenůa, Česká republika, 2007–2010 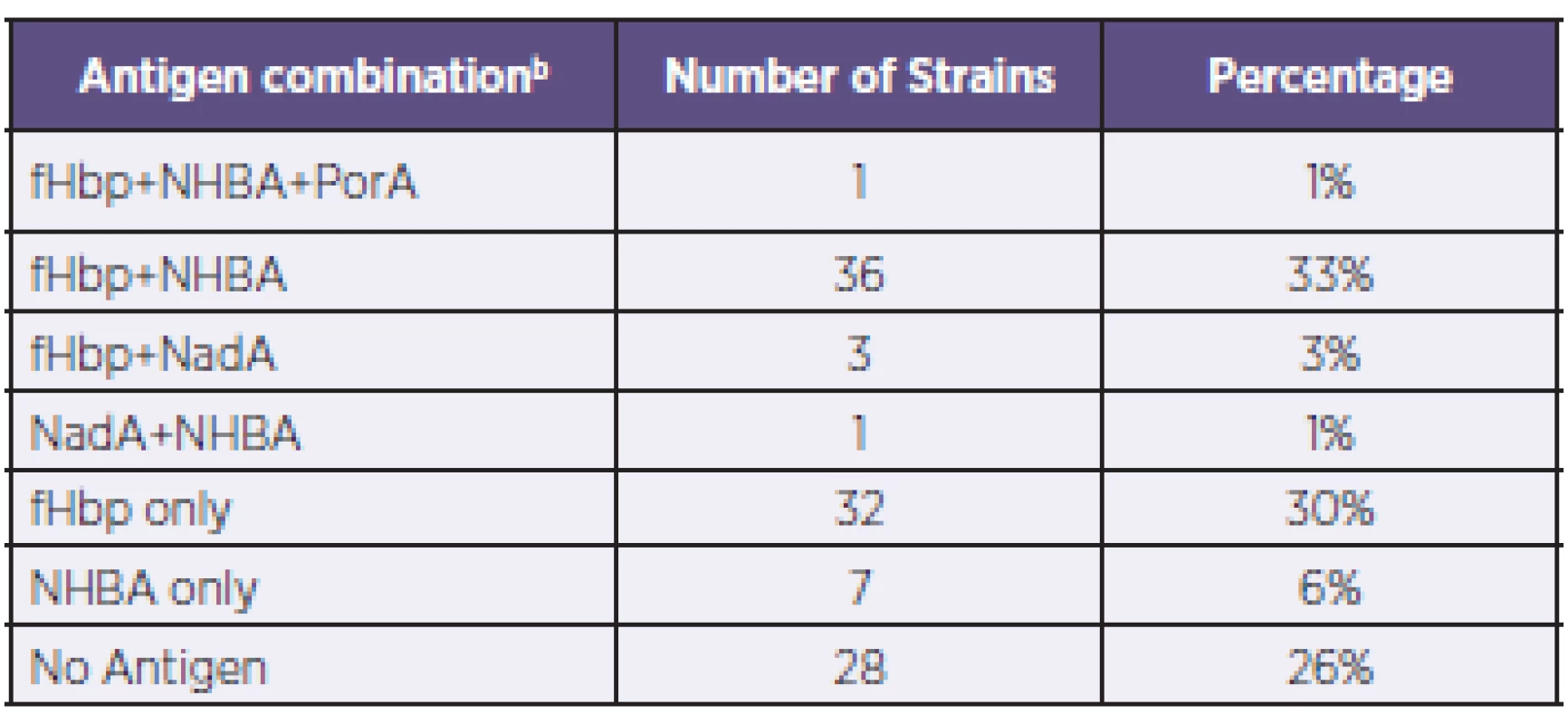
a: Only the actual combinations observed are presented. b: The antigen combinations listed are expressed at levels above the associated PBT. a: Uvedeny jsou pouze zjištěné kombinace. b: Uvedené kombinace antigenů jsou exprimovány v hladině přesahující spojený pozitivní baktericidní práh (PBT). Genotyping of strains was also conducted using MLST – Table 2, a standard typing system that relies on the direct sequencing of seven housekeeping genes that encode intracellular, cytoplasmic enzymes. Although MLST has limited utility in evaluating 4CMenB coverage, it provides information that can increase our understanding of the evolution of the bacterium and of the impact of mass vaccination on carriage and prevalence of the hypervirulent lineages. The most represented clonal complex (cc) in the Czech 2007–2010 collection is cc-41/44 in which 90% of strains are predicted by MATS to be covered. An equal proportion (19%) of strains have yet to be assigned to a clonal complex; of these, 76% are predicted by MATS to be covered.
2. Distribution of clonal complexes and coverage within each complex in the N. meningitidis B strains, Czech Republic, 2007–2010 Tabulka 2. Distribuce klonálních komplexů a jejich pokrytí vakcínou u kmenů N. meningitidis B, Česká republika, 2007–2010 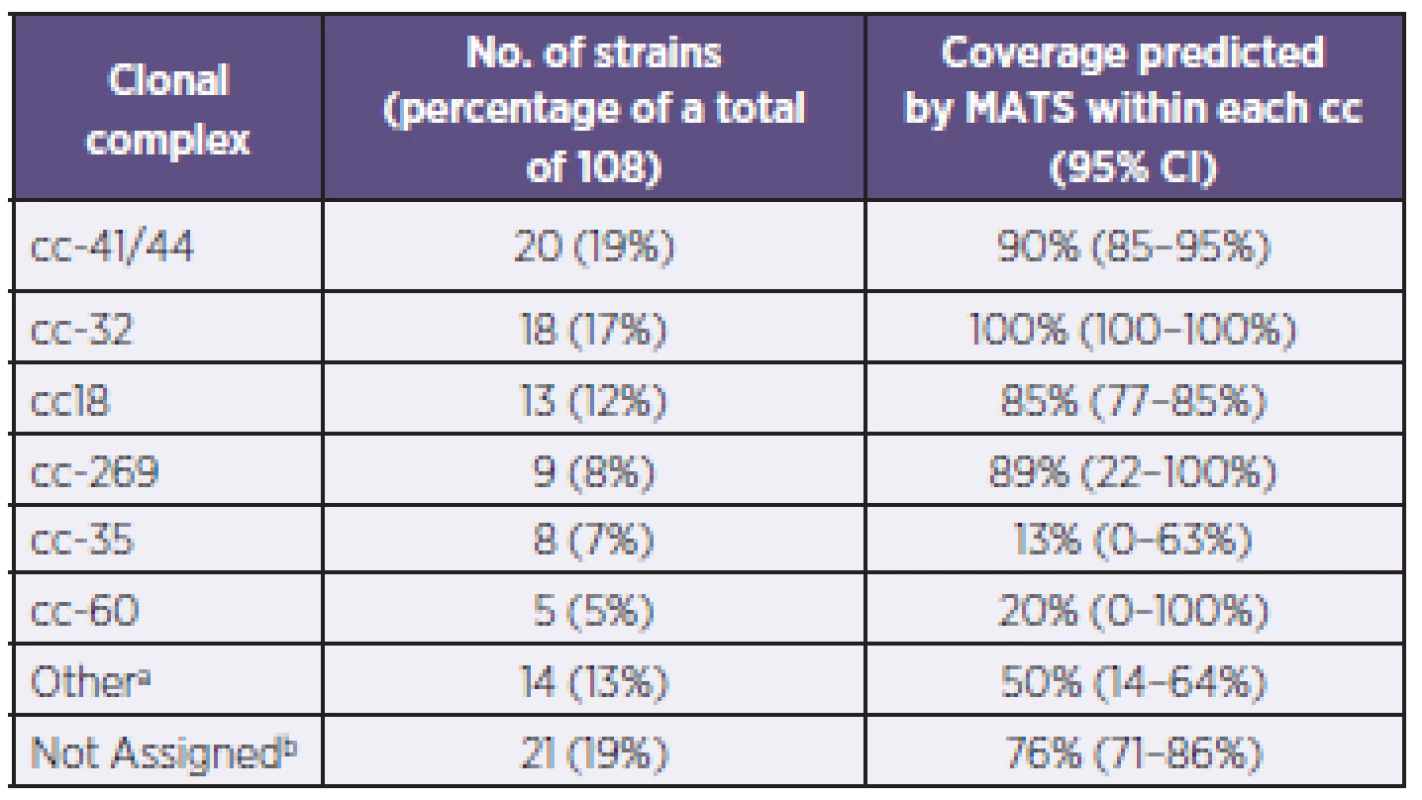
a: “Other”: refers to established ccs that have less than 5 strains within each cc. These include: cc-103 (3 strains), cc-1157 (2 strains), cc-174 (2 strains), cc-213 (2 strains), cc-865 (2 strains), cc-11, cc-162, and cc-461 (one strain each). b: “Not Assigned”: Sequence Type (ST) of strains in this collection has been identified or established, but does not belong to a clonal complex (cc) a: “jiný”: odkazuje na ustavené klonální komplexy o méně než 5 kmenech. Jedná se o cc-103 (3 kmeny), cc-1157 (2 kmeny), cc-174 (2 kmeny), cc-213 (2 kmeny), cc-865 (2 kmeny), cc-11, cc-162 a cc-461 (po jednom kmeni). b: “nezařazený”: Sekvenční typ (ST) kmenů v tomto souboru byl určen, ale nepatří k žádnému klonálnímu komplexu (cc) DISCUSSION AND CONCLUSION
As MATS does not account for the activity of bactericidal antibodies generated from non-PorA components of OMV or synergistic effects of multiple components of 4CMenB, this is considered to be a conservative estimate of vaccine strain coverage.
A recent investigation revealed that the MATS could underestimate 4CMenB vaccine strain coverage [12]. The authors have concluded that MATS is a conservative predictive tool. However, the EMGM (European Monitoring Group on Meningococci) recommends its use for the surveillance after implementation of the 4CMenB vaccine [13]. Considering that the performance of MATS has been established in few reference laboratories in Europe, these laboratories should provide MATS testing of selected isolates on request.
Based on MATS analysis of 108 invasive meningococcal serogroup B strains collected during 2007–2010 in the Czech Republic, an estimated 74% (95% CI: 59–87%) of strains are expected to be covered by bactericidal antibody responses to the 4CMenB vaccine. Of the strains predicted to be covered (n = 80), roughly half (41 strains) are likely to be covered by more than one antigen. Thirty-two strains (30%) were covered by the fHbp antigen only and 36 strains (33%) by combination of the fHbp + NHBA antigens. In 28 strains (26%) no antigen coverage was found.
Coverage by more than one antigen provides a measure of redundancy that ensures 4CMenB can still be effective even if one target antigen is mutated or down-regulated.
MATS analysis has shown that the new MenB protein-based vaccine could protect against a substantial proportion of IMD caused by N. meningitidis B in the Czech Republic. Continued detailed surveillance of IMD will be essential if the MenB vaccine is introduced to the country.
Acknowledgement: Sequence characterisation of N. meningitidis isolates was supported by grant IGA NT 11424-4/2010 of the Internal Grant Agency of the Ministry of Health of the Czech Republic.
Do redakce došlo dne 20.12. 2013.
Adresa pro korespondenci:
MUDr. Pavla Křížová, CSc.
Státní zdravotní ústav
Šrobárova 48
100 00 Praha 10
e-mail: pavla.krizova@szu.cz
Sources
1. http://www.ecdc.europa.eu/en/publications/Publications/Annual-Epidemiological-Report-2012.pdf. Access: December 29, 2013.
2. Harrison LH, Trotter CL, Ramsay ME. Global epidemiology of meningococcal disease. Vaccine, 2009;27(Suppl 2):B51–63.
3. Halperin SA, Bettinger JA, Greenwood,B et al. The changing and dynamic epidemiology of meningococcal disease. Vaccine, 2012;30(Suppl 2):B26–36.
4. Křížová P, Vacková Z, Musílek M, Kozáková J. Invazivní meningokokové onemocnění v České republice – surveillance a doporučení k vakcinační strategii. Epidemiologie, mikrobiologie, imunologie, 2013;62(4):138–147.
5. Rappuoli R. Reverse vaccinology. Curr Opin Microbiol, 2000;3(5):445–450.
6. Rappuoli R, Covacci A. Reverse vaccinology and genomics. Science, 2003;302(5645):602.
7. Serruto D, Bottomley MJ, Ram S et al. The new multicomponent vaccine against meningococcal serogroup B, 4CMenB: immunological, functional and structural characterization of antigens. Vaccine, 2012;30Suppl 2:B87–97.
8. Donnelly J, Medini D, Boccadifuoco G et al. Qualitative and quantitative assessment of meningococcal antigens to evaluace the potential strain coverage of protein-based vaccines. Proc Natl Acad Sci USA, 2010;107(45):19490–19495.
9. Vogel U, Taha MK, Vazquez JA et al. Predicted strain coverage of a meningococcal multicomponent vaccine (4CMenB) in Europe: a qualitative and quantitative assessment. Lancet Infect Dis, 2013;13(5):416–425.
10. http://pubmlst.org/neisseria/ Access: December 29, 2013.
11. Plikaytis BD, Stella M, Boccadifuoco G et al. Interlaboratory standardization of the sandwich enzyme-linked immunosorbent assay designed for MATS, a rapid, reproducible method for estimating the strain coverage of investigational vaccines. Clinical and Vaccine Imunology, 2012;19 : 1609–1617.
12. Frosi G, Biolchi A, Lo Sapio M et al. Bactericidal antipody against a representative epidemiological meningococcal serogroup B panel confirms that MATS underestimates 4CMenB vaccine strain coverage. Vaccine, 2013;31(43):4968–49674.
13. http://emgm.eu/downloads/Statement_of_the_EMGM_Society_on_Bexsero_final_Jan_25_13.pdf. Access: December 29, 2013.
Labels
Hygiene and epidemiology Medical virology Clinical microbiology
Article was published inEpidemiology, Microbiology, Immunology

2014 Issue 2-
All articles in this issue
- Phylogenetic and molecular analysis of A/H1N1pdm influenza viruses isolated in the epidemic season 2012/2013 from hospitalised patients with symptoms of influenza-like illness
- An increase in the prevalence of syphilis in women in Eastern Bohemia – 30 years of surveillance
- Diagnosis of Clostridium difficile infections: Comparative study of two immuno enzyme assays with confirmation by PCR and culture followed by PCR ribotyping
- A point prevalence survey of healthcare-associated infections in the Slovak Republic – a part of the EU project
- Nosocomial transmission of listeriosis
- Diversity of human Salmonella isolates in the South Moravian Region in 2009–2012
- Candida dubliniensis in clinical specimens and possibilities for identification
- Natural antibodies against α(1,3) galactosyl epitope in the serum of cancer patients
- Cytolethal distending toxins
- Evaluation of the importance of a ready-made, gentamicin-impregnated spacer in relation to bacteriological findings in patients with periprosthetic joint infections
- Q fever – an occupational disease leading to disability – case report
- Measles re-emerging in the Ústí Region
- Predicted strain coverage of a new protein-based meningococcal vaccine in the Czech Republic
- Post-mortem analysis of Candida albicans breakthrough infection during echinocandin treatment in haematopoietic stem cell transplant recipient
- Viral gastroenteritis in Eastern Bohemia Region of the Czech Republic
- Seroprevalence study of hepatitis E virus infection in two districts of the Czech Republic
- Epidemiology, Microbiology, Immunology
- Journal archive
- Current issue
- Online only
- About the journal
Most read in this issue- Candida dubliniensis in clinical specimens and possibilities for identification
- A point prevalence survey of healthcare-associated infections in the Slovak Republic – a part of the EU project
- Q fever – an occupational disease leading to disability – case report
- Diagnosis of Clostridium difficile infections: Comparative study of two immuno enzyme assays with confirmation by PCR and culture followed by PCR ribotyping
Login#ADS_BOTTOM_SCRIPTS#Forgotten passwordEnter the email address that you registered with. We will send you instructions on how to set a new password.
- Career

